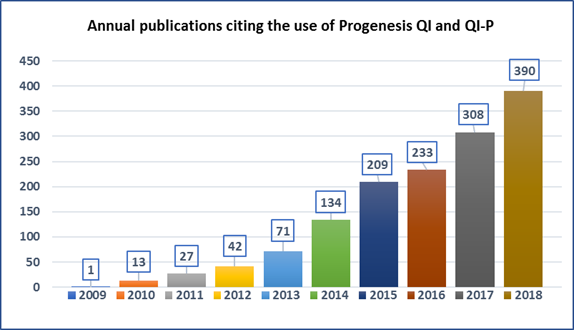
Progenesis QI for proteomics software references
2019
Abdulwahab RA, Alaiya A, Shinwari Z, Allaith AAA, Giha HA (2019) LCMS/MS proteomic analysis revealed novel associations of 37 proteins with T2DM and notable upregulation of immunoglobulins. Int J Mol Med 43:2118-2132
Abdulwahab RA, Allaith AAA, Shinwari Z, Alaiya A, Giha HA (2019) Association of TATA box-binding protein-associated factor RNA polymerase I subunit C (TAF1C) with T2DM. Gene 706:43-51
Al-Majdoub ZM, Al Feteisi H, Achour B, Warwood S, Neuhoff S, Rostami-Hodjegan A, Barber J (2019) Proteomic Quantification of Human Blood-Brain Barrier SLC and ABC Transporters in Healthy Individuals and Dementia Patients. Mol Pharm 16:1220-1233
Almeida FA, Vale EM, Reis RS, Santa-Catarina C, Silveira V (2019) LED lamps enhance somatic embryo maturation in association with the differential accumulation of proteins in the Carica papaya L. 'Golden' embryogenic callus. Plant Physiol Biochem 143:109-118
Al-Mutairy EA, Imtiaz FA, Khalid M, Al Qattan S, Saleh S, Mahmoud LM, Al-Saif MM, et al. (2019) An atypical pulmonary fibrosis is associated with co-inheritance of mutations in the calcium binding protein genes S100A3 and S100A13. Eur Respir J 54 (in press)
Aloui C, Barlier C, Claverol S, Fagan J, Awounou D, Tavernier E, Guyotat D, Hamzeh-Cognasse H, Cognasse F, Garraud O, Laradi S (2019) Differential protein expression of blood platelet components associated with adverse transfusion reactions. J Proteomics 194:25-36
Araujo-Gomes N, Romero-Gavilan F, Zhang Y, Martinez-Ramos C, Elortza F, Azkargorta M, Martin de Llano JJ, Gurruchaga M, Goni I, van den Beucken J, Suay J (2019) Complement proteins regulating macrophage polarisation on biomaterials. Colloids Surf B Biointerfaces 181:125-133
Arffman RK, Saraswat M, Joenvaara S, Khatun M, Agarwal R, Tohmola T, Sundstrom-Poromaa I, Renkonen R, Piltonen TT (2019) Thromboinflammatory changes in plasma proteome of pregnant women with PCOS detected by quantitative label-free proteomics. Sci Rep 9:17578
Bagi Z, Couch Y, Broskova Z, Perez-Balderas F, Yeo T, Davis S, Fischer R, Sibson NR, Davis BG, Anthony DC (2019) Extracellular vesicle integrins act as a nexus for platelet adhesion in cerebral microvessels. Sci Rep 9:15847
Banfi C, Brioschi M, Karjalainen MK, Huusko JM, Gianazza E, Agostoni P (2019) Immature surfactant protein-B impairs the antioxidant capacity of HDL. Int J Cardiol 285:53-58
Banova Vulic R, Zduriencikova M, Tyciakova S, Benada O, Dubrovcakova M, Lakota J, Skultety L (2019) Silencing of carbonic anhydrase I enhances the malignant potential of exosomes secreted by prostatic tumour cells. J Cell Mol Med 23:3641-3655
Bardell D, Milner PI, Goljanek-Whysall K, Peffers MJ (2019) Differences in plasma and peritoneal fluid proteomes identifies potential biomarkers associated with survival following strangulating small intestinal disease. Equine Vet J 51:727-732
Barjaktarovic Z, Merl-Pham J, Braga-Tanaka I, Tanaka S, Hauck SM, Saran A, Mancuso M, Atkinson MJ, Tapio S, Azimzadeh O (2019) Hyperacetylation of Cardiac Mitochondrial Proteins Is Associated with Metabolic Impairment and Sirtuin Downregulation after Chronic Total Body Irradiation of ApoE (-/-) Mice. Int J Mol Sci 20 (in press)
Baud A, Little D, Wen TQ, Heywood WE, Gissen P, Mills K (2019) An Optimized Method for the Proteomic Analysis of Low Volumes of Cell Culture Media and the Secretome: The Application and the Demonstration of Altered Protein Expression in iPSC-Derived Neuronal Cell Lines from Parkinson's Disease Patients. J Proteome Res 18:1198-1207
Bazile J, Picard B, Chambon C, Valais A, Bonnet M (2019) Pathways and biomarkers of marbling and carcass fat deposition in bovine revealed by a combination of gel-based and gel-free proteomic analyses. Meat Sci 156:146-155
Beker MC, Caglayan B, Caglayan AB, Kelestemur T, Yalcin E, Caglayan A, Kilic U, Baykal AT, Reiter RJ, Kilic E (2019) Interaction of melatonin and Bmal1 in the regulation of PI3K/AKT pathway components and cellular survival. Sci Rep 9:19082
Belchi-Navarro S, Almagro L, Bru-Martinez R, Pedreno MA (2019) Changes in the secretome of Vitis vinifera cv. Monastrell cell cultures treated with cyclodextrins and methyl jasmonate. Plant Physiol Biochem 135:520-527
Bell M, Blais JM (2019) "-Omics" workflow for paleolimnological and geological archives: A review. Sci Total Environ 672:438-455
Berg P, McConnell EW, Hicks LM, Popescu SC, Popescu GV (2019) Evaluation of linear models and missing value imputation for the analysis of peptide-centric proteomics. BMC Bioinformatics 20:102
Beyer JA, Szollossi A, Byles E, Fischer R, Armitage JP (2019) Mechanism of Signalling and Adaptation through the Rhodobacter sphaeroides Cytoplasmic Chemoreceptor Cluster. Int J Mol Sci 20 (in press)
Bharucha T, Gangadharan B, Kumar A, de Lamballerie X, Newton PN, Winterberg M, Dubot-Peres A, Zitzmann N (2019) Mass spectrometry-based proteomic techniques to identify cerebrospinal fluid biomarkers for diagnosing suspected central nervous system infections. A systematic review. J Infect 79:407-418
Bi B, Choi HP, Hyeon SJ, Sun S, Su N, Liu Y, Lee J, Kowall NW, McKee AC, Yang JH, Ryu H (2019) Quantitative Proteomic Analysis Reveals Impaired Axonal Guidance Signaling in Human Postmortem Brain Tissues of Chronic Traumatic Encephalopathy. Exp Neurobiol 28:362-375
Biswal AK, McConnell EW, Werth EG, Lo SF, Yu SM, Hicks LM, Jones AM (2019) The Nucleotide-Dependent Interactome of Rice Heterotrimeric G-Protein alpha -Subunit. Proteomics 19:e1800385
Blank-Landeshammer B, Teichert I, Marker R, Nowrousian M, Kuck U, Sickmann A (2019) Combination of Proteogenomics with Peptide De Novo Sequencing Identifies New Genes and Hidden Posttranscriptional Modifications. MBio 10 (in press)
Boccard J, Tonoli D, Strajhar P, Jeanneret F, Odermatt A, Rudaz S (2019) Removal of batch effects using stratified subsampling of metabolomic data for in vitro endocrine disruptors screening. Talanta 195:77-86
Bonnet J, Garcia C, Leger T, Couquet MP, Vignoles P, Vatunga G, Ndung'u J, Boudot C, Bisser S, Courtioux B (2019) Proteome characterization in various biological fluids of Trypanosoma brucei gambiense-infected subjects. J Proteomics 196:150-161
Bouillier C, Cosentino G, Leger T, Rincheval V, Richard CA, Desquesnes A, Sitterlin D, Blouquit-Laye S, Eleouet JF, Gault E, Rameix-Welti MA (2019) The Interactome analysis of the Respiratory Syncytial Virus protein M2-1 suggests a new role in viral mRNA metabolism post-transcription. Sci Rep 9:15258
Bowazolo C, Tse SPK, Beauchemin M, Lo SC, Rivoal J, Morse D (2019) Label-free MS/MS analyses of the dinoflagellate Lingulodinium identifies rhythmic proteins facilitating adaptation to a diurnal LD cycle. Sci Total Environ:135430
Brandao-Teles C, de Almeida V, Cassoli JS, Martins-de-Souza D (2019) Oligodendrocytes: Potential of Discovering New Treatment Targets. Front Pharmacol 10:186
Brault ML, Petit JD, Immel F, Nicolas WJ, Glavier M, Brocard L, Gaston A, Fouche M, Hawkins TJ, Crowet JM, Grison MS, Germain V, Rocher M, Kraner M, Alva V, Claverol S, Paterlini A, Helariutta Y, Deleu M, Lins L, Tilsner J, Bayer EM (2019) Multiple C2 domains and transmembrane region proteins (MCTPs) tether membranes at plasmodesmata. EMBO Rep 20:e47182
Breton J, Legrand R, Achamrah N, Chan P, do Rego JL, do Rego JC, Coeffier M, Dechelotte P, Fetissov SO (2019) Proteome modifications of gut microbiota in mice with activity-based anorexia and starvation: Role in ATP production. Nutrition 67-68:110557
Brunoro GVF, Carvalho PC, Barbosa VC, Pagnoncelli D, De Moura Gallo CV, Perales J, Zahedi RP, Valente RH, Neves-Ferreira A (2019) Differential proteomic comparison of breast cancer secretome using a quantitative paired analysis workflow. BMC Cancer 19:365
Burlina S, Banfi C, Brioschi M, Visentin S, Dalfra MG, Traldi P, Lapolla A (2019) Is the placental proteome impaired in well-controlled gestational diabetes? J Mass Spectrom 54:359-365
Calderon-Rodriguez SI, Sanabria-Salas MC, Umana-Perez A (2019) A comparative proteomic study of plasma in Colombian childhood acute lymphoblastic leukemia. PLoS One 14:e0221509
Canals R, Hammarlof DL, Kroger C, Owen SV, Fong WY, Lacharme-Lora L, Zhu X, Wenner N, Carden SE, Honeycutt J, Monack DM, Kingsley RA, Brownridge P, Chaudhuri RR, Rowe WPM, Predeus AV, Hokamp K, Gordon MA, Hinton JCD (2019) Adding function to the genome of African Salmonella Typhimurium ST313 strain D23580. PLoS Biol 17:e3000059
Cao TH, Jones DJL, Voors AA, Quinn PA, Sandhu JK, Chan DCS, Parry HM, Mohan M, Mordi IR, Sama IE, Anker SD, Cleland JG, Dickstein K, Filippatos G, Hillege HL, Metra M, Ponikowski P, Samani NJ, Van Veldhuisen DJ, Zannad F, Lang CC, Ng LL (2019) Plasma proteomic approach in patients with heart failure: insights into pathogenesis of disease progression and potential novel treatment targets. Eur J Heart Fail (in press)
Chan BD, Wong WY, Lee MM, Cho WC, Yee BK, Kwan YW, Tai WC (2019) Exosomes in Inflammation and Inflammatory Disease. Proteomics 19:e1800149
Chang YT, Chang MC, Tsai YJ, Ferng C, Shih HC, Kuo YP, Chen CH, Tsai IL (2019) Method development of immunoglobulin G purification from micro-volumes of human serum for untargeted and targeted proteomics-based antibody repertoire studies. J Food Drug Anal 27:475-482
Chapman EA, Lyon M, Simpson D, Mason D, Beynon RJ, Moots RJ, Wright HL (2019) Caught in a Trap? Proteomic Analysis of Neutrophil Extracellular Traps in Rheumatoid Arthritis and Systemic Lupus Erythematosus. Front Immunol 10:423
Chapman TP, Corridoni D, Shiraishi S, Pandey S, Aulicino A, Wigfield S, do Carmo Costa M, Thezenas ML, Paulson H, Fischer R, Kessler BM, Simmons A (2019) Ataxin-3 Links NOD2 and TLR2 Mediated Innate Immune Sensing and Metabolism in Myeloid Cells. Front Immunol 10:1495
Charkoftaki G, Thompson DC, Golla JP, Garcia-Milian R, Lam TT, Engel J, Vasiliou V (2019) Integrated multi-omics approach reveals a role of ALDH1A1 in lipid metabolism in human colon cancer cells. Chem Biol Interact 304:88-96
Chen F, Ma R, Chen XL (2019) Advances of Metabolomics in Fungal Pathogen-Plant Interactions. Metabolites 9
Chen X, Liu H, Sun W, Guo Z, Lang J (2019) Elevated urine histone 4 levels in women with ovarian endometriosis revealed by discovery and parallel reaction monitoring proteomics. J Proteomics 204:103398
Cheng HW, Hsiao CT, Chen YQ, Huang CM, Chan SI, Chiou A, Kuo JC (2019) Centrosome guides spatial activation of Rac to control cell polarization and directed cell migration. Life Sci Alliance 2 (in press)
Cho SY, Protzman RA, Kim YO, Vaidya B, Oh MJ, Kwon J, Kim D (2019) Elucidation of mechanism for host response to VHSV infection at varying temperatures in vitro and in vivo through proteomic analysis. Fish Shellfish Immunol 88:244-253
Coleman O, Suda S, Meiller J, Henry M, Riedl M, Barron N, Clynes M, Meleady P (2019) Increased growth rate and productivity following stable depletion of miR-7 in a mAb producing CHO cell line causes an increase in proteins associated with the Akt pathway and ribosome biogenesis. J Proteomics 195:23-32
Coliva G, Duarte S, Perez-Sala D, Fedorova M (2019) Impact of inhibition of the autophagy-lysosomal pathway on biomolecules carbonylation and proteome regulation in rat cardiac cells. Redox Biol:101123
Corridoni D, Shiraishi S, Chapman T, Steevels T, Muraro D, Thezenas ML, Prota G, Chen JL, Gileadi U, Ternette N, Cerundolo V, Simmons A (2019) NOD2 and TLR2 Signal via TBK1 and PI31 to Direct Cross-Presentation and CD8 T Cell Responses. Front Immunol 10:958
Couto N, Al-Majdoub ZM, Achour B, Wright PC, Rostami-Hodjegan A, Barber J (2019) Quantification of Proteins Involved in Drug Metabolism and Disposition in the Human Liver Using Label-Free Global Proteomics. Mol Pharm 16:632-647
Cury NF, RCC ES, Fayad M, Fontes W, Ricart CAO, Castro MS, Silveira C, Valle de Sousa M, Pereira LAR (2019) Proteome Dataset of Qualea grandiflora Mart. (Vochysiaceae) by LC-MS/MS Label-Free Identification in Response to Aluminum. Proteomics 19:e1900148
De Clerck L, Taelman J, Popovic M, Willems S, Van der Jeught M, Heindryckx B, De Sutter P, Marks H, Deforce D, Dhaenens M (2019) Untargeted histone profiling during naive conversion uncovers conserved modification markers between mouse and human. Sci Rep 9:17240
De Meyer F, Eeckhaut V, Ducatelle R, Dhaenens M, Daled S, Dedeurwaerder A, De Gussem M, Haesebrouck F, Deforce D, Van Immerseel F (2019) Host intestinal biomarker identification in a gut leakage model in broilers. Vet Res 50:46
de Oliveira Gorgulho Silva C, de Castro Moreira Dos Santos Junior A, Santana RH, Kruger RH, Fontes W, de Sousa MV, Ricart CAO, Filho EXF (2019) Mild hydrothermal pretreatment of sugarcane bagasse enhances the production of holocellulases by Aspergillus niger. J Ind Microbiol Biotechnol 46:1517-1529
de Pablos LM, Ferreira TR, Dowle AA, Forrester S, Parry E, Newling K, Walrad PB (2019) The mRNA-bound Proteome of Leishmania mexicana: Novel Genetic Insight into an Ancient Parasite. Mol Cell Proteomics 18:1271-1284
Derricott H, Luu L, Fong WY, Hartley CS, Johnston LJ, Armstrong SD, Randle N, Duckworth CA, Campbell BJ, Wastling JM, Coombes JL (2019) Developing a 3D intestinal epithelium model for livestock species. Cell Tissue Res 375:409-424
Dhaenens L, Lierman S, De Clerck L, Govaert E, Deforce D, Tilleman K, De Sutter P (2019) Endometrial stromal cell proteome mapping in repeated implantation failure and recurrent pregnancy loss cases and fertile women. Reprod Biomed Online 38:442-454
Dietrich J, Roth M, Konig S, Geerling G, Mertsch S, Schrader S (2019) Analysis of lacrimal gland derived mesenchymal stem cell secretome and its impact on epithelial cell survival. Stem Cell Res 38:101477
