How has data processing been improved in Progenesis QI for proteomics?
Progenesis QI for proteomics performs automatic reference run selection and alignment and reports alignment quality scores for your imported runs. This increases objectivity of analysis and frees-up your research time.
Reference run selection and alignment can be started simultaneously as you start to load your runs, so you can:
- Spend time away from the computer screen as Progenesis QI for proteomics automatically performs the first key data analysis steps, confident they are applied objectively every time
- Let the software select the reference run that gives the best alignment quality scores across all your runs for increased objectivity and reproducibility of data analysis
- Reduce automated processing time by selecting a single run or a limited set of runs, which routinely produce the best alignment scores, for the software to choose a reference from
- Review quality assurance scores for alignment so you can be confident that the next stage of automatic peak picking and normalisation will achieve the best analysis results possible
- Visualise alignment quality with a simple red/orange/green colour-coded grid overlaid across each run
- Define threshold scores for alignment to objectively reject poor quality runs and help improve reproducibility of results between experiments
- Immediately see the effect of adding manual alignment vectors has on the alignment scores, should editing be required
- Maintain a check on LC-MS system performance with each run displayed as an ion intensity map once loaded
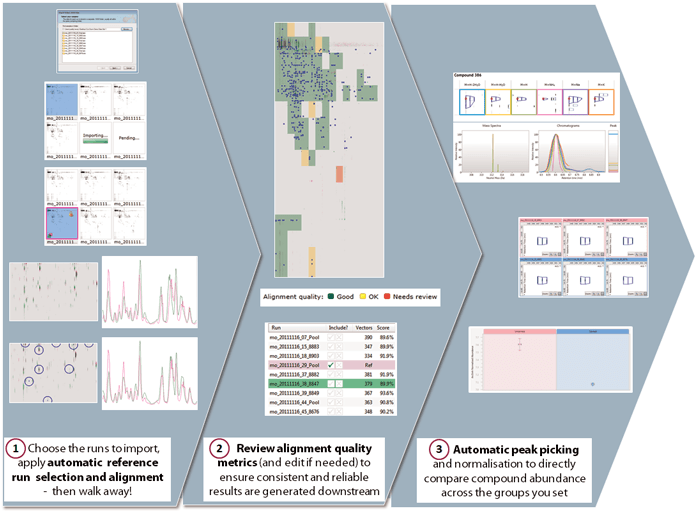
The workflow to show where the initial data processing steps, (1), have been automated and how alignment can be a simple review before applying automatic peak picking, a feature already available in previous versions of Progenesis QI for proteomics.