Why is alignment so important?
Alignment makes it possible to accurately compare the peptides in your runs by correcting any drift in retention times. This is done non-linearly across the entire run to give the accuracy needed for fast, statistically robust analysis.
Once alignment has been performed, each run in the experiment will be perfectly aligned so that an ion on run A will be in the same location as the matching ion in run B. The screenshots below show a single peptide ion in two runs being aligned. Each image shows both runs overlaid on each other; one run is displayed in green and the other is pink. When the runs are not yet aligned (shown on the left), the separate ions are clearly visible. After alignment (shown on the right), the pink and green ions combine to make black when perfectly overlaid.

Overlay before alignment
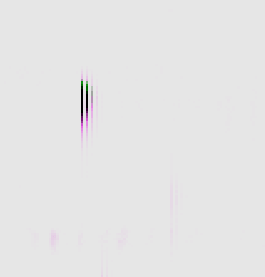
Overlay after alignment
A single set of peptide ion outlines is then added to all runs; as all runs are used during peak picking, compounds will be found, even if they are missing from one or more runs. This means that Progenesis measures a zero abundance only if the ion is truly missing. Peak picking in this manner, from an aggregate image, also reduces the instances of false positives.
Another benefit is that while good alignment means more efficient peak picking in terms of accuracy, it also speeds up software performance and reduces the demand on PC resources.
Alignment is arguably the most important part of the workflow, so it’s worth spending a bit of time at this stage to review the quality of your alignment and make adjustments where necessary.