Dorgau B, Felemban M, Hilgen G, Kiening M, Zerti D, Hunt NC, Doherty M, Whitfield P, Hallam D, White K, Ding Y, Krasnogor N, Al-Aama J, Asfour HZ, Sernagor E, Lako M (2019) Decellularised extracellular matrix-derived peptides from neural retina and retinal pigment epithelium enhance the expression of synaptic markers and light responsiveness of human pluripotent stem cell derived retinal organoids. Biomaterials 199:63-75
Dowling P, Zweyer M, Raucamp M, Henry M, Meleady P, Swandulla D, Ohlendieck K (2019) Proteomic and cell biological profiling of the renal phenotype of the mdx-4cv mouse model of Duchenne muscular dystrophy. Eur J Cell Biol:151059
Duijvesz D, Rodriguez-Blanco G, Hoogland AM, Verhoef EI, Dekker LJ, Roobol MJ, van Leenders G, Luider TM, Jenster G (2019) Differential tissue expression of extracellular vesicle-derived proteins in prostate cancer. Prostate 79:1032-1042
Duval C, Macabiou C, Garcia C, Lesuisse E, Camadro JM, Auchere F (2019) The adaptive response to iron involves changes in energetic strategies in the pathogen Candida albicans. Microbiologyopen:e970
Egloff P, Zimmermann I, Arnold FM, Hutter CAJ, Morger D, Opitz L, Poveda L, Keserue HA, Panse C, Roschitzki B, Seeger MA (2019) Engineered peptide barcodes for in-depth analyses of binding protein libraries. Nat Methods 16:421-428
Einer C, Leitzinger C, Lichtmannegger J, Eberhagen C, Rieder T, Borchard S, Wimmer R, Denk G, Popper B, Neff F, Polishchuk EV, Polishchuk RS, Hauck SM, von Toerne C, Muller JC, Karst U, Baral BS, DiSpirito AA, Kremer AE, Semrau J, Weiss KH, Hohenester S, Zischka H (2019) A High-Calorie Diet Aggravates Mitochondrial Dysfunction and Triggers Severe Liver Damage in Wilson Disease Rats. Cell Mol Gastroenterol Hepatol 7:571-596
El-Khateeb E, Vasilogianni AM, Alrubia S, Al-Majdoub ZM, Couto N, Howard M, Barber J, Rostami-Hodjegan A, Achour B (2019) Quantitative mass spectrometry-based proteomics in the era of model-informed drug development: Applications in translational pharmacology and recommendations for best practice. Pharmacol Ther 203:107397
Engstrom P, Burke TP, Mitchell G, Ingabire N, Mark KG, Golovkine G, Iavarone AT, Rape M, Cox JS, Welch MD (2019) Evasion of autophagy mediated by Rickettsia surface protein OmpB is critical for virulence. Nat Microbiol 4:2538-2551
Eshraghi M, Gombar R, De Repentigny Y, Vacratsis PO, Kothary R (2019) Pathologic Alterations in the Proteome of Synaptosomes from a Mouse Model of Spinal Muscular Atrophy. J Proteome Res 18:3042-3051
Esteves P, Dard L, Brillac A, Hubert C, Sarlak S, Rousseau B, Dumon E, Izotte J, Bonneu M, Lacombe D, Dupuy JW, Amoedo N, Rossignol R (2019) Nuclear control of lung cancer cells migration, invasion and bioenergetics by eukaryotic translation initiation factor 3F. Oncogene (in press)
Fernandez J, Machado AK, Lyonnais S, Chamontin C, Gartner K, Leger T, Henriquet C, Garcia C, Portilho DM, Pugniere M, Chaloin L, Muriaux D, Yamauchi Y, Blaise M, Nisole S, Arhel NJ (2019) Transportin-1 binds to the HIV-1 capsid via a nuclear localization signal and triggers uncoating. Nat Microbiol 4:1840-1850
Fernandez-Boo S, Gervais O, Prado-Alvarez M, Chollet B, Claverol S, Lecadet C, Dubreuil C, Arzul I (2019) Is pallial mucus involved in Ostrea edulis defenses against the parasite Bonamia ostreae? J Invertebr Pathol 169:107259
Fernandez-Tussy P, Fernandez-Ramos D, Lopitz-Otsoa F, Simon J, Barbier-Torres L, Gomez-Santos B, Nunez-Garcia M, et al. (2019) miR-873-5p targets mitochondrial GNMT-Complex II interface contributing to non-alcoholic fatty liver disease. Mol Metab 29:40-54
Fingas F, Volke D, Hassert R, Fornefett J, Funk S, Baums CG, Hoffmann R (2019) Sensitive and immunogen-specific serological detection of Rodentibacter pneumotropicus infections in mice. BMC Microbiol 19:43
Fisher P, Spencer H, Thomas-Oates J, Wood AJ, Ungar D (2019) Modeling Glycan Processing Reveals Golgi-Enzyme Homeostasis upon Trafficking Defects and Cellular Differentiation. Cell Rep 27:1231-1243 e1236
Ford MM, Smythers AL, McConnell EW, Lowery SC, Kolling DRJ, Hicks LM (2019) Inhibition of TOR in Chlamydomonas reinhardtii Leads to Rapid Cysteine Oxidation Reflecting Sustained Physiological Changes. Cells 8
Frischknecht L, Britschgi C, Galliker P, Christinat Y, Vichalkovski A, Gstaiger M, Kovacs WJ, Krek W (2019) BRAF inhibition sensitizes melanoma cells to alpha-amanitin via decreased RNA polymerase II assembly. Sci Rep 9:7779
Gannesen AV, Zdorovenko EL, Botchkova EA, Hardouin J, Massier S, Kopitsyn DS, Gorbachevskii MV, Kadykova AA, Shashkov AS, Zhurina MV, Netrusov AI, Knirel YA, Plakunov VK, Feuilloley MGJ (2019) Composition of the Biofilm Matrix of Cutibacterium acnes Acneic Strain RT5. Front Microbiol 10:1284
Gartner K, Klahn S, Watanabe S, Mikkat S, Scholz I, Hess WR, Hagemann M (2019) Cytosine N4-Methylation via M.Ssp6803II Is Involved in the Regulation of Transcription, Fine- Tuning of DNA Replication and DNA Repair in the Cyanobacterium Synechocystis sp. PCC 6803. Front Microbiol 10:1233
Gentric G, Kieffer Y, Mieulet V, Goundiam O, Bonneau C, Nemati F, Hurbain I, Raposo G, Popova T, Stern MH, Lallemand-Breitenbach V, Muller S, Caneque T, Rodriguez R, Vincent-Salomon A, de The H, Rossignol R, Mechta-Grigoriou F (2019) PML-Regulated Mitochondrial Metabolism Enhances Chemosensitivity in Human Ovarian Cancers. Cell Metab 29:156-173 e110
Ghousein A, Mosca N, Cartier F, Charpentier J, Dupuy JW, Raymond AA, Bioulac-Sage P, Grosset CF (2019) miR-4510 blocks hepatocellular carcinoma development through RAF1 targeting and RAS/RAF/MEK/ERK signalling inactivation. Liver Int 40:240-251
Gilbert HTJ, Mallikarjun V, Dobre O, Jackson MR, Pedley R, Gilmore AP, Richardson SM, Swift J (2019) Nuclear decoupling is part of a rapid protein-level cellular response to high-intensity mechanical loading. Nat Commun 10:4149
Gilgunn S, El-Sabbahy H, Albrecht S, Gaikwad M, Corrigan K, Deakin L, Jellum G, Bones J (2019) Identification and tracking of problematic host cell proteins removed by a synthetic, highly functionalized nonwoven media in downstream bioprocessing of monoclonal antibodies. J Chromatogr A 1595:28-38
Gonzalez-Morales A, Zabaleta A, Garcia-Moure M, Alonso MM, Fernandez-Irigoyen J, Santamaria E (2019) Oncolytic adenovirus Delta-24-RGD induces a widespread glioma proteotype remodeling during autophagy. J Proteomics 194:168-178
Graham LC, Naldrett MJ, Kohama SG, Smith C, Lamont DJ, McColl BW, Gillingwater TH, Skehel P, Urbanski HF, Wishart TM (2019) Regional Molecular Mapping of Primate Synapses during Normal Healthy Aging. Cell Rep 27:1018-1026 e1014
Guo XH, Cheng XL, Liu WX, Li MH, Wei F, Ma SC (2019) Identification of velvet antler and its mixed varieties by UPLC-QTOF-MS combined with principal component analysis. J Pharm Biomed Anal 165:18-23
Gutsch A, Sergeant K, Keunen E, Prinsen E, Guerriero G, Renaut J, Hausman JF, Cuypers A (2019) Does long-term cadmium exposure influence the composition of pectic polysaccharides in the cell wall of Medicago sativa stems? BMC Plant Biol 19:271
Hadjidemetriou M, Al-Ahmady Z, Buggio M, Swift J, Kostarelos K (2019) A novel scavenging tool for cancer biomarker discovery based on the blood-circulating nanoparticle protein corona. Biomaterials 188:118-129
Ham AS, Chojnowska K, Tintignac LA, Lin S, Schmidt A, Ham DJ, Sinnreich M, Ruegg MA (2019) mTORC1 signalling is not essential for the maintenance of muscle mass and function in adult sedentary mice. J Cachexia Sarcopenia Muscle (in press)
Hayashi N, Doi H, Kurata Y, Kagawa H, Atobe Y, Funakoshi K, Tada M, Katsumoto A, Tanaka K, Kunii M, Nakamura H, Takahashi K, Takeuchi H, Koyano S, Kimura Y, Hirano H, Tanaka F (2019) Proteomic analysis of exosome-enriched fractions derived from cerebrospinal fluid of amyotrophic lateral sclerosis patients. Neurosci Res (in press)
Haythorne E, Rohm M, van de Bunt M, Brereton MF, Tarasov AI, Blacker TS, Sachse G, Silva Dos Santos M, Terron Exposito R, Davis S, Baba O, Fischer R, Duchen MR, Rorsman P, MacRae JI, Ashcroft FM (2019) Diabetes causes marked inhibition of mitochondrial metabolism in pancreatic beta-cells. Nat Commun 10:2474
Hiroshima Y, Kasajima R, Kimura Y, Komura D, Ishikawa S, Ichikawa Y, Bouvet M, Yamamoto N, Oshima T, Morinaga S, Singh SR, Hoffman RM, Endo I, Miyagi Y (2019) Novel targets identified by integrated cancer-stromal interactome analysis of pancreatic adenocarcinoma. Cancer Lett 469:217-227
Hladik D, Dalke C, von Toerne C, Hauck SM, Azimzadeh O, Philipp J, Ung MC, Schlattl H, Rossler U, Graw J, Atkinson MJ, Tapio S (2019) CREB Signaling Mediates Dose-Dependent Radiation Response in the Murine Hippocampus Two Years after Total Body Exposure. J Proteome Res (in press)
Hu C, Sun J, Du J, Wen D, Lu H, Zhang H, Xue Y, Zhang A, Yang C, Zeng L, Jiang J (2019) The Hippo-YAP Pathway Regulates the Proliferation of Alveolar Epithelial Progenitors after Acute Lung Injury. Cell Biol Int (in press)
Imrie L, Le Bihan T, O'Toole A, Hickner PV, Dunn WA, Weise B, Rund SSC (2019) Genome annotation improvements from cross-phyla proteogenomics and time-of-day differences in malaria mosquito proteins using untargeted quantitative proteomics. PLoS One 14:e0220225
Irshad Z, Xue M, Ashour A, Larkin JR, Thornalley PJ, Rabbani N (2019) Activation of the unfolded protein response in high glucose treated endothelial cells is mediated by methylglyoxal. Sci Rep 9:7889
Jaleel A, Aneesh Kumar A, Ajith Kumar GS, Surendran A, Kartha CC (2019) Label-free quantitative proteomics analysis reveals distinct molecular characteristics in endocardial endothelium. Mol Cell Biochem 451:1-10
James KL, Kung JW, Crable BR, Mouttaki H, Sieber JR, Nguyen HH, Yang Y, Xie Y, Erde J, Wofford NQ, Karr EA, Loo JA, Ogorzalek Loo RR, Gunsalus RP, McInerney MJ (2019) Syntrophus aciditrophicus uses the same enzymes in a reversible manner to degrade and synthesize aromatic and alicyclic acids. Environ Microbiol 21:1833-1846
Jing J, Du Z, Ji S, Han K (2019) Urinary proteome analysis of acute hypercoagulable state in rat model induced by epsilon-aminocaproic acid. Biomed Pharmacother 110:275-284
Jourdan J, Walz A, Matile H, Schmidt A, Wu J, Wang X, Dong Y, Vennerstrom JL, Schmidt RS, Wittlin S, Maser P (2019) Stochastic Protein Alkylation by Antimalarial Peroxides. ACS Infect Dis 5:2067-2075
Jung MH, Chico V, Ciordia S, Mena MC, Jung SJ, Ortega-Villaizan MDM (2019) The Megalocytivirus RBIV Induces Apoptosis and MHC Class I Presentation in Rock Bream (Oplegnathus fasciatus) Red Blood Cells. Front Immunol 10:160
Kadri S, El Ayed M, Cosette P, Jouenne T, Elkhaoui S, Zekri S, Limam F, Aouani E, Mokni M (2019) Neuroprotective effect of grape seed extract on brain ischemia: a proteomic approach. Metab Brain Dis 34:889-907
Kay LJ, Sangal V, Black GW, Soundararajan M (2019) Proteomics and bioinformatics analyses identify novel cellular roles outside mitochondrial function for human miro GTPases. Mol Cell Biochem 451:21-35
Kelly PS, Dorival-Garcia N, Pare S, Carillo S, Ta C, Alarcon Miguez A, Coleman O, Harper E, Shannon M, Henry M, Connolly L, Clynes M, Meleady P, Bones J, Barron N (2019) Improvements in single-use bioreactor film material composition leads to robust and reliable Chinese hamster ovary cell performance. Biotechnol Prog:e2824
Kleinwort KJH, Hauck SM, Degroote RL, Scholz AM, Holzel C, Maertlbauer EP, Deeg C (2019) Peripheral blood bovine lymphocytes and MAP show distinctly different proteome changes and immune pathways in host-pathogen interaction. PeerJ 7:e8130
Koelschbach JS, Mouttaki H, Merl-Pham J, Arnold ME, Meckenstock RU (2019) Identification of naphthalene carboxylase subunits of the sulfate-reducing culture N47. Biodegradation 30:147-160
Konig S, Hadrian K, Schlatt S, Wistuba J, Thanos S, Bohm MRR (2019) Topographic protein profiling of the age-related proteome in the retinal pigment epithelium of Callithrix jacchus with respect to macular degeneration. J Proteomics 191:1-15
Kundu K, Marozava S, Ehrl B, Merl-Pham J, Griebler C, Elsner M (2019) Defining lower limits of biodegradation: atrazine degradation regulated by mass transfer and maintenance demand in Arthrobacter aurescens TC1. Isme J 13:2236-2251
Lampaki D, Diepold A, Glatter T (2019) A Serial Sample Processing Strategy with Improved Performance for in-Depth Quantitative Analysis of Type III Secretion Events in Pseudomonas aeruginosa. J Proteome Res (in press)
Lecourieux D, Kappel C, Claverol S, Pieri P, Feil R, Lunn JE, Bonneu M, Wang L, Gomes E, Delrot S, Lecourieux F (2019) Proteomic and metabolomic profiling underlines the stage- and time-dependent effects of high temperature on grape berry metabolism. J Integr Plant Biol (in press)
Lelandais G, Denecker T, Garcia C, Danila N, Leger T, Camadro JM (2019) Label-free quantitative proteomics in Candida yeast species: technical and biological replicates to assess data reproducibility. BMC Res Notes 12:470
Ley D, Pereira S, Pedersen LE, Arnsdorf J, Hefzi H, Davy AM, Ha TK, Wulff T, Kildegaard HF, Andersen MR (2019) Reprogramming AA catabolism in CHO cells with CRISPR/Cas9 genome editing improves cell growth and reduces by product secretion. Metab Eng 56:120-129
Li B, Takahashi D, Kawamura Y, Uemura M (2019) Plasma membrane proteome analyses of Arabidopsis thaliana suspension-cultured cells during cold or ABA treatment: Relationship with freezing tolerance and growth phase. J Proteomics 211:103528
Li L, Zhao Z, Jiang W, Guo J, Zhang S (2019) Identification and functional characterization of Lys-trimethylation of lactate dehydrogenase A. Onco Targets Ther 12:5395-5404
Liljedahl L, Pedersen MH, McGuire JN, James P (2019) The impact of the glucagon-like peptide 1 receptor agonist liraglutide on the streptozotocin-induced diabetic mouse kidney proteome. Physiol Rep 7:e13994
Lisitsa AV, Petushkova NA, Levitsky LI, Zgoda VG, Larina OV, Kisrieva YS, Frankevich VE, Gamidov SI (2019) [Comparative Analysis of the Performance of Mascot and IdentiPy Algorithms on a Benchmark Dataset Obtained by Tandem Mass Spectrometry Analysis of Testicular Biopsies]. Mol Biol (Mosk) 53:166-176
