How do I use Progenesis LC‑MS to analyse fractionated biological samples?
Analysis of fractionated samples in Progenesis LC‑MS is a 2-step process:
- Create a series of experiments, each containing the sample runs from a single fraction. Once that is done, each experiment will contain its own set of identified proteins and peptides.
- Collate the proteins for each sample by creating a multi-fraction experiment into which you import the single-fraction experiments from the first step.
Let's look at these in a little more detail with a simple example.
Fractionating the samples
For illustrative purposes, let's consider an experiment in which gel fractionation has been used. In the experiment, protein expression in 3 subjects treated with a novel drug is being compared against expression in 3 control subjects. In an attempt to increase peptide coverage, each sample is electrophoretically fractionated in a 1D gel, producing 4 gel slices for each sample:
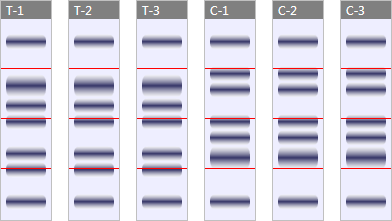
(In the simplified diagram above, the red lines indicate where the gel is sliced to produce the fractions. In reality, you may choose to slice between bands, but for this example, all fractions are of an equal size.)
Consequently, we'll end up with 4 runs for each sample, giving us a total of 24 sample runs going through our LC‑MS machine.
Analysing the separate fractions
Once we have collected the run data, we'll need to create 4 separate experiments in Progenesis LC‑MS, using the runs as shown here:
1st experiment:

2nd experiment:

3rd experiment:

4th experiment:

We'll refer to these experiments as our single-fraction experiments. To create each of the single-fraction experiments, you can do either of the following in the Progenesis LC‑MS start-up screen:
- Select the Perform analysis tab, and click the New button at the bottom-left of the screen
- Select the Combine analysed fractions tab, and click the Analyse a single fraction button
Those buttons are exactly equivalent; a new experiment is created and can be analysed in the standard Progenesis workflow (as described in the online tutorial). Which button you use is a matter of preference. Note, however, that the Perform analysis tab has the benefit of also listing all of your recently-used single-fraction experiments, helping you to keep track of your progress in analysing them.
Recombining the samples
When all single-fraction experiments have been fully analysed — including identification of proteins and the resolution of any peptide conflicts — we can recombine all our of protein information in one multi-fraction experiment.
Again, from the Progenesis LC‑MS start-up screen, select the Combine analysed fractions tab. Now, click the Recombine analysed fractions button at the left of the screen. This creates a new multi-fraction experiment and activates the Import Data screen at the start of the fractionation workflow:
From here, it's a simple matter of:
- Selecting and importing your single-fraction experiments
- Grouping the runs in the single-fraction experiments by the sample they contain
- Setting up the experimental conditions (e.g. treated vs. control)
The Review Proteins then gives you the tools to explore the statistics and protein behaviour for the whole experiment.