Liu S, Liu X, Wu F, Zhang X, Zhang H, Gao D, Bi D, Qu H, Ge J, Xu Y, Zhao Z (2019) HADHA overexpression disrupts lipid metabolism and inhibits tumor growth in clear cell renal cell carcinoma. Exp Cell Res 384:111558
Liu X, Song Y, Guo Z, Sun W, Liu J (2019) A comprehensive profile and inter-individual variations analysis of the human normal amniotic fluid proteome. J Proteomics 192:1-9
Lo M, Zhu J, Hansen SG, Carroll T, Farr Zuend C, Noel-Romas L, Ma ZM, Fritts L, Huang ML, Sun S, Huang Y, Koelle DM, Picker LJ, Burgener A, Corey L, Miller CJ (2019) Acute Infection and Subsequent Subclinical Reactivation of Herpes Simplex Virus 2 after Vaginal Inoculation of Rhesus Macaques. J Virol 93
Lopes EM, Linhares RG, de Oliveira Pires L, Castro RN, Souza G, Koblitz MGB, Cameron LC, Macedo AF (2019) Vanilla bahiana, a contribution from the Atlantic Forest biodiversity for the production of vanilla: A proteomic approach through high-definition nanoLC/MS. Food Res Int 120:148-156
Luo M, Willis WT, Coletta DK, Langlais PR, Mengos A, Ma W, Finlayson J, Wagner GR, Shi CX, Mandarino LJ (2019) Deletion of the Mitochondrial Protein VWA8 Induces Oxidative Stress and an HNF4alpha Compensatory Response in Hepatocytes. Biochemistry 58:4983-4996
Luther A, Urfer M, Zahn M, Muller M, Wang SY, Mondal M, Vitale A, et al. (2019) Chimeric peptidomimetic antibiotics against Gram-negative bacteria. Nature 576:452-458
Luu L, Johnston LJ, Derricott H, Armstrong SD, Randle N, Hartley CS, Duckworth CA, Campbell BJ, Wastling JM, Coombes JL (2019) An Open-Format Enteroid Culture System for Interrogation of Interactions Between Toxoplasma gondii and the Intestinal Epithelium. Front Cell Infect Microbiol 9:300
Mages K, Grassmann F, Jagle H, Rupprecht R, Weber BHF, Hauck SM, Grosche A (2019) The agonistic TSPO ligand XBD173 attenuates the glial response thereby protecting inner retinal neurons in a murine model of retinal ischemia. J Neuroinflammation 16:43
Martinez Fernandez A, Regazzoni L, Brioschi M, Gianazza E, Agostoni P, Aldini G, Banfi C (2019) Pro-oxidant and pro-inflammatory effects of glycated albumin on cardiomyocytes. Free Radic Biol Med 144:245-255
Martinez-Herrero MDC, Garijo-Toledo MM, Gonzalez F, Bilic I, Liebhart D, Ganas P, Hess M, Gomez-Munoz MT (2019) Membrane associated proteins of two Trichomonas gallinae clones vary with the virulence. PLoS One 14:e0224032
Miki Y, Takahashi D, Kawamura Y, Uemura M (2019) Temporal proteomics of Arabidopsis plasma membrane during cold- and de-acclimation. J Proteomics 197:71-81
Milkovska-Stamenova S, Hoffmann R (2019) Diversity of advanced glycation end products in the bovine milk proteome. Amino Acids 51:891-901
Modic M, Grosch M, Rot G, Schirge S, Lepko T, Yamazaki T, Lee FCY, Rusha E, Shaposhnikov D, Palo M, Merl-Pham J, Cacchiarelli D, Rogelj B, Hauck SM, von Mering C, Meissner A, Lickert H, Hirose T, Ule J, Drukker M (2019) Cross-Regulation between TDP-43 and Paraspeckles Promotes Pluripotency-Differentiation Transition. Mol Cell 74:951-965 e913
Mogielnicka-Brzozowska M, Prochowska S, Nizanski W, Bromke MA, Wisniewski J, Olejnik B, Kuzborska A, Fraser L, Mlynarz P, Kordan W (2019) Proteome of cat semen obtained after urethral catheterization. Theriogenology 141:68-81
Moro A, Driscoll TP, Boraas LC, Armero W, Kasper DM, Baeyens N, Jouy C, Mallikarjun V, Swift J, Ahn SJ, Lee D, Zhang J, Gu M, Gerstein M, Schwartz M, Nicoli S (2019) MicroRNA-dependent regulation of biomechanical genes establishes tissue stiffness homeostasis. Nat Cell Biol 21:348-358
Moyer TB, Heil LR, Kirkpatrick CL, Goldfarb D, Lefever WA, Parsley NC, Wommack AJ, Hicks LM (2019) PepSAVI-MS Reveals a Proline-rich Antimicrobial Peptide in Amaranthus tricolor. J Nat Prod 82:2744-2753
Murphy S, Zweyer M, Henry M, Meleady P, Mundegar RR, Swandulla D, Ohlendieck K (2019) Proteomic analysis of the sarcolemma-enriched fraction from dystrophic mdx-4cv skeletal muscle. J Proteomics 191:212-227
Murphy S, Zweyer M, Raucamp M, Henry M, Meleady P, Swandulla D, Ohlendieck K (2019) Proteomic profiling of the mouse diaphragm and refined mass spectrometric analysis of the dystrophic phenotype. J Muscle Res Cell Motil 40:9-28
Nazar RN, Castroverde CDM, Xu X, Kurosky A, Robb J (2019) Wounding induces tomato Ve1 R-gene expression. Planta 249:1779-1797
Nie Q, Xing M, Chen H, Hu J, Nie S (2019) Metabolomics and Lipidomics Profiling Reveals Hypocholesterolemic and Hypolipidemic Effects of Arabinoxylan on Type 2 Diabetic Rats. J Agric Food Chem 67:10614-10623
Novais FJ, Pires PRL, Alexandre PA, Dromms RA, Iglesias AH, Ferraz JBS, Styczynski MP, Fukumasu H (2019) Identification of a metabolomic signature associated with feed efficiency in beef cattle. BMC Genomics 20:8
Nowill AE, Fornazin MC, Spago MC, Dorgan Neto V, Pinheiro VRP, Alexandre SSS, Moraes EO, Souza G, Eberlin MN, Marques LA, Meurer EC, Franchi GC, Jr., de Campos-Lima PO (2019) Immune Response Resetting in Ongoing Sepsis. J Immunol 203:1298-1312
Nybo T, Davies MJ, Rogowska-Wrzesinska A (2019) Analysis of protein chlorination by mass spectrometry. Redox Biol 26:101236
Obradovic MMS, Hamelin B, Manevski N, Couto JP, Sethi A, Coissieux MM, Munst S, Okamoto R, Kohler H, Schmidt A, Bentires-Alj M (2019) Glucocorticoids promote breast cancer metastasis. Nature 567:540-544
Ochsner AM, Hemmerle L, Vonderach T, Nussli R, Bortfeld-Miller M, Hattendorf B, Vorholt JA (2019) Use of rare-earth elements in the phyllosphere colonizer Methylobacterium extorquens PA1. Mol Microbiol 111:1152-1166
Oliveira PVS, Garcia-Rosa S, Sachetto ATA, Moretti AIS, Debbas V, De Bessa TC, Silva NT, Pereira ADC, Martins-de-Souza D, Santoro ML, Laurindo FRM (2019) Protein disulfide isomerase plasma levels in healthy humans reveal proteomic signatures involved in contrasting endothelial phenotypes. Redox Biol 22:101142
Othman Z, Fernandes H, Groot AJ, Luider TM, Alcinesio A, Pereira DM, Guttenplan APM, Yuan H, Habibovic P (2019) The role of ENPP1/PC-1 in osteoinduction by calcium phosphate ceramics. Biomaterials 210:12-24
Padilla CA, Barcena JA, Lopez-Grueso MJ, Requejo-Aguilar R (2019) The regulation of TORC1 pathway by the yeast chaperones Hsp31 is mediated by SFP1 and affects proteasomal activity. Biochim Biophys Acta Gen Subj 1863:534-546
Pallett R, Leslie LJ, Lambert PA, Milic I, Devitt A, Marshall LJ (2019) Anaerobiosis influences virulence properties of Pseudomonas aeruginosa cystic fibrosis isolates and the interaction with Staphylococcus aureus. Sci Rep 9:6748
Pandey R, Zhou M, Islam S, Chen B, Barker NK, Langlais P, Srivastava A, Luo M, Cooke LS, Weterings E, Mahadevan D (2019) Carcinoembryonic antigen cell adhesion molecule 6 (CEACAM6) in Pancreatic Ductal Adenocarcinoma (PDA): An integrative analysis of a novel therapeutic target. Sci Rep 9:18347
Parks C, Giorgianni F, Jones BC, Beranova-Giorgianni S, Moore Ii BM, Mulligan MK (2019) Comparison and Functional Genetic Analysis of Striatal Protein Expression Among Diverse Inbred Mouse Strains. Front Mol Neurosci 12:128
Pauly D, Agarwal D, Dana N, Schafer N, Biber J, Wunderlich KA, Jabri Y, Straub T, Zhang NR, Gautam AK, Weber BHF, Hauck SM, Kim M, Curcio CA, Stambolian D, Li M, Grosche A (2019) Cell-Type-Specific Complement Expression in the Healthy and Diseased Retina. Cell Rep 29:2835-2848 e2834
Paunas FTI, Finne K, Leh S, Osman TA, Marti HP, Berven F, Vikse BE (2019) Characterization of glomerular extracellular matrix in IgA nephropathy by proteomic analysis of laser-captured microdissected glomeruli. BMC Nephrol 20:410
Pawlowski K, Pires JAA, Faulconnier Y, Chambon C, Germon P, Boby C, Leroux C (2019) Mammary Gland Transcriptome and Proteome Modifications by Nutrient Restriction in Early Lactation Holstein Cows Challenged with Intra-Mammary Lipopolysaccharide. Int J Mol Sci 20
Personnic N, Striednig B, Lezan E, Manske C, Welin A, Schmidt A, Hilbi H (2019) Quorum sensing modulates the formation of virulent Legionella persisters within infected cells. Nat Commun 10:5216
Phan V, Cox D, Cipriani S, Spendiff S, Buchkremer S, O'Connor E, Horvath R, Goebel HH, Hathazi D, Lochmuller H, Straka T, Rudolf R, Weis J, Roos A (2019) SIL1 deficiency causes degenerative changes of peripheral nerves and neuromuscular junctions in fish, mice and human. Neurobiol Dis 124:218-229
Piasecka A, Kachlicki P, Stobiecki M (2019) Analytical Methods for Detection of Plant Metabolomes Changes in Response to Biotic and Abiotic Stresses. Int J Mol Sci 20
Prasad B, Achour B, Artursson P, Hop C, Lai Y, Smith PC, Barber J, Wisniewski JR, Spellman D, Uchida Y, Zientek MA, Unadkat JD, Rostami-Hodjegan A (2019) Toward a Consensus on Applying Quantitative Liquid Chromatography-Tandem Mass Spectrometry Proteomics in Translational Pharmacology Research: A White Paper. Clin Pharmacol Ther 106:525-543
Price MN, Ray J, Iavarone AT, Carlson HK, Ryan EM, Malmstrom RR, Arkin AP, Deutschbauer AM (2019) Oxidative Pathways of Deoxyribose and Deoxyribonate Catabolism. mSystems 4
Prieto-Bonete G, Perez-Carceles MD, Maurandi-Lopez A, Perez-Martinez C, Luna A (2019) Association between protein profile and postmortem interval in human bone remains. J Proteomics 192:54-63
Prokopec SD, Lu A, Lee SCS, Yao CQ, Sun RX, Watson JD, Soliymani R, de Borja R, Wong A, Sam M, Zuzarte P, McPherson JD, Okey AB, Pohjanvirta R, Boutros PC (2019) Comparative toxicoproteogenomics of mouse and rat liver identifies TCDD-resistance genes. Arch Toxicol 93:2961-2978
Prozeller D, Pereira J, Simon J, Mailander V, Morsbach S, Landfester K (2019) Prevention of Dominant IgG Adsorption on Nanocarriers in IgG-Enriched Blood Plasma by Clusterin Precoating. Adv Sci (Weinh) 6:1802199
Qin W, Li L, Wang T, Huang H, Gao Y (2019) Urine Proteome Changes in a TNBS-Induced Colitis Rat Model. Proteomics Clin Appl:e1800100
Raasakka A, Linxweiler H, Brophy PJ, Sherman DL, Kursula P (2019) Direct Binding of the Flexible C-Terminal Segment of Periaxin to beta4 Integrin Suggests a Molecular Basis for CMT4F. Front Mol Neurosci 12:84
Rahm M, Merl-Pham J, Adamski J, Hauck SM (2019) Time-resolved phosphoproteomic analysis elucidates hepatic 11,12-Epoxyeicosatrienoic acid signaling pathways. Prostaglandins Other Lipid Mediat 146:106387
Reese HR, Shanahan CC, Proulx C, Menegatti S (2019) Peptide science: A "rule model" for new generations of peptidomimetics. Acta Biomater (in press)
Riba A, Di Nanni N, Mittal N, Arhne E, Schmidt A, Zavolan M (2019) Protein synthesis rates and ribosome occupancies reveal determinants of translation elongation rates. Proc Natl Acad Sci U S A 116:15023-15032
Ribeiro DG, de Almeida RF, Fontes W, de Souza Castro M, de Sousa MV, Ricart CAO, da Cunha RNV, Lopes R, Scherwinski-Pereira JE, Mehta A (2019) Stress and cell cycle regulation during somatic embryogenesis plays a key role in oil palm callus development. J Proteomics 192:137-146
Ribeiro DM, Planchon S, Leclercq CC, Raundrup K, Alves SP, Bessa RJB, Renaut J, Almeida AM (2019) The muscular, hepatic and adipose tissues proteomes in muskox (Ovibos moschatus): Differences between males and females. J Proteomics 208:103480
Rodrigues JM, Lasa B, Betti M, Fernandez-Irigoyen J, Santamaria E, Gonzalez-Murua C, Aparicio-Tejo PM, Marino D (2019) Multi-omic and physiologic approach to understand Lotus japonicus response upon exposure to 3,4 dimethylpyrazole phosphate nitrification inhibitor. Sci Total Environ 660:1201-1209
Romero-Gavilan F, Araujo-Gomes N, Cerqueira A, Garcia-Arnaez I, Martinez-Ramos C, Azkargorta M, Iloro I, Elortza F, Gurruchaga M, Suay J, Goni I (2019) Proteomic analysis of calcium-enriched sol-gel biomaterials. J Biol Inorg Chem 24:563-574
Romero-Gavilan F, Araujo-Gomes N, Garcia-Arnaez I, Martinez-Ramos C, Elortza F, Azkargorta M, Iloro I, Gurruchaga M, Suay J, Goni I (2019) The effect of strontium incorporation into sol-gel biomaterials on their protein adsorption and cell interactions. Colloids Surf B Biointerfaces 174:9-16
Rosendale M, Van TNN, Grillo-Bosch D, Sposini S, Claverie L, Gauthereau I, Claverol S, Choquet D, Sainlos M, Perrais D (2019) Functional recruitment of dynamin requires multimeric interactions for efficient endocytosis. Nat Commun 10:4462
Sahebekhtiari N, Saraswat M, Joenvaara S, Jokinen R, Lovric A, Kaye S, Mardinoglu A, Rissanen A, Kaprio J, Renkonen R, Pietilainen KH (2019) Plasma Proteomics Analysis Reveals Dysregulation of Complement Proteins and Inflammation in Acquired Obesity-A Study on Rare BMI-Discordant Monozygotic Twin Pairs. Proteomics Clin Appl:e1800173
Salleh MZ, Karuppiah V, Snee M, Thistlethwaite A, Levy CW, Knight D, Derrick JP (2019) Structure and Properties of a Natural Competence-Associated Pilin Suggest a Unique Pilus Tip-Associated DNA Receptor. MBio 10 (in press)
Sander T, Farke N, Diehl C, Kuntz M, Glatter T, Link H (2019) Allosteric Feedback Inhibition Enables Robust Amino Acid Biosynthesis in E. coli by Enforcing Enzyme Overabundance. Cell Syst 8:66-75 e68
Sander T, Wang CY, Glatter T, Link H (2019) CRISPRi-Based Downregulation of Transcriptional Feedback Improves Growth and Metabolism of Arginine Overproducing E. coli. ACS Synth Biol 8:1983-1990
Santos C, Nogueira FCS, Domont GB, Fontes W, Prado GS, Habibi P, Santos VO, Oliveira-Neto OB, Grossi-de-Sa MF, Jorrin-Novo JV, Franco OL, Mehta A (2019) Proteomic Analysis and Functional Validation of a Brassica oleracea Endochitinase Involved in Resistance to Xanthomonas campestris. Front Plant Sci 10:414
Santos Rezende J, Zivanovic M, Costa de Novaes MI, Chen ZY (2019) The AVR4 effector is involved in cercosporin biosynthesis and likely affects the virulence of Cercospora cf. flagellaris on soybean. Mol Plant Pathol 21:53-65
Schada von Borzyskowski L, Severi F, Kruger K, Hermann L, Gilardet A, Sippel F, Pommerenke B, Claus P, Cortina NS, Glatter T, Zauner S, Zarzycki J, Fuchs BM, Bremer E, Maier UG, Amann RI, Erb TJ (2019) Marine Proteobacteria metabolize glycolate via the beta-hydroxyaspartate cycle. Nature 575:500-504
Schelstraete W, Clerck L, Govaert E, Millecam J, Devreese M, Deforce D, Bocxlaer JV, Croubels S (2019) Characterization of Porcine Hepatic and Intestinal Drug Metabolizing CYP450: Comparison with Human Orthologues from A Quantitative, Activity and Selectivity Perspective. Sci Rep 9:9233
Schmal Z, Isermann A, Hladik D, von Toerne C, Tapio S, Rube CE (2019) DNA damage accumulation during fractionated low-dose radiation compromises hippocampal neurogenesis. Radiother Oncol 137:45-54
Semis M, Gugiu GB, Bernstein EA, Bernstein KE, Kalkum M (2019) The Plethora of Angiotensin-Converting Enzyme-Processed Peptides in Mouse Plasma. Anal Chem 91:6440-6453
Sepil I, Hopkins BR, Dean R, Thezenas ML, Charles PD, Konietzny R, Fischer R, Kessler BM, Wigby S (2019) Quantitative Proteomics Identification of Seminal Fluid Proteins in Male Drosophila melanogaster. Mol Cell Proteomics 18:S46-S58
Shah AD, Goode RJA, Huang C, Powell DR, Schittenhelm RB (2019) LFQ-Analyst: An Easy-To-Use Interactive Web Platform To Analyze and Visualize Label-Free Proteomics Data Preprocessed with MaxQuant. J Proteome Res (in press)
Sheng Y, Capri J, Waring A, Valentine JS, Whitelegge J (2019) Exposure of Solvent-Inaccessible Regions in the Amyloidogenic Protein Human SOD1 Determined by Hydroxyl Radical Footprinting. J Am Soc Mass Spectrom 30:218-226
Sibai M, Altuntas E, Suzek BE, Sahin B, Parlayan C, Ozturk G, Baykal AT, Demircan T (2019) Comparison of protein expression profile of limb regeneration between neotenic and metamorphic axolotl. Biochem Biophys Res Commun (in press)
Sobczak M, Pitt AR, Spickett CM, Robaszkiewicz A (2019) PARP1 Co-Regulates EP300-BRG1-Dependent Transcription of Genes Involved in Breast Cancer Cell Proliferation and DNA Repair. Cancers (Basel) 11 (in press)
Soste M, Charmpi K, Lampert F, Gerez JA, van Oostrum M, Malinovska L, Boersema PJ, Prymaczok NC, Riek R, Peter M, Vanni S, Beyer A, Picotti P (2019) Proteomics-Based Monitoring of Pathway Activity Reveals that Blocking Diacylglycerol Biosynthesis Rescues from Alpha-Synuclein Toxicity. Cell Syst 9:309-320 e308
Srisawat K, Hesketh K, Cocks M, Strauss J, Edwards BJ, Lisboa PJ, Shepherd S, Burniston JG (2019) Reliability of Protein Abundance and Synthesis Measurements in Human Skeletal Muscle. Proteomics:e1900194
Stella M, Chinello C, Cazzaniga A, Smith A, Galli M, Piga I, Grasso A, Grasso M, Del Puppo M, Varallo M, Bovo G, Magni F (2019) Histology-guided proteomic analysis to investigate the molecular profiles of clear cell Renal Cell Carcinoma grades. J Proteomics 191:38-47
Sumrall ET, Shen Y, Keller AP, Rismondo J, Pavlou M, Eugster MR, Boulos S, Disson O, Thouvenot P, Kilcher S, Wollscheid B, Cabanes D, Lecuit M, Grundling A, Loessner MJ (2019) Phage resistance at the cost of virulence: Listeria monocytogenes serovar 4b requires galactosylated teichoic acids for InlB-mediated invasion. PLoS Pathog 15:e1008032
Taheri N, Fallman M, Wai SN, Fahlgren A (2019) Accumulation of virulence-associated proteins in Campylobacter jejuni Outer Membrane Vesicles at human body temperature. J Proteomics 195:33-40
Tahrioui A, Duchesne R, Bouffartigues E, Rodrigues S, Maillot O, Tortuel D, Hardouin J, Taupin L, Groleau MC, Dufour A, Deziel E, Brenner-Weiss G, Feuilloley M, Orange N, Lesouhaitier O, Cornelis P, Chevalier S (2019) Extracellular DNA release, quorum sensing, and PrrF1/F2 small RNAs are key players in Pseudomonas aeruginosa tobramycin-enhanced biofilm formation. NPJ Biofilms Microbiomes 5:15
Takahashi D, Gorka M, Erban A, Graf A, Kopka J, Zuther E, Hincha DK (2019) Both cold and sub-zero acclimation induce cell wall modification and changes in the extracellular proteome in Arabidopsis thaliana. Sci Rep 9:2289
Tang X, Li J, Zhao WG, Sun H, Guo Z, Jing L, She Z, Yuan T, Liu SN, Liu Q, Fu Y, Sun W (2019) Comprehensive map and functional annotation of the mouse white adipose tissue proteome. PeerJ 7:e7352
Tavares GC, Pereira FL, Barony GM, Rezende CP, da Silva WM, de Souza G, Verano-Braga T, de Carvalho Azevedo VA, Leal CAG, Figueiredo HCP (2019) Delineation of the pan-proteome of fish-pathogenic Streptococcus agalactiae strains using a label-free shotgun approach. BMC Genomics 20:11
Thoms F, Jennings GT, Maudrich M, Vogel M, Haas S, Zeltins A, Hofmann-Lehmann R, Riond B, Grossmann J, Hunziker P, Fettelschoss-Gabriel A, Senti G, Kundig TM, Bachmann MF (2019) Immunization of cats to induce neutralizing antibodies against Fel d 1, the major feline allergen in human subjects. J Allergy Clin Immunol 144:193-203
Tomlinson L, Lu ZQ, Bentley RA, Colley HE, Murdoch C, Webb SD, Cross MJ, Copple IM, Sharma P (2019) Attenuation of doxorubicin-induced cardiotoxicity in a human in vitro cardiac model by the induction of the NRF-2 pathway. Biomed Pharmacother 112:108637
Tse K, Hammond D, Simpson D, Beynon RJ, Beamer E, Tymianski M, Salter MW, Sills GJ, Thippeswamy T (2019) The impact of postsynaptic density 95 blocking peptide (Tat-NR2B9c) and an iNOS inhibitor (1400W) on proteomic profile of the hippocampus in C57BL/6J mouse model of kainate-induced epileptogenesis. J Neurosci Res 97:1378-1392
Uhrig RG, Schlapfer P, Roschitzki B, Hirsch-Hoffmann M, Gruissem W (2019) Diurnal changes in concerted plant protein phosphorylation and acetylation in Arabidopsis organs and seedlings. Plant J 99:176-194
Unsworth AJ, Bombik I, Pinto-Fernandez A, McGouran JF, Konietzny R, Zahedi RP, Watson SP, Kessler BM, Pears CJ (2019) Human Platelet Protein Ubiquitylation and Changes following GPVI Activation. Thromb Haemost 119:104-116
Uppal S, Liu T, Poliakov E, Gentleman S, Redmond TM (2019) The dual roles of RPE65 S-palmitoylation in membrane association and visual cycle function. Sci Rep 9:5218
Vadgama N, Lamont D, Hardy J, Nasir J, Lovering RC (2019) Distinct proteomic profiles in monozygotic twins discordant for ischaemic stroke. Mol Cell Biochem 456:157-165
van der Ende EL, Meeter LH, Stingl C, van Rooij JGJ, Stoop MP, Nijholt DAT, Sanchez-Valle R, Graff C, Oijerstedt L, Grossman M, McMillan C, Pijnenburg YAL, Laforce R, Jr., Binetti G, Benussi L, Ghidoni R, Luider TM, Seelaar H, van Swieten JC (2019) Novel CSF biomarkers in genetic frontotemporal dementia identified by proteomics. Ann Clin Transl Neurol 6:698-707
van Huizen NA, van Rosmalen J, Dekker LJM, Coebergh van den Braak RRJ, Verhoef C, JNM IJ, Luider TM (2019) Identification of a Collagen Marker in Urine Improves the Detection of Colorectal Liver Metastases. J Proteome Res (in press)
Vileigas DF, Harman VM, Freire PP, Marciano CLC, Sant'Ana PG, de Souza SLB, Mota GAF, da Silva VL, Campos DHS, Padovani CR, Okoshi K, Beynon RJ, Santos LD, Cicogna AC (2019) Landscape of heart proteome changes in a diet-induced obesity model. Sci Rep 9:18050
von Ruden EL, Zellinger C, Gedon J, Walker A, Bierling V, Deeg CA, Hauck SM, Potschka H (2019) Regulation of Alzheimer's disease-associated proteins during epileptogenesis. Neuroscience 424:102-120
Walter J, Huwiler F, Fortes C, Grossmann J, Roschitzki B, Hu J, Naegeli H, Laczko E, Bleul U (2019) Analysis of the equine "cumulome" reveals major metabolic aberrations after maturation in vitro. BMC Genomics 20:588
Waterworth WM, Wilson M, Wang D, Nuhse T, Warward S, Selley J, West CE (2019) Phosphoproteomic analysis reveals plant DNA damage signalling pathways with a functional role for histone H2AX phosphorylation in plant growth under genotoxic stress. Plant J 100:1007-1021
Weeraphan C, Phongdara A, Chaiyawat P, Diskul-Na-Ayudthaya P, Chokchaichamnankit D, Verathamjamras C, Netsirisawan P, Yingchutrakul Y, Roytrakul S, Champattanachai V, Svasti J, Srisomsap C (2019) Phosphoproteome Profiling of Isogenic Cancer Cell-Derived Exosome Reveals HSP90 as a Potential Marker for Human Cholangiocarcinoma. Proteomics 19:e1800159
Wei J, Ni N, Meng W, Gao Y (2019) Early urine proteome changes in the Walker-256 tail-vein injection rat model. Sci Rep 9:13804
Weinstabl H, Treu M, Rinnenthal J, Zahn SK, Ettmayer P, Bader G, Dahmann G, et al. (2019) Intracellular Trapping of the Selective Phosphoglycerate Dehydrogenase (PHGDH) Inhibitor BI-4924 Disrupts Serine Biosynthesis. J Med Chem 62:7976-7997
Wen M, Jin Y, Zhang H, Sun X, Kuai Y, Tan W (2019) Proteomic Analysis of Rat Cerebral Cortex in the Subacute to Long-Term Phases of Focal Cerebral Ischemia-Reperfusion Injury. J Proteome Res 18:3099-3118
Witzke KE, Grosserueschkamp F, Jutte H, Horn M, Roghmann F, von Landenberg N, Bracht T, Kallenbach-Thieltges A, Kafferlein H, Bruning T, Schork K, Eisenacher M, Marcus K, Noldus J, Tannapfel A, Sitek B, Gerwert K (2019) Integrated Fourier Transform Infrared Imaging and Proteomics for Identification of a Candidate Histochemical Biomarker in Bladder Cancer. Am J Pathol 189:619-631
Wu AY, Ueda K, Lai CP (2019) Proteomic Analysis of Extracellular Vesicles for Cancer Diagnostics. Proteomics 19:e1800162
Wu CY, Hsu WL, Tsai MH, Chai CY, Yen CJ, Chen CH, Lu JH, Yu HS, Yoshioka T (2019) A potential new approach for treating systemic sclerosis: Dedifferentiation of SSc fibroblasts and change in the microenvironment by blocking store-operated Ca2+ entry. PLoS One 14:e0213400
Xue J, Scotti E, Stoffel M (2019) CDK8 Regulates Insulin Secretion and Mediates Postnatal and Stress-Induced Expression of Neuropeptides in Pancreatic beta Cells. Cell Rep 28:2892-2904 e2897
Yalcin E, Beker MC, Turkseven S, Caglayan B, Gurel B, Kilic U, Caglayan AB, Kalkan R, Baykal AT, Kelestemur T, Kilic E (2019) Evidence that melatonin downregulates Nedd4-1 E3 ligase and its role in cellular survival. Toxicol Appl Pharmacol 379:114686
Yang SK, Yusoff K, Ajat M, Thomas W, Abushelaibi A, Akseer R, Lim SE, Lai KS (2019) Disruption of KPC-producing Klebsiella pneumoniae membrane via induction of oxidative stress by cinnamon bark (Cinnamomum verum J. Presl) essential oil. PLoS One 14:e0214326
Yli-Karjanmaa M, Clausen BH, Degn M, Novrup HG, Ellman DG, Toft-Jensen P, Szymkowski DE, Stensballe A, Meyer M, Brambilla R, Lambertsen KL (2019) Topical Administration of a Soluble TNF Inhibitor Reduces Infarct Volume After Focal Cerebral Ischemia in Mice. Front Neurosci 13:781
Yoshida M, Satoh A, Lin JB, Mills KF, Sasaki Y, Rensing N, Wong M, Apte RS, Imai SI (2019) Extracellular Vesicle-Contained eNAMPT Delays Aging and Extends Lifespan in Mice. Cell Metab 30:329-342 e325
Zhang F, Li X, Ni Y, Shan G, Gao Y (2019) Preliminary study of the urinary proteome in Li and Han ethnic individuals from Hainan. Sci China Life Sci (in press)
Zhang Y, Natale R, Domingues APJ, Toleco MR, Siemiatkowska B, Fabregas N, Fernie AR (2019) Rapid Identification of Protein-Protein Interactions in Plants. Curr Protoc Plant Biol 4:e20099
2018
Aguilar-Hernandez V, Loyola-Vargas VM (2018) Advanced Proteomic Approaches to Elucidate Somatic Embryogenesis. Front Plant Sci 9:1658
Al Akeel R, Mateen A, Syed R, Alqahtani MS, Alqahtani AS (2018) Alanine rich peptide from Populus trichocarpa inhibit growth of Staphylococcus aureus via targetting its extracellular domain of Sensor Histidine Kinase YycGex protein. Microb Pathog 121:115-122
Al Feteisi H, Al-Majdoub ZM, Achour B, Couto N, Rostami-Hodjegan A, Barber J (2018) Identification and quantification of blood-brain barrier transporters in isolated rat brain microvessels. J Neurochem 146:670-685
Albrecht S, Kaisermayer C, Gallagher C, Farrell A, Lindeberg A, Bones J (2018) Proteomics in biomanufacturing control: Protein dynamics of CHO-K1 cells and conditioned media during apoptosis and necrosis. Biotechnol Bioeng 115:1509-1520
Alcoriza-Balaguer MI, Garcia-Canaveras JC, Lopez A, Conde I, Juan O, Carretero J, Lahoz A (2018) LipidMS: an R package for lipid annotation in untargeted liquid chromatography-data independent acquisition-mass spectrometry lipidomics. Anal Chem (in press)
Amadeo F, Boschetti F, Polvani G, Banfi C, Pesce M, Santoro R (2018) Aortic valve cell seeding into decellularized animal pericardium by perfusion-assisted bioreactor. J Tissue Eng Regen Med 12:1481-1493
Amanzougaghene N, Fenollar F, Nappez C, Ben-Amara A, Decloquement P, Azza S, Bechah Y, Chabriere E, Raoult D, Mediannikov O (2018) Complexin in ivermectin resistance in body lice. PLoS Genet 14:e1007569
Anitua E, de la Fuente M, Muruzabal F, Sanchez-Avila RM, Merayo-Lloves J, Azkargorta M, Elortza F, Orive G (2018) Differential profile of protein expression on human keratocytes treated with autologous serum and plasma rich in growth factors (PRGF). PLoS One 13:e0205073
Araujo-Gomes N, Romero-Gavilan F, Garcia-Arnaez I, Martinez-Ramos C, Sanchez-Perez AM, Azkargorta M, Elortza F, de Llano JJM, Gurruchaga M, Goni I, Suay J (2018) Osseointegration mechanisms: a proteomic approach. J Biol Inorg Chem 23:459-470
Arquint C, Cubizolles F, Morand A, Schmidt A, Nigg EA (2018) The SKP1-Cullin-F-box E3 ligase betaTrCP and CDK2 cooperate to control STIL abundance and centriole number. Open Biol 8 (in press)
Asadian M, Dhaenens M, Onyshchenko I, De Waele S, Declercq H, Cools P, Devreese B, Deforce D, Morent R, De Geyter N (2018) Plasma Functionalization of Polycaprolactone Nanofibers Changes Protein Interactions with Cells, Resulting in Increased Cell Viability. ACS Appl Mater Interfaces (in press)
Bailey J, Davis S, Shaw A, Diotallevi M, Fischer R, Benson MA, Zhu H, Brown J, Bhattacharya S, Kessler BM, Channon KM, Crabtree MJ (2018) Tetrahydrobiopterin modulates ubiquitin conjugation to UBC13/UBE2N and proteasome activity by S-nitrosation. Sci Rep 8:14310
Bao K, Bostanci N, Thurnheer T, Grossmann J, Wolski WE, Thay B, Belibasakis GN, Oscarsson J (2018) Aggregatibacter actinomycetemcomitans H-NS promotes biofilm formation and alters protein dynamics of other species within a polymicrobial oral biofilm. NPJ Biofilms Microbiomes 4:12
Barrachina MN, Sueiro AM, Casas V, Izquierdo I, Hermida-Nogueira L, Guitian E, Casanueva FF, Abian J, Carrascal M, Pardo M, Garcia A (2018) A Combination of Proteomic Approaches Identifies A Panel of Circulating Extracellular Vesicle Proteins Related to the Risk of Suffering Cardiovascular Disease in Obese Patients. Proteomics:e1800248
Bastaert F, Kheir S, Saint-Criq V, Villeret B, Dang PM, El-Benna J, Sirard JC, Voulhoux R, Sallenave JM (2018) Pseudomonas aeruginosa LasB Subverts Alveolar Macrophage Activity by Interfering With Bacterial Killing Through Downregulation of Innate Immune Defense, Reactive Oxygen Species Generation, and Complement Activation. Front Immunol 9:1675
Bauza-Martinez J, Aletti F, Pinto BB, Ribas V, Odena MA, Diaz R, Romay E, Ferrer R, Kistler EB, Tedeschi G, Schmid-Schonbein GW, Herpain A, Bendjelid K, de Oliveira E (2018) Proteolysis in septic shock patients: plasma peptidomic patterns are associated with mortality. Br J Anaesth 121:1065-1074
Bazaid AS, Forbes S, Humphreys GJ, Ledder RG, O'Cualain R, McBain AJ (2018) Fatty Acid Supplementation Reverses the Small Colony Variant Phenotype in Triclosan-Adapted Staphylococcus aureus: Genetic, Proteomic and Phenotypic Analyses. Sci Rep 8:3876
Belchi-Navarro S, Almagro L, Bru-Martinez R, Pedreno MA (2018) Changes in the secretome of Vitis vinifera cv. Monastrell cell cultures treated with cyclodextrins and methyl jasmonate. Plant Physiol Biochem (in press)
Bell L, Calder B, Hiller R, Klein A, Soares NC, Stoychev SH, Vorster BC, Tabb DL (2018) Challenges and Opportunities for Biological Mass Spectrometry Core Facilities in the Developing World. J Biomol Tech 29:4-15
Berenguer M, Darnaudery M, Claverol S, Bonneu M, Lacombe D, Rooryck C (2018) Prenatal retinoic acid exposure reveals candidate genes for craniofacial disorders. Sci Rep 8:17492
Berle M, Ghila L, Vethe H, Chaudhry A, Garberg H, Beisland C, Haaland OA, Oveland E, Halvorsen OJ, Davidsson T, Chera S (2018) Novel protein signatures suggest progression to muscular invasiveness in bladder cancer. PLoS One 13:e0206475
Bonnet J, Garcia C, Leger T, Couquet MP, Vignoles P, Vatunga G, Ndung'u J, Boudot C, Bisser S, Courtioux B (2018) Proteome characterization in various biological fluids of Trypanosoma brucei gambiense-infected subjects. J Proteomics (in press)
Bostanci N, Bao K, Li X, Maekawa T, Grossmann J, Panse C, Briones RA, Resuello RRG, Tuplano JV, Garcia CAG, Reis ES, Lambris JD, Hajishengallis G (2018) Gingival Exudatome Dynamics Implicate Inhibition of the Alternative Complement Pathway in the Protective Action of the C3 Inhibitor Cp40 in Nonhuman Primate Periodontitis. J Proteome Res 17:3153-3175
Bostanci N, Belibasakis GN (2018) Gingival crevicular fluid and its immune mediators in the proteomic era. Periodontol 2000 76:68-84
Bradley F, Birse K, Hasselrot K, Noel-Romas L, Introini A, Wefer H, Seifert M, Engstrand L, Tjernlund A, Broliden K, Burgener AD (2018) The vaginal microbiome amplifies sex hormone-associated cyclic changes in cervicovaginal inflammation and epithelial barrier disruption. Am J Reprod Immunol 80:e12863
Bricio-Moreno L, Sheridan VH, Goodhead I, Armstrong S, Wong JKL, Waters EM, Sarsby J, Panagiotou S, Dunn J, Chakraborty A, Fang Y, Griswold KE, Winstanley C, Fothergill JL, Kadioglu A, Neill DR (2018) Evolutionary trade-offs associated with loss of PmrB function in host-adapted Pseudomonas aeruginosa. Nat Commun 9:2635
Buffon G, Blasi E, Rativa AGS, Lamb TI, Gastmann R, Adamski JM, Schwambach J, Ricachenevsky FK, Heringer AS, Silveira V, Lopes MCB, Sperotto RA (2018) Unraveling Rice Tolerance Mechanisms Against Schizotetranychus oryzae Mite Infestation. Front Plant Sci 9:1341
Buhner S, Hahne H, Hartwig K, Li Q, Vignali S, Ostertag D, Meng C, Hormannsperger G, Braak B, Pehl C, Frieling T, Barbara G, De Giorgio R, Demir IE, Ceyhan GO, Zeller F, Boeckxstaens G, Haller D, Kuster B, Schemann M (2018) Protease signaling through protease activated receptor 1 mediate nerve activation by mucosal supernatants from irritable bowel syndrome but not from ulcerative colitis patients. PLoS One 13:e0193943
Burschel S, Kreuzer Decovic D, Nuber F, Stiller M, Hofmann M, Zupok A, Siemiatkowska B, Gorka M, Leimkuhler S, Friedrich T (2018) Iron-sulfur cluster carrier proteins involved in the assembly of Escherichia coli NADH:ubiquinone oxidoreductase (complex I). Mol Microbiol (in press)
Cassoli JS, Brandao-Teles C, Santana AG, Souza G, Martins-de-Souza D (2018) Ion Mobility-Enhanced Data-Independent Acquisitions Enable a Deep Proteomic Landscape of Oligodendrocytes. Proteomics 18 (in press)
Ceballos-Laita L, Gutierrez-Carbonell E, Imai H, Abadia A, Uemura M, Abadia J, Lopez-Millan AF (2018) Effects of manganese toxicity on the protein profile of tomato (Solanum lycopersicum) roots as revealed by two complementary proteomic approaches, two-dimensional electrophoresis and shotgun analysis. J Proteomics 185:51-63
Ceballos-Laita L, Gutierrez-Carbonell E, Takahashi D, Abadia A, Uemura M, Abadia J, Lopez-Millan AF (2018) Data on xylem sap proteins from Mn- and Fe-deficient tomato plants obtained using shotgun proteomics. Data Brief 17:512-516
Ceballos-Laita L, Gutierrez-Carbonell E, Takahashi D, Abadia A, Uemura M, Abadia J, Lopez-Millan AF (2018) Effects of Fe and Mn deficiencies on the protein profiles of tomato (Solanum lycopersicum) xylem sap as revealed by shotgun analyses. J Proteomics 170:117-129
Chevreux G, Tilly N, Bihoreau N (2018) Quantification of proteins by data independent acquisition: Performance assessment of the Hi3 methodology. Anal Biochem 549:184-187
Cobbaut M, Derua R, Parker PJ, Waelkens E, Janssens V, Van Lint J (2018) Protein kinase D displays intrinsic Tyr autophosphorylation activity: insights into mechanism and regulation. FEBS Lett (in press)
Costello A, Coleman O, Lao NT, Henry M, Meleady P, Barron N, Clynes M (2018) Depletion of endogenous miRNA-378-3p increases peak cell density of CHO DP12 cells and is correlated with elevated levels of ubiquitin carboxyl-terminal hydrolase 14. J Biotechnol 288:30-40
Daulat AM, Puvirajesinghe TM, Camoin L, Borg JP (2018) Mapping Cellular Polarity Networks Using Mass Spectrometry-based Strategies. J Mol Biol
De A, Pompilio A, Francis J, Sutcliffe IC, Black GW, Lupidi G, Petrelli D, Vitali LA (2018) Antidiabetic "gliptins" affect biofilm formation by Streptococcus mutans. Microbiol Res 209:79-85
Denard J, Rouillon J, Leger T, Garcia C, Lambert MP, Griffith G, Jenny C, Camadro JM, Garcia L, Svinartchouk F (2018) AAV-8 and AAV-9 Vectors Cooperate with Serum Proteins Differently Than AAV-1 and AAV-6. Mol Ther Methods Clin Dev 10:291-302
Depluverez S, Daled S, De Waele S, Planckaert S, Schoovaerts J, Deforce D, Devreese B (2018) Microfluidics-based LC-MS MRM approach for the relative quantification of Burkholderia cenocepacia secreted virulence factors. Rapid Commun Mass Spectrom (in press)
Derricott H, Luu L, Fong WY, Hartley CS, Johnston LJ, Armstrong SD, Randle N, Duckworth CA, Campbell BJ, Wastling JM, Coombes JL (2018) Developing a 3D intestinal epithelium model for livestock species. Cell Tissue Res (in press)
Dhaenens L, Lierman S, De Clerck L, Govaert E, Deforce D, Tilleman K, De Sutter P (2018) Endometrial stromal cell proteome mapping in repeated implantation failure and recurrent pregnancy loss cases and fertile women. Reprod Biomed Online (in press)
do Carmo FLR, Silva WM, Tavares GC, Ibraim IC, Cordeiro BF, Oliveira ER, Rabah H, Cauty C, da Silva SH, Canario Viana MV, Caetano ACB, Dos Santos RG, de Oliveira Carvalho RD, Jardin J, Pereira FL, Folador EL, Le Loir Y, Figueiredo HCP, Jan G, Azevedo V (2018) Mutation of the Surface Layer Protein SlpB Has Pleiotropic Effects in the Probiotic Propionibacterium freudenreichii CIRM-BIA 129. Front Microbiol 9:1807
Dong MX, Feng X, Xu XM, Hu L, Liu Y, Jia SY, Li B, Chen W, Wei YD (2018) Integrated Analysis Reveals Altered Lipid and Glucose Metabolism and Identifies NOTCH2 as a Biomarker for Parkinson's Disease Related Depression. Front Mol Neurosci 11:257
Dornadula S, Thiruppathi S, Palanisamy R, Umapathy D, Suzuki T, R KM (2018) Differential proteomic profiling identifies novel molecular targets of pterostilbene against experimental diabetes. J Cell Physiol 234:1996-2012
Draime A, Bridoux L, Belpaire M, Pringels T, Degand H, Morsomme P, Rezsohazy R (2018) The O-GlcNAc transferase OGT interacts with and post-translationally modifies the transcription factor HOXA1. FEBS Lett 592:1185-1201
Dreier RF, Ahrne E, Broz P, Schmidt A (2018) Global Ion Suppression Limits the Potential of Mass Spectrometry Based Phosphoproteomics. J Proteome Res (in press)
El-Daher MT, Cagnard N, Gil M, Da Cruz MC, Leveau C, Sepulveda F, Zarhrate M, Tores F, Legoix P, Baulande S, de Villartay JP, Almouzni G, Quivy JP, Fischer A, de Saint Basile G (2018) Tetratricopeptide repeat domain 7A is a nuclear factor that modulates transcription and chromatin structure. Cell Discov 4:61
Finamore F, Reny JL, Malacarne S, Fontana P, Sanchez JC (2018) Shotgun proteomics data on the impact of hyperglycaemia on platelet protein acetylation by aspirin. Data Brief 21:2475-2481
Fingas F, Volke D, Bielefeldt P, Hassert R, Hoffmann R (2018) Detection of mammalian orthoreovirus type-3 (Reo-3) infections in mice based on serotype-specific hemagglutination protein sigma-1. Virol J 15:114
Fiorillo M, Peiris-Pages M, Sanchez-Alvarez R, Bartella L, Di Donna L, Dolce V, Sindona G, Sotgia F, Cappello AR, Lisanti MP (2018) Bergamot natural products eradicate cancer stem cells (CSCs) by targeting mevalonate, Rho-GDI-signalling and mitochondrial metabolism. Biochim Biophys Acta 1859:984-996
Firmino M, Weis SN, Souza JMF, Gomes BRB, Mol AR, Mortari MR, Souza GEP, Coca GC, Williams TCR, Fontes W, Ricart CAO, de Sousa MV, Veiga-Souza FH (2018) Label-free quantitative proteomics of rat hypothalamus under fever induced by LPS and PGE2. J Proteomics 187:182-199
Flechard M, Duchesne R, Tahrioui A, Bouffartigues E, Depayras S, Hardouin J, Lagy C, Maillot O, Tortuel D, Azuama CO, Clamens T, Duclairoir-Poc C, Catel-Ferreira M, Gicquel G, Feuilloley MGJ, Lesouhaitier O, Heipieper HJ, Groleau MC, Deziel E, Cornelis P, Chevalier S (2018) The absence of SigX results in impaired carbon metabolism and membrane fluidity in Pseudomonas aeruginosa. Sci Rep 8:17212
Frik J, Merl-Pham J, Plesnila N, Mattugini N, Kjell J, Kraska J, Gomez RM, Hauck SM, Sirko S, Gotz M (2018) Cross-talk between monocyte invasion and astrocyte proliferation regulates scarring in brain injury. EMBO Rep 19 (in press)
Froyset AK, Edson AJ, Gharbi N, Khan EA, Dondorp D, Bai Q, Tiraboschi E, Suster ML, Connolly JB, Burton EA, Fladmark KE (2018) Astroglial DJ-1 over-expression up-regulates proteins involved in redox regulation and is neuroprotective in vivo. Redox Biol 16:237-247
Gago S, Overton NLD, Ben-Ghazzi N, Novak-Frazer L, Read ND, Denning DW, Bowyer P (2018) Lung colonization by Aspergillus fumigatus is controlled by ZNF77. Nat Commun 9:3835
Garalla HM, Lertkowit N, Tiszlavicz L, Reisz Z, Holmberg C, Beynon R, Simpson D, Varga A, Kumar JD, Dodd S, Pritchard DM, Moore AR, Rosztoczy AI, Wittman T, Simpson A, Dockray GJ, Varro A (2018) Matrix metalloproteinase (MMP)-7 in Barrett's esophagus and esophageal adenocarcinoma: expression, metabolism, and functional significance. Physiol Rep 6:e13683
Garcia-Murria MJ, Sudhani HPK, Marin-Navarro J, Sanchez Del Pino MM, Moreno J (2018) Dissecting the individual contribution of conserved cysteines to the redox regulation of RubisCO. Photosynth Res 137:251-262
Gautier F, Eliasova K, Leple JC, Vondrakova Z, Lomenech AM, Le Mette C, Label P, Costa G, Trontin JF, Teyssier C, Lelu-Walter MA (2018) Repetitive somatic embryogenesis induced cytological and proteomic changes in embryogenic lines of Pseudotsuga menziesii [Mirb.]. BMC Plant Biol 18:164
Gaviard C, Broutin I, Cosette P, De E, Jouenne T, Hardouin J (2018) Lysine Succinylation and Acetylation in Pseudomonas aeruginosa. J Proteome Res 17:2449-2459
Geilfus CM, Lan J, Carpentier S (2018) Dawn regulates guard cell proteins in Arabidopsis thaliana that function in ATP production from fatty acid beta-oxidation. Plant Mol Biol 98:525-543
Gennaro VJ, Stanek TJ, Peck AR, Sun Y, Wang F, Qie S, Knudsen KE, Rui H, Butt T, Diehl JA, McMahon SB (2018) Control of CCND1 ubiquitylation by the catalytic SAGA subunit USP22 is essential for cell cycle progression through G1 in cancer cells. Proc Natl Acad Sci U S A 115:E9298-E9307
Gentric G, Kieffer Y, Mieulet V, Goundiam O, Bonneau C, Nemati F, Hurbain I, Raposo G, Popova T, Stern MH, Lallemand-Breitenbach V, Muller S, Caneque T, Rodriguez R, Vincent-Salomon A, de The H, Rossignol R, Mechta-Grigoriou F (2018) PML-Regulated Mitochondrial Metabolism Enhances Chemosensitivity in Human Ovarian Cancers. Cell Metab 29:156-173 e110
Ghezraoui H, Oliveira C, Becker JR, Bilham K, Moralli D, Anzilotti C, Fischer R, Deobagkar-Lele M, Sanchiz-Calvo M, Fueyo-Marcos E, Bonham S, Kessler BM, Rottenberg S, Cornall RJ, Green CM, Chapman JR (2018) 53BP1 cooperation with the REV7-shieldin complex underpins DNA structure-specific NHEJ. Nature 560:122-127
Gillan V, Simpson DM, Kinnaird J, Maitland K, Shiels B, Devaney E (2018) Characterisation of infection associated microRNA and protein cargo in extracellular vesicles of Theileria annulata infected leukocytes. Cell Microbiol 21:e12969
Gioria S, Urban P, Hajduch M, Barboro P, Cabaleiro N, La Spina R, Chassaigne H (2018) Proteomics study of silver nanoparticles on Caco-2 cells. Toxicol In Vitro 50:347-372
Girard L, Birse K, Holm JB, Gajer P, Humphrys MS, Garber D, Guenthner P, Noel-Romas L, Abou M, McCorrister S, Westmacott G, Wang L, Rohan LC, Matoba N, McNicholl J, Palmer KE, Ravel J, Burgener AD (2018) Impact of the griffithsin anti-HIV microbicide and placebo gels on the rectal mucosal proteome and microbiome in non-human primates. Sci Rep 8:8059
Goel RK, Meyer M, Paczkowska M, Reimand J, Vizeacoumar F, Lam TT, Lukong KE (2018) Global phosphoproteomic analysis identifies SRMS-regulated secondary signaling intermediates. Proteome Sci 16:16
Gonzalez Coraspe JA, Weis J, Anderson ME, Munchberg U, Lorenz K, Buchkremer S, Carr S, Zahedi RP, Brauers E, Michels H, Sunada Y, Lochmuller H, Campbell KP, Freier E, Hathazi D, Roos A (2018) Biochemical and pathological changes result from mutated Caveolin-3 in muscle. Skelet Muscle 8:28
Gonzalez-Morales A, Zabaleta A, Garcia-Moure M, Alonso MM, Fernandez-Irigoyen J, Santamaria E (2018) Oncolytic adenovirus Delta-24-RGD induces a widespread glioma proteotype remodeling during autophagy. J Proteomics (in press)
Grube L, Dellen R, Kruse F, Schwender H, Stuhler K, Poschmann G (2018) Mining the Secretome of C2C12 Muscle Cells: Data Dependent Experimental Approach To Analyze Protein Secretion Using Label-Free Quantification and Peptide Based Analysis. J Proteome Res 17:879-890
Gueugneau M, d'Hose D, Barbe C, de Barsy M, Lause P, Maiter D, Bindels LB, Delzenne NM, Schaeffer L, Gangloff YG, Chambon C, Coudy-Gandilhon C, Bechet D, Thissen JP (2018) Increased Serpina3n release into circulation during glucocorticoid-mediated muscle atrophy. J Cachexia Sarcopenia Muscle (in press)
Guo Z, Wang Z, Lu C, Yang S, Sun H, Reziw, Guo Y, Sun W, Yue H (2018) Analysis of the differential urinary protein profile in IgA nephropathy patients of Uygur ethnicity. BMC Nephrol 19:358
Hadjidemetriou M, Al-Ahmady Z, Buggio M, Swift J, Kostarelos K (2018) A novel scavenging tool for cancer biomarker discovery based on the blood-circulating nanoparticle protein corona. Biomaterials 188:118-129
Hakobyan A, Liesack W, Glatter T (2018) Crude-MS Strategy for in-Depth Proteome Analysis of the Methane-Oxidizing Methylocystis sp. strain SC2. J Proteome Res 17:3086-3103
Hakuno D, Kimura M, Ito S, Satoh J, Nakashima Y, Horie T, Kuwabara Y, Nishiga M, Ide Y, Baba O, Nishi H, Nakao T, Nishino T, Nakazeki F, Koyama S, Hanada R, Randolph RR, Endo J, Kimura T, Ono K (2018) Hepatokine alpha1-Microglobulin Signaling Exacerbates Inflammation and Disturbs Fibrotic Repair in Mouse Myocardial Infarction. Sci Rep 8:16749
Heath N, Grant L, De Oliveira TM, Rowlinson R, Osteikoetxea X, Dekker N, Overman R (2018) Rapid isolation and enrichment of extracellular vesicle preparations using anion exchange chromatography. Sci Rep 8:5730
Heavey S, Dowling P, Moore G, Barr MP, Kelly N, Maher SG, Cuffe S, Finn SP, O'Byrne KJ, Gately K (2018) Development and characterisation of a panel of phosphatidylinositide 3-kinase - mammalian target of rapamycin inhibitor resistant lung cancer cell lines. Sci Rep 8:1652
Hemadou A, Laroche-Traineau J, Antoine S, Mondon P, Fontayne A, Le Priol Y, Claverol S, Sanchez S, Cerutti M, Ottones F, Clofent-Sanchez G, Jacobin-Valat MJ (2018) An innovative flow cytometry method to screen human scFv-phages selected by in vivo phage-display in an animal model of atherosclerosis. Sci Rep 8:15016
Henry M, Gallagher C, Kelly RM, Frye CC, Osborne MD, Brady CP, Barron N, Clynes M, Meleady P (2018) Clonal variation in productivity and proteolytic clipping of an Fc-fusion protein in CHO cells: Proteomic analysis suggests a role for defective protein folding and the UPR. J Biotechnol 281:21-30
Hindupur SK, Colombi M, Fuhs SR, Matter MS, Guri Y, Adam K, Cornu M, Piscuoglio S, Ng CKY, Betz C, Liko D, Quagliata L, Moes S, Jenoe P, Terracciano LM, Heim MH, Hunter T, Hall MN (2018) The protein histidine phosphatase LHPP is a tumour suppressor. Nature 555:678-682
Holm M, Saraswat M, Joenvaara S, Ristimaki A, Haglund C, Renkonen R (2018) Colorectal cancer patients with different C-reactive protein levels and 5-year survival times can be differentiated with quantitative serum proteomics. PLoS One 13:e0195354
Ibort P, Imai H, Uemura M, Aroca R (2018) Proteomic analysis reveals that tomato interaction with plant growth promoting bacteria is highly determined by ethylene perception. J Plant Physiol 220:43-59
Imamovic L, Ellabaan MMH, Dantas Machado AM, Citterio L, Wulff T, Molin S, Krogh Johansen H, Sommer MOA (2018) Drug-Driven Phenotypic Convergence Supports Rational Treatment Strategies of Chronic Infections. Cell 172:121-134 e114
Infante A, Rodriguez CI (2018) Secretome analysis of in vitro aged human mesenchymal stem cells reveals IGFBP7 as a putative factor for promoting osteogenesis. Sci Rep 8:4632
Izzi-Engbeaya C, Comninos AN, Clarke SA, Jomard A, Yang L, Jones S, Abbara A, et al. (2018) The Effects of Kisspeptin on beta-cell Function, Serum Metabolites and Appetite in Humans. Diabetes Obes Metab (in press)
Jaleel A, Aneesh Kumar A, Ajith Kumar GS, Surendran A, Kartha CC (2018) Label-free quantitative proteomics analysis reveals distinct molecular characteristics in endocardial endothelium. Mol Cell Biochem (in press)
Jamin A, Berthelot L, Couderc A, Chemouny JM, Boedec E, Dehoux L, Abbad L, Dossier C, Daugas E, Monteiro RC, Deschenes G (2018) Autoantibodies against podocytic UCHL1 are associated with idiopathic nephrotic syndrome relapses and induce proteinuria in mice. J Autoimmun 89:149-161
Janes MR, Zhang J, Li LS, Hansen R, Peters U, Guo X, Chen Y, et al. (2018) Targeting KRAS Mutant Cancers with a Covalent G12C-Specific Inhibitor. Cell 172:578-589 e517
Janowski M, Zoschke R, Scharff LB, Martinez Jaime S, Ferrari C, Proost S, Ng Wei Xiong J, Omranian N, Musialak-Lange M, Nikoloski Z, Graf A, Schottler MA, Sampathkumar A, Vaid N, Mutwil M (2018) AtRsgA from Arabidopsis thaliana is important for maturation of the small subunit of the chloroplast ribosome. Plant J 96:404-420
Jeitany M, Leroy C, Tosti P, Lafitte M, Le Guet J, Simon V, Bonenfant D, Robert B, Grillet F, Mollevi C, El Messaoudi S, Otandault A, Canterel-Thouennon L, Busson M, Thierry AR, Martineau P, Pannequin J, Roche S, Sirvent A (2018) Inhibition of DDR1-BCR signalling by nilotinib as a new therapeutic strategy for metastatic colorectal cancer. EMBO Mol Med 10 (in press)
Jing J, Du Z, Ji S, Han K (2019) Urinary proteome analysis of acute hypercoagulable state in rat model induced by epsilon-aminocaproic acid. Biomed Pharmacother 110:275-284
Joenvaara S, Saraswat M, Kuusela P, Saraswat S, Agarwal R, Kaartinen J, Jarvinen A, Renkonen R (2018) Quantitative N-glycoproteomics reveals altered glycosylation levels of various plasma proteins in bloodstream infected patients. PLoS One 13:e0195006
Jones MC, Askari JA, Humphries JD, Humphries MJ (2018) Cell adhesion is regulated by CDK1 during the cell cycle. J Cell Biol 217:3203-3218
Juvvadi PR, Moseley MA, Hughes CJ, Soderblom EJ, Lennon S, Perkins SR, Thompson JW, Geromanos SJ, Wildgoose J, Richardson K, Langridge JI, Vissers JPC, Steinbach WJ (2018) Scanning Quadrupole Data-Independent Acquisition, Part B: Application to the Analysis of the Calcineurin-Interacting Proteins during Treatment of Aspergillus fumigatus with Azole and Echinocandin Antifungal Drugs. J Proteome Res 17:780-793
Karri V, Ramos D, Martinez JB, Odena A, Oliveira E, Coort SL, Evelo CT, Mariman ECM, Schuhmacher M, Kumar V (2018) Differential protein expression of hippocampal cells associated with heavy metals (Pb, As, and MeHg) neurotoxicity: Deepening into the molecular mechanism of neurodegenerative diseases. J Proteomics 187:106-125
Kaushik P, Henry M, Clynes M, Meleady P (2018) The Expression Pattern of the Phosphoproteome Is Significantly Changed During the Growth Phases of Recombinant CHO Cell Culture. Biotechnol J 13:e1700221
Kay LJ, Sangal V, Black GW, Soundararajan M (2018) Proteomics and bioinformatics analyses identify novel cellular roles outside mitochondrial function for human miro GTPases. Mol Cell Biochem (in press)
Keck M, van Dijk RM, Deeg CA, Kistler K, Walker A, von Ruden EL, Russmann V, Hauck SM, Potschka H (2018) Proteomic profiling of epileptogenesis in a rat model: Focus on cell stress, extracellular matrix and angiogenesis. Neurobiol Dis 112:119-135
Kirkpatrick CL, Parsley NC, Bartges TE, Cooke ME, Evans WS, Heil LR, Smith TJ, Hicks LM (2018) Fungal Secretome Analysis via PepSAVI-MS: Identification of the Bioactive Peptide KP4 from Ustilago maydis. J Am Soc Mass Spectrom 29:859-865
Kirkpatrick CL, Parsley NC, Bartges TE, Wing CE, Kommineni S, Kristich CJ, Salzman NH, Patrie SM, Hicks LM (2018) Exploring bioactive peptides from bacterial secretomes using PepSAVI-MS: identification and characterization of Bac-21 from Enterococcus faecalis pPD1. Microb Biotechnol (in press)
Kisrieva YS, Petushkova NA, Samenkova NF, Kuznetsova GP, Larina OV, Teryaeva NB, Zgoda VG, Karuzina, II, Usachev DU, Belyaev AY (2018) Analysis of Blood Plasma Protein Composition in Patients with Cerebral Ischemia. Bull Exp Biol Med 165:22-26
Koirala D, Beranova-Giorgianni S, Giorgianni F (2018) Data-independent proteome analysis of ARPE-19 cells. Data Brief 20:333-336
Konig S, Hadrian K, Schlatt S, Wistuba J, Thanos S, Bohm MRR (2019) Topographic protein profiling of the age-related proteome in the retinal pigment epithelium of Callithrix jacchus with respect to macular degeneration. J Proteomics 191:1-15
Kotini M, Barriga EH, Leslie J, Gentzel M, Rauschenberger V, Schambony A, Mayor R (2018) Gap junction protein Connexin-43 is a direct transcriptional regulator of N-cadherin in vivo. Nat Commun 9:3846
Krahmer J, Goralogia GS, Kubota A, Zardilis A, Johnson RS, Song YH, MacCoss MJ, Le Bihan T, Halliday KJ, Imaizumi T, Millar AJ (2018) Time-resolved interaction proteomics of the GIGANTEA protein under diurnal cycles in Arabidopsis. FEBS Lett (in press)
Lachen-Montes M, Gonzalez-Morales A, Iloro I, Elortza F, Ferrer I, Gveric D, Fernandez-Irigoyen J, Santamaria E (2018) Unveiling the olfactory proteostatic disarrangement in Parkinson's disease by proteome-wide profiling. Neurobiol Aging 73:123-134
Lalowski MM, Bjork S, Finckenberg P, Soliymani R, Tarkia M, Calza G, Blokhina D, Tulokas S, Kankainen M, Lakkisto P, Baumann M, Kankuri E, Mervaala E (2018) Characterizing the Key Metabolic Pathways of the Neonatal Mouse Heart Using a Quantitative Combinatorial Omics Approach. Front Physiol 9:365
Le Faouder J, Gigante E, Leger T, Albuquerque M, Beaufrere A, Soubrane O, Dokmak S, Camadro JM, Cros J, Paradis V (2018) Proteomic Landscape of Cholangiocarcinomas Reveals Three Different Subgroups According to Their Localization and the Aspect of Non-Tumor Liver. Proteomics Clin Appl:e1800128
Lepper MF, Ohmayer U, von Toerne C, Maison N, Ziegler AG, Hauck SM (2018) Proteomic Landscape of Patient-Derived CD4+ T Cells in Recent-Onset Type 1 Diabetes. J Proteome Res 17:618-634
Leutert M, Menzel S, Braren R, Rissiek B, Hopp AK, Nowak K, Bisceglie L, Gehrig P, Li H, Zolkiewska A, Koch-Nolte F, Hottiger MO (2018) Proteomic Characterization of the Heart and Skeletal Muscle Reveals Widespread Arginine ADP-Ribosylation by the ARTC1 Ectoenzyme. Cell Rep 24:1916-1929 e1915
Li J, Lu Q, Lu P (2018) Quantitative proteomics analysis of vitreous body from type 2 diabetic patients with proliferative diabetic retinopathy. BMC Ophthalmol 18:151
Li W, Papilloud A, Lozano-Montes L, Zhao N, Ye X, Zhang X, Sandi C, Rainer G (2018) Stress Impacts the Regulation Neuropeptides in the Rat Hippocampus and Prefrontal Cortex. Proteomics 18:e1700408
Light SH, Su L, Rivera-Lugo R, Cornejo JA, Louie A, Iavarone AT, Ajo-Franklin CM, Portnoy DA (2018) A flavin-based extracellular electron transfer mechanism in diverse Gram-positive bacteria. Nature 562:140-144
Lock JG, Jones MC, Askari JA, Gong X, Oddone A, Olofsson H, Goransson S, Lakadamyali M, Humphries MJ, Stromblad S (2018) Reticular adhesions are a distinct class of cell-matrix adhesions that mediate attachment during mitosis. Nat Cell Biol 20:1290-1302
Lucero Camacho-Morales R, Garcia-Fontana C, Fernandez-Irigoyen J, Santamaria E, Gonzalez-Lopez J, Manzanera M, Aranda E (2018) Anthracene drives sub-cellular proteome-wide alterations in the degradative system of Penicillium oxalicum. Ecotoxicol Environ Saf 159:127-135
Luscher A, Moynie L, Auguste PS, Bumann D, Mazza L, Pletzer D, Naismith JH, Kohler T (2018) TonB-Dependent Receptor Repertoire of Pseudomonas aeruginosa for Uptake of Siderophore-Drug Conjugates. Antimicrob Agents Chemother 62 (in press)
Luu L, Matthews ZJ, Armstrong SD, Powell P, Wileman T, Wastling JM, Coombes JL (2018) Proteomic Profiling of Enteroid Cultures Skewed Towards Development of Specific Epithelial Lineages. Proteomics:e1800132
Maciel VL, Jr., Caldas-Bussiere MC, Silveira V, Reis RS, Rios AFL, Paes de Carvalho CS (2018) l-arginine alters the proteome of frozen-thawed bovine sperm during in vitro capacitation. Theriogenology 119:1-9
Malheiros JM, Braga CP, Grove RA, Ribeiro FA, Calkins CR, Adamec J, Chardulo LAL (2018) Influence of oxidative damage to proteins on meat tenderness using a proteomics approach. Meat Sci 148:64-71
McConnell EW, Werth EG, Hicks LM (2018) The phosphorylated redox proteome of Chlamydomonas reinhardtii: Revealing novel means for regulation of protein structure and function. Redox Biol 17:35-46
Mi Y, Sun C, Wei B, Sun F, Guo Y, Hu Q, Ding W, Zhu L, Xia G (2018) Coatomer subunit beta 2 (COPB2), identified by label-free quantitative proteomics, regulates cell proliferation and apoptosis in human prostate carcinoma cells. Biochem Biophys Res Commun 495:473-480
Miki Y, Takahashi D, Kawamura Y, Uemura M (2018) Temporal proteomics of Arabidopsis plasma membrane during cold- and de-acclimation. J Proteomics (in press)
Milkovska-Stamenova S, Krieg L, Hoffmann R (2018) Products of Early and Advanced Glycation in the Soy Milk Proteome. Mol Nutr Food Res:e1800725
Millership SJ, Da Silva Xavier G, Choudhury AI, Bertazzo S, Chabosseau P, Pedroni SM, Irvine EE, Montoya A, Faull P, Taylor WR, Kerr-Conte J, Pattou F, Ferrer J, Christian M, John RM, Latreille M, Liu M, Rutter GA, Scott J, Withers DJ (2018) Neuronatin regulates pancreatic beta cell insulin content and secretion. J Clin Invest 128:3369-3381
Monteiro R, Chafsey I, Leroy S, Chambon C, Hebraud M, Livrelli V, Pizza M, Pezzicoli A, Desvaux M (2018) Differential biotin labelling of the cell envelope proteins in lipopolysaccharidic diderm bacteria: Exploring the proteosurfaceome of Escherichia coli using sulfo-NHS-SS-biotin and sulfo-NHS-PEG4-bismannose-SS-biotin. J Proteomics 181:16-23
Morra R, Del Carratore F, Muhamadali H, Horga LG, Halliwell S, Goodacre R, Breitling R, Dixon N (2018) Translation Stress Positively Regulates MscL-Dependent Excretion of Cytoplasmic Proteins. MBio 9 (in press)
Muller J, Prozeller D, Ghazaryan A, Kokkinopoulou M, Mailander V, Morsbach S, Landfester K (2018) Beyond the protein corona - lipids matter for biological response of nanocarriers. Acta Biomater 71:420-431
Muller J, Simon J, Rohne P, Koch-Brandt C, Mailander V, Morsbach S, Landfester K (2018) Denaturation via Surfactants Changes Composition of Protein Corona. Biomacromolecules 19:2657-2664
Muraleedharan S, Freitas C, Mann P, Glatter T, Ringgaard S (2018) A cell length-dependent transition in MinD-dynamics promotes a switch in division-site placement and preservation of proliferating elongated Vibrio parahaemolyticus swarmer cells. Mol Microbiol (in press)
Murphy S, Zweyer M, Henry M, Meleady P, Mundegar RR, Swandulla D, Ohlendieck K (2018) Proteomic analysis of the sarcolemma-enriched fraction from dystrophic mdx-4cv skeletal muscle. J Proteomics (in press)
Murphy S, Zweyer M, Henry M, Meleady P, Mundegar RR, Swandulla D, Ohlendieck K (2018) Proteomic profiling of liver tissue from the mdx-4cv mouse model of Duchenne muscular dystrophy. Clin Proteomics 15:34
Murray CE, Gami-Patel P, Gkanatsiou E, Brinkmalm G, Portelius E, Wirths O, Heywood W, Blennow K, Ghiso J, Holton JL, Mills K, Zetterberg H, Revesz T, Lashley T (2018) The presubiculum is preserved from neurodegenerative changes in Alzheimer's disease. Acta Neuropathol Commun 6:62
Nakamura H, Takahashi-Jitsuki A, Makihara H, Asano T, Kimura Y, Nakabayashi J, Yamashita N, Kawamoto Y, Nakamura F, Ohshima T, Hirano H, Tanaka F, Goshima Y (2018) Proteome and behavioral alterations in phosphorylation-deficient mutant Collapsin Response Mediator Protein2 knock-in mice. Neurochem Int (in press)
Nazar RN, Xu X, Shittu H, Kurosky A, Robb J (2018) Tomato Ve resistance locus; defense or growth. Planta 247:1339-1350
Nazar RN, Xu X, Kurosky A, Robb J (2018) Antagonistic function of the Ve R-genes in tomato. Plant Mol Biol 98:67-79
Novakova S, Danchenko M, Skultety L, Fialova I, Leskova A, Beke G, Flores-Ramirez G, Glasa M (2018) Photosynthetic and Stress Responsive Proteins Are Altered More Effectively in Nicotiana benthamiana Infected with Plum pox virus Aggressive PPV-CR versus Mild PPV-C Cherry-Adapted Isolates. J Proteome Res 17:3114-3127
Novy K, Kilcher S, Omasits U, Bleck CKE, Beerli C, Vowinckel J, Martin CK, Syedbasha M, Maiolica A, White I, Mercer J, Wollscheid B (2018) Proteotype profiling unmasks a viral signalling network essential for poxvirus assembly and transcriptional competence. Nat Microbiol 3:588-599
Nunes PT, Gomez-Mendoza DP, Rezende CP, Figueiredo HCP, Ribeiro AM (2018) Thalamic Proteome Changes and Behavioral Impairments in Thiamine-deficient Rats. Neuroscience 385:181-197
Nybo T, Cai H, Chuang CY, Gamon LF, Rogowska-Wrzesinska A, Davies MJ (2018) Chlorination and oxidation of human plasma fibronectin by myeloperoxidase-derived oxidants, and its consequences for smooth muscle cell function. Redox Biol 19:388-400
Nybo T, Dieterich S, Gamon LF, Chuang CY, Hammer A, Hoefler G, Malle E, Rogowska-Wrzesinska A, Davies MJ (2018) Chlorination and oxidation of the extracellular matrix protein laminin and basement membrane extracts by hypochlorous acid and myeloperoxidase. Redox Biol 20:496-513
Ohman T, Tamene F, Goos H, Loukovaara S, Varjosalo M (2018) Systems pathology analysis identifies neurodegenerative nature of age-related vitreoretinal interface diseases. Aging Cell:e12809
Oppong-Danquah E, Parrot D, Blumel M, Labes A, Tasdemir D (2018) Molecular Networking-Based Metabolome and Bioactivity Analyses of Marine-Adapted Fungi Co-cultivated With Phytopathogens. Front Microbiol 9:2072
Padilla CA, Barcena JA, Lopez-Grueso MJ, Requejo-Aguilar R (2018) The regulation of TORC1 pathway by the yeast chaperones Hsp31 is mediated by SFP1 and affects proteasomal activity. Biochim Biophys Acta Gen Subj 1863:534-546
Paes W, Dowle A, Coldwell J, Leech A, Ganderton T, Brzozowski A (2018) The Chlamydia trachomatis PmpD adhesin forms higher order structures through disulphide-mediated covalent interactions. PLoS One 13:e0198662
Palomo L, Mleczko JE, Azkargorta M, Conde-Vancells J, Gonzalez E, Elortza F, Royo F, Falcon-Perez JM (2018) Abundance of Cytochromes in Hepatic Extracellular Vesicles Is Altered by Drugs Related With Drug-Induced Liver Injury. Hepatol Commun 2:1064-1079
Pao PJ, Emri E, Abdirahman SB, Soorma T, Zeng HH, Hauck SM, Thompson RB, Lengyel I (2018) The effects of zinc supplementation on primary human retinal pigment epithelium. J Trace Elem Med Biol 49:184-191
Parsley NC, Kirkpatrick CL, Crittenden CM, Rad JG, Hoskin DW, Brodbelt JS, Hicks LM (2018) PepSAVI-MS reveals anticancer and antifungal cycloviolacins in Viola odorata. Phytochemistry 152:61-70
Pastor Y, Camacho AI, Zuniga-Ripa A, Merchan A, Rosas P, Irache JM, Gamazo C (2018) Towards a subunit vaccine from a Shigella flexneri DeltatolR mutant. Vaccine 36:7509-7519
Peng M, He Q, Li S, Li L, Ma H (2018) Integrated analysis of proteomics-delineated and metabolomics-delineated hepatic metabolic responses to (-)-hydroxycitric acid in chick embryos. J Cell Biochem (in press)
Phan V, Cox D, Cipriani S, Spendiff S, Buchkremer S, O'Connor E, Horvath R, Goebel HH, Hathazi D, Lochmuller H, Straka T, Rudolf R, Weis J, Roos A (2018) SIL1 deficiency causes degenerative changes of peripheral nerves and neuromuscular junctions in fish, mice and human. Neurobiol Dis 124:218-229
Phan V, Schmidt J, Matyash V, Malchow S, Thanisch M, Lorenz C, Diepolder I, Schulz JB, Stenzel W, Roos A, Gess B (2018) Characterization of Naive and Vitamin C-Treated Mouse Schwann Cell Line MSC80: Induction of the Antioxidative Thioredoxin Related Transmembrane Protein 1. J Proteome Res 17:2925-2936
Piazza I, Kochanowski K, Cappelletti V, Fuhrer T, Noor E, Sauer U, Picotti P (2018) A Map of Protein-Metabolite Interactions Reveals Principles of Chemical Communication. Cell 172:358-372 e323
Piltti J, Bygdell J, Qu C, Lammi MJ (2018) Effects of long-term low oxygen tension in human chondrosarcoma cells. J Cell Biochem 119:2320-2332
Planckaert S, Jourdan S, Francis IM, Deflandre B, Rigali S, Devreese B (2018) Proteomic Response to Thaxtomin Phytotoxin Elicitor Cellobiose and to Deletion of Cellulose Utilization Regulator CebR in Streptomyces scabies. J Proteome Res 17:3837-3852
Poleti MD, Regitano LCA, Souza G, Cesar ASM, Simas RC, Silva-Vignato B, Oliveira GB, Andrade SCS, Cameron LC, Coutinho LL (2018) Longissimus dorsi muscle label-free quantitative proteomic reveals biological mechanisms associated with intramuscular fat deposition. J Proteomics 179:30-41
Pontremoli M, Brioschi M, Baetta R, Ghilardi S, Banfi C (2018) Identification of DKK-1 as a novel mediator of statin effects in human endothelial cells. Sci Rep 8:16671
Prieto-Bonete G, Perez-Carceles MD, Maurandi-Lopez A, Perez-Martinez C, Luna A (2018) Association between protein profile and postmortem interval in human bone remains. J Proteomics 192:54-63
Pringels L, Broeckx V, Boonen K, Landuyt B, Schoofs L (2018) Abundant plasma protein depletion using ammonium sulfate precipitation and Protein A affinity chromatography. J Chromatogr B Analyt Technol Biomed Life Sci 1089:43-59
Procopio N, Chamberlain AT, Buckley M (2018) Exploring Biological and Geological Age-related Changes through Variations in Intra- and Intertooth Proteomes of Ancient Dentine. J Proteome Res 17:1000-1013
Procopio N, Williams A, Chamberlain AT, Buckley M (2018) Forensic proteomics for the evaluation of the post-mortem decay in bones. J Proteomics 177:21-30
Puente-Marin S, Nombela I, Chico V, Ciordia S, Mena MC, Coll J, Mercado L, Ortega-Villaizan MDM (2018) Rainbow Trout Erythrocytes ex vivo Transfection With a DNA Vaccine Encoding VHSV Glycoprotein G Induces an Antiviral Immune Response. Front Immunol 9:2477
Qin W, Du Z, Gao Y (2018) Collection and preservation of urinary proteins, using a fluff pulp diaper. Sci China Life Sci 61:671-674
Raimondo F, Chinello C, Stella M, Santorelli L, Magni F, Pitto M (2018) Effects of Hematuria on the Proteomic Profile of Urinary Extracellular Vesicles: Technical Challenges. J Proteome Res 17:2572-2580
Ramirez-Tejero JA, Martinez-Lara E, Rus A, Camacho MV, Del Moral ML, Siles E (2018) Insight into the biological pathways underlying fibromyalgia by a proteomic approach. J Proteomics 186:47-55
Renthal R, Lohmeyer K, Borges LMF, Perez de Leon AA (2018) Surface lipidome of the lone star tick, Amblyomma americanum, provides leads on semiochemicals and lipid metabolism. Ticks Tick Borne Dis 10:138-145
Ribeiro DG, de Almeida RF, Fontes W, de Souza Castro M, de Sousa MV, Ricart CAO, da Cunha RNV, Lopes R, Scherwinski-Pereira JE, Mehta A (2018) Stress and cell cycle regulation during somatic embryogenesis plays a key role in oil palm callus development. J Proteomics 192:137-146
Rockenbach MF, Correa CCG, Heringer AS, Freitas ILJ, Santa-Catarina C, do Amaral-Junior AT, Silveira V (2018) Differentially abundant proteins associated with heterosis in the primary roots of popcorn. PLoS One 13:e0197114
Romero-Gavilan F, Araujo-Gomes N, Garcia-Arnaez I, Martinez-Ramos C, Elortza F, Azkargorta M, Iloro I, Gurruchaga M, Suay J, Goni I (2018) The effect of strontium incorporation into sol-gel biomaterials on their protein adsorption and cell interactions. Colloids Surf B Biointerfaces 174:9-16
Romero-Gavilan F, Araujo-Gomes N, Sanchez-Perez AM, Garcia-Arnaez I, Elortza F, Azkargorta M, de Llano JJM, Carda C, Gurruchaga M, Suay J, Goni I (2018) Bioactive potential of silica coatings and its effect on the adhesion of proteins to titanium implants. Colloids Surf B Biointerfaces 162:316-325
Roy M, Sorokina O, Skene N, Simonnet C, Mazzo F, Zwart R, Sher E, Smith C, Armstrong JD, Grant SGN (2018) Proteomic analysis of postsynaptic proteins in regions of the human neocortex. Nat Neurosci 21:130-138
Ruland T, Wolbert J, Gottschalk MG, Konig S, Schulte-Mecklenbeck A, Minnerup J, Meuth SG, Gross CC, Wiendl H, Meyer Zu Horste G (2018) Cerebrospinal Fluid Concentrations of Neuronal Proteins Are Reduced in Primary Angiitis of the Central Nervous System. Front Neurol 9:407
Ruzafa N, Pereiro X, Lepper MF, Hauck SM, Vecino E (2018) A Proteomics Approach to Identify Candidate Proteins Secreted by Muller Glia that Protect Ganglion Cells in the Retina. Proteomics 18:e1700321
Sayd T, Dufour C, Chambon C, Buffiere C, Remond D, Sante-Lhoutellier V (2018) Combined in vivo and in silico approaches for predicting the release of bioactive peptides from meat digestion. Food Chem 249:111-118
Schauer M, Kleinwort KJH, Degroote RL, Wiedemann C, Kremmer E, Hauck SM, Deeg CA (2018) Interaction of septin 7 and DOCK8 in equine lymphocytes reveals novel insights into signaling pathways associated with autoimmunity. Sci Rep 8:12332
Schlegel I, Renz P, Simon J, Lieberwirth I, Pektor S, Bausbacher N, Miederer M, Mailander V, Munoz-Espi R, Crespy D, Landfester K (2018) Highly Loaded Semipermeable Nanocapsules for Magnetic Resonance Imaging. Macromol Biosci 18:e1700387
Schott K, Fuchs NV, Derua R, Mahboubi B, Schnellbacher E, Seifried J, Tondera C, Schmitz H, Shepard C, Brandariz-Nunez A, Diaz-Griffero F, Reuter A, Kim B, Janssens V, Konig R (2018) Dephosphorylation of the HIV-1 restriction factor SAMHD1 is mediated by PP2A-B55alpha holoenzymes during mitotic exit. Nat Commun 9:2227
Seaton DD, Graf A, Baerenfaller K, Stitt M, Millar AJ, Gruissem W (2018) Photoperiodic control of the Arabidopsis proteome reveals a translational coincidence mechanism. Mol Syst Biol 14:e7962
Sepil I, Hopkins BR, Dean R, Thezenas ML, Charles PD, Konietzny R, Fischer R, Kessler B, Wigby S (2018) Quantitative proteomics identification of seminal fluid proteins in male Drosophila melanogaster. Mol Cell Proteomics (in press)
Severi L, Losi L, Fonda S, Taddia L, Gozzi G, Marverti G, Magni F, Chinello C, Stella M, Sheouli J, Braicu EI, Genovese F, Lauriola A, Marraccini C, Gualandi A, D'Arca D, Ferrari S, Costi MP (2018) Proteomic and Bioinformatic Studies for the Characterization of Response to Pemetrexed in Platinum Drug Resistant Ovarian Cancer. Front Pharmacol 9:454
Sheng Y, Capri J, Waring A, Valentine JS, Whitelegge J (2018) Exposure of Solvent-Inaccessible Regions in the Amyloidogenic Protein Human SOD1 Determined by Hydroxyl Radical Footprinting. J Am Soc Mass Spectrom (in press)
Sherman DL, Brophy PJ (2018) A murine model of Charcot-Marie-Tooth disease 4F reveals a role for the C-terminus of periaxin in the formation and stabilization of Cajal bands. Wellcome Open Res 3:20
Shorrock HK, van der Hoorn D, Boyd PJ, Llavero Hurtado M, Lamont DJ, Wirth B, Sleigh JN, Schiavo G, Wishart TM, Groen EJN, Gillingwater TH (2018) UBA1/GARS-dependent pathways drive sensory-motor connectivity defects in spinal muscular atrophy. Brain 141:2878-2894
Silva WM, Sousa CS, Oliveira LC, Soares SC, Souza G, Tavares GC, Resende CP, Folador EL, Pereira FL, Figueiredo H, Azevedo V (2018) Comparative proteomic analysis of four biotechnological strains Lactococcus lactis through label-free quantitative proteomics. Microb Biotechnol (in press)
Smith J, Davey G, Polom K, Roviello F, Bones J (2018) Mining the acidic serum proteome utilizing off-gel isoelectric focusing and label free quantitative liquid chromatography mass spectrometry. J Chromatogr A 1566:32-43
Smits K, Willems S, Van Steendam K, Van De Velde M, De Lange V, Ververs C, Roels K, Govaere J, Van Nieuwerburgh F, Peelman L, Deforce D, Van Soom A (2018) Proteins involved in embryo-maternal interaction around the signalling of maternal recognition of pregnancy in the horse. Sci Rep 8:5249
Soares ALC, Geilfus CM, Carpentier SC (2018) Genotype-Specific Growth and Proteomic Responses of Maize Toward Salt Stress. Front Plant Sci 9:661
Sobotzki N, Schafroth MA, Rudnicka A, Koetemann A, Marty F, Goetze S, Yamauchi Y, Carreira EM, Wollscheid B (2018) HATRIC-based identification of receptors for orphan ligands. Nat Commun 9:1519
Soethoudt M, Alachouzos G, van Rooden EJ, Moya-Garzon MD, van den Berg R, Heitman LH, van der Stelt M (2018) Development of a Cannabinoid-Based Photoaffinity Probe to Determine the Delta(8/9)-Tetrahydrocannabinol Protein Interaction Landscape in Neuroblastoma Cells. Cannabis Cannabinoid Res 3:136-151
Sopena-Torres S, Jorda L, Sanchez-Rodriguez C, Miedes E, Escudero V, Swami S, Lopez G, Pislewska-Bednarek M, Lassowskat I, Lee J, Gu Y, Haigis S, Alexander D, Pattathil S, Munoz-Barrios A, Bednarek P, Somerville S, Schulze-Lefert P, Hahn MG, Scheel D, Molina A (2018) YODA MAP3K kinase regulates plant immune responses conferring broad-spectrum disease resistance. New Phytol 218:661-680
Soto-Acosta R, Xie X, Shan C, Baker CK, Shi PY, Rossi SL, Garcia-Blanco MA, Bradrick S (2018) Fragile X mental retardation protein is a Zika virus restriction factor that is antagonized by subgenomic flaviviral RNA. Elife 7
Steevensz A, Gombar R, Vergilino R, Cristescu ME, Vacratsis PO (2018) Proteomic Profile of Daphnia pulex using Data-Independent Acquisition Mass Spectrometry and Ion Mobility Separation. Proteomics:e1700460
Stegmann M, Barclay AN, Metcalfe C (2018) Reduction of leucocyte cell surface disulfide bonds during immune activation is dynamic as revealed by a quantitative proteomics workflow (SH-IQ). Open Biol 8
Steindor M, Nkwouano V, Stefanski A, Stuehler K, Ioerger TR, Bogumil D, Jacobsen M, Mackenzie CR, Kalscheuer R (2018) A proteomics approach for the identification of species-specific immunogenic proteins in the Mycobacterium abscessus complex. Microbes Infect (in press)
Stella M, Chinello C, Cazzaniga A, Smith A, Galli M, Piga I, Grasso A, Grasso M, Del Puppo M, Varallo M, Bovo G, Magni F (2019) Histology-guided proteomic analysis to investigate the molecular profiles of clear cell Renal Cell Carcinoma grades. J Proteomics 191:38-47
Stuhr M, Blank-Landeshammer B, Reymond CE, Kollipara L, Sickmann A, Kucera M, Westphal H (2018) Disentangling thermal stress responses in a reef-calcifier and its photosymbionts by shotgun proteomics. Sci Rep 8:3524
Subramanian V, Borchard S, Azimzadeh O, Sievert W, Merl-Pham J, Mancuso M, Pasquali E, Multhoff G, Popper B, Zischka H, Atkinson MJ, Tapio S (2018) PPARalpha Is Necessary for Radiation-Induced Activation of Noncanonical TGFbeta Signaling in the Heart. J Proteome Res 17:1677-1689
Sung Y, Spagou K, Kafeza M, Kyriakides M, Dharmarajah B, Shalhoub J, Diaz JA, Wakefield TW, Holmes E, Davies AH (2018) Deep Vein Thrombosis Exhibits Characteristic Serum and Vein Wall Metabolic Phenotypes in the Inferior Vena Cava Ligation Mouse Model. Eur J Vasc Endovasc Surg 55:703-713
Suo B, Yang H, Wang Y, Lv H, Li Z, Xu C, Ai Z (2018) Comparative Proteomic and Morphological Change Analyses of Staphylococcus aureus During Resuscitation From Prolonged Freezing. Front Microbiol 9:866
Swart C, Martinez-Jaime S, Gorka M, Zander K, Graf A (2018) Hit-Gel: Streamlining in-gel protein digestion for high-throughput proteomics experiments. Sci Rep 8:8582
Szelechowski M, Amoedo N, Obre E, Leger C, Allard L, Bonneu M, Claverol S, Lacombe D, Oliet S, Chevallier S, Le Masson G, Rossignol R (2018) Metabolic Reprogramming in Amyotrophic Lateral Sclerosis. Sci Rep 8:3953
Taheri N, Mahmud A, Sandblad L, Fallman M, Wai SN, Fahlgren A (2018) Campylobacter jejuni bile exposure influences outer membrane vesicles protein content and bacterial interaction with epithelial cells. Sci Rep 8:16996
Tavares GC, Carvalho AF, Pereira FL, Rezende CP, Azevedo VAC, Leal CAG, Figueiredo HCP (2018) Transcriptome and Proteome of Fish-Pathogenic Streptococcus agalactiae Are Modulated by Temperature. Front Microbiol 9:2639
Telot L, Rousseau E, Lesuisse E, Garcia C, Morlet B, Leger T, Camadro JM, Serre V (2018) Quantitative proteomics in Friedreich's ataxia B-lymphocytes: A valuable approach to decipher the biochemical events responsible for pathogenesis. Biochim Biophys Acta 1864:997-1009
Theron L, Sayd T, Chambon C, Venien A, Viala D, Astruc T, Vautier A, Sante-Lhoutellier V (2018) Deciphering PSE-like muscle defect in cooked hams: A signature from the tissue to the molecular scale. Food Chem 270:359-366
Thompson AG, Gray E, Mager I, Fischer R, Thezenas ML, Charles PD, Talbot K, El Andaloussi S, Kessler BM, Wood M, Turner MR (2018) UFLC-Derived CSF Extracellular Vesicle Origin and Proteome. Proteomics 18:e1800257
Torregrossa MM, MacDonald M, Stone KL, Lam TT, Nairn AC, Taylor JR (2018) Phosphoproteomic analysis of cocaine memory extinction and reconsolidation in the nucleus accumbens. Psychopharmacology (Berl) (in press)
Tse SPK, Beauchemin M, Morse D, Lo SCL (2018) Refining Transcriptome Gene Catalogs by MS-Validation of Expressed Proteins. Proteomics 18 (in press)
Tuhkuri A, Saraswat M, Makitie A, Mattila P, Silen R, Dickinson A, Carpen T, Tohmola T, Joenvaara S, Renkonen S (2018) Patients with early-stage oropharyngeal cancer can be identified with label-free serum proteomics. Br J Cancer 119:200-212
Turapov O, Forti F, Kadhim B, Ghisotti D, Sassine J, Straatman-Iwanowska A, Bottrill AR, Moynihan PJ, Wallis R, Barthe P, Cohen-Gonsaud M, Ajuh P, Vollmer W, Mukamolova GV (2018) Two Faces of CwlM, an Essential PknB Substrate, in Mycobacterium tuberculosis. Cell Rep 25:57-67 e55
Turlo AJ, Ashraf Kharaz Y, Clegg PD, Anderson J, Peffers MJ (2018) Donor age affects proteome composition of tenocyte-derived engineered tendon. BMC Biotechnol 18:2
Umadevi P, Soumya M, George JK, Anandaraj M (2018) Proteomics assisted profiling of antimicrobial peptide signatures from black pepper (Piper nigrum L.). Physiol Mol Biol Plants 24:379-387
Valko PO, Roschitzki B, Faigle W, Grossmann J, Panse C, Biro P, Dambach M, Spahn DR, Weller M, Martin R, Baumann CR (2018) In search of cerebrospinal fluid biomarkers of fatigue in multiple sclerosis: A proteomics study. J Sleep Res:e12721
Van Bael S, Zels S, Boonen K, Beets I, Schoofs L, Temmerman L (2018) A Caenorhabditis elegans Mass Spectrometric Resource for Neuropeptidomics. J Am Soc Mass Spectrom 29:879-889
van Mierlo G, Dirks RAM, De Clerck L, Brinkman AB, Huth M, Kloet SL, Saksouk N, Kroeze LI, Willems S, Farlik M, Bock C, Jansen JH, Deforce D, Vermeulen M, Dejardin J, Dhaenens M, Marks H (2018) Integrative Proteomic Profiling Reveals PRC2-Dependent Epigenetic Crosstalk Maintains Ground-State Pluripotency. Cell Stem Cell 24:123-137 e128
van Wesemael J, Hueber Y, Kissel E, Campos N, Swennen R, Carpentier S (2018) Homeolog expression analysis in an allotriploid non-model crop via integration of transcriptomics and proteomics. Sci Rep 8:1353
Vetterli SU, Zerbe K, Muller M, Urfer M, Mondal M, Wang SY, Moehle K, Zerbe O, Vitale A, Pessi G, Eberl L, Wollscheid B, Robinson JA (2018) Thanatin targets the intermembrane protein complex required for lipopolysaccharide transport in Escherichia coli. Sci Adv 4:eaau2634
Viana AGA, Martins AMA, Pontes AH, Fontes W, Castro MS, Ricart CAO, Sousa MV, Kaya A, Topper E, Memili E, Moura AA (2018) Proteomic landscape of seminal plasma associated with dairy bull fertility. Sci Rep 8:16323
Victorio VCM, Souza G, Santos MCB, Vega AR, Cameron LC, Ferreira MSL (2018) Differential expression of albumins and globulins of wheat flours of different technological qualities revealed by nanoUPLC-UDMS(E). Food Chem 239:1027-1036
Volpato V, Smith J, Sandor C, Ried JS, Baud A, Handel A, Newey SE, et al. (2018) Reproducibility of Molecular Phenotypes after Long-Term Differentiation to Human iPSC-Derived Neurons: A Multi-Site Omics Study. Stem Cell Reports 11:897-911
Wang Y, Feng R, He C, Su H, Ma H, Wan JB (2018) An integrated strategy to improve data acquisition and metabolite identification by time-staggered ion lists in UHPLC/Q-TOF MS-based metabolomics. J Pharm Biomed Anal 157:171-179
Waniwan JT, Chen YJ, Capangpangan R, Weng SH (2018) Glycoproteomic Alterations in Drug-Resistant Nonsmall Cell Lung Cancer Cells Revealed by Lectin Magnetic Nanoprobe-Based Mass Spectrometry. J Proteome Res 17:3761-3773
Weber C, Simon J, Mailander V, Morsbach S, Landfester K (2018) Preservation of the soft protein corona in distinct flow allows identification of weakly bound proteins. Acta Biomater 76:217-224
Werth EG, McConnell EW, Couso Lianez I, Perrine Z, Crespo JL, Umen JG, Hicks LM (2018) Investigating the effect of target of rapamycin kinase inhibition on the Chlamydomonas reinhardtii phosphoproteome: from known homologs to new targets. New Phytol (in press)
Willforss J, Chawade A, Levander F (2018) NormalyzerDE: Online tool for improved normalization of omics expression data and high-sensitivity differential expression analysis. J Proteome Res (in press)
Witte H, Schreiner D, Scheiffele P (2018) A Sam68-dependent alternative splicing program shapes postsynaptic protein complexes. Eur J Neurosci (in press)
Witzel K, Matros A, Moller ALB, Ramireddy E, Finnie C, Peukert M, Rutten T, Herzog A, Kunze G, Melzer M, Kaspar-Schoenefeld S, Schmulling T, Svensson B, Mock HP (2018) Plasma membrane proteome analysis identifies a role of barley membrane steroid binding protein in root architecture response to salinity. Plant Cell Environ 41:1311-1330
Wong EHC, Dong YY, Coray M, Cortada M, Levano S, Schmidt A, Brand Y, Bodmer D, Muller L (2018) Inner ear exosomes and their potential use as biomarkers. PLoS One 13:e0198029
Wu AY, Ueda K, Lai CP (2018) Proteomic Analysis of Extracellular Vesicles for Cancer Diagnostics. Proteomics:e1800162
Yakhine-Diop SMS, Rodriguez-Arribas M, Martinez-Chacon G, Uribe-Carretero E, Gomez-Sanchez R, Aiastui A, Lopez de Munain A, Bravo-San Pedro JM, Niso-Santano M, Gonzalez-Polo RA, Fuentes JM (2018) Acetylome in Human Fibroblasts From Parkinson's Disease Patients. Front Cell Neurosci 12:97
Yamamoto R, Kawahara M, Ito S, Satoh J, Tatsumi G, Hishizawa M, Suzuki T, Andoh A (2018) Selective dissociation between LSD1 and GFI1B by a LSD1 inhibitor NCD38 induces the activation of ERG super-enhancer in erythroleukemia cells. Oncotarget 9:21007-21021
Yao Z, Jia X, Megger DA, Chen J, Liu Y, Li J, Sitek B, Yuan Z (2018) Label-free Proteomic Analysis of Exosomes Secreted from THP-1-derived Macrophages Treated with IFN-alpha Identifies Antiviral Proteins Enriched in Exosomes. J Proteome Res (in press)
Ye M, Veyrat N, Xu H, Hu L, Turlings TCJ, Erb M (2018) An herbivore-induced plant volatile reduces parasitoid attraction by changing the smell of caterpillars. Sci Adv 4:eaar4767
Yoon C, Song H, Yin T, Bausch-Fluck D, Frei AP, Kattman S, Dubois N, Witty AD, Hewel JA, Guo H, Emili A, Wollscheid B, Keller G, Zandstra PW (2018) FZD4 Marks Lateral Plate Mesoderm and Signals with NORRIN to Increase Cardiomyocyte Induction from Pluripotent Stem Cell-Derived Cardiac Progenitors. Stem Cell Reports 10:87-100
Yuzhalin AE, Gordon-Weeks AN, Tognoli ML, Jones K, Markelc B, Konietzny R, Fischer R, Muth A, O'Neill E, Thompson PR, Venables PJ, Kessler BM, Lim SY, Muschel RJ (2018) Colorectal cancer liver metastatic growth depends on PAD4-driven citrullination of the extracellular matrix. Nat Commun 9:4783
Zhang H, Jiang JM, Zheng D, Yuan M, Wang ZY, Zhang HM, Zheng CW, Xiao LB, Xu HX (2018) A multidimensional analytical approach based on time-decoupled online comprehensive two-dimensional liquid chromatography coupled with ion mobility quadrupole time-of-flight mass spectrometry for the analysis of ginsenosides from white and red ginsengs. J Pharm Biomed Anal 163:24-33
Zhang L, Li Y, Gao Y (2018) Early changes in the urine proteome in a diethyldithiocarbamate-induced chronic pancreatitis rat model. J Proteomics 186:8-14
Zhao M, Yang Y, Guo Z, Shao C, Sun H, Zhang Y, Sun Y, Liu Y, Song Y, Zhang L, Li Q, Liu J, Li M, Gao Y, Sun W (2018) A Comparative Proteomics Analysis of Five Body Fluids: Plasma, Urine, Cerebrospinal Fluid, Amniotic Fluid, and Saliva. Proteomics Clin Appl:e1800008
Zufferey F, Rahban R, Garcia A, Gagnebin Y, Boccard J, Tonoli D, Jeanneret F, Stettler E, Senn A, Nef S, Rudaz S, Rossier MF (2018) Steroid profiles in both blood serum and seminal plasma are not correlated and do not reflect sperm quality: Study on the male reproductive health of fifty young Swiss men. Clin Biochem 62:39-46
2017
Abreu TF, Sumitomo BN, Nishiyama MY, Jr., Oliveira UC, Souza GH, Kitano ES, Zelanis A, Serrano SM, Junqueira-de-Azevedo I, Silva PI, Jr., Tashima AK (2017) Peptidomics of Acanthoscurria gomesiana spider venom reveals new toxins with potential antimicrobial activity. J Proteomics 151:232-242
Al Shweiki MR, Monchgesang S, Majovsky P, Thieme D, Trutschel D, Hoehenwarter W (2017) Assessment of Label-Free Quantification in Discovery Proteomics and Impact of Technological Factors and Natural Variability of Protein Abundance. J Proteome Res 16:1410-1424
Al-Bajalan MMM, Xia D, Armstrong S, Randle N, Wastling JM (2017) Toxoplasma gondii and Neospora caninum induce different host cell responses at proteome-wide phosphorylation events; a step forward for uncovering the biological differences between these closely related parasites. Parasitol Res 116:2707-2719
Al-Idrus A, Carpentier SC, Ahmad MT, Panis B, Mohamed Z (2017) Elucidation of the compatible interaction between banana and Meloidogyne incognita via high-throughput proteome profiling. PLoS One 12:e0178438
Al-Mayyahi RS, Sterio LD, Connolly JB, Adams CF, Al-Tumah WA, Sen J, Emes RD, Hart SR, Chari DM (2017) A proteomic investigation into mechanisms underpinning corticosteroid effects on neural stem cells. Mol Cell Neurosci 86:30-40
Almeida AM, Nanni P, Ferreira AM, Fortes C, Grossmann J, Bessa RJ, Costa P (2017) The longissimus thoracis muscle proteome in Alentejana bulls as affected by growth path. J Proteomics 152:206-215
Almozyan S, Colak D, Mansour F, Alaiya A, Al-Harazi O, Qattan A, Al-Mohanna F, Al-Alwan M, Ghebeh H (2017) PD-L1 promotes OCT4 and Nanog expression in breast cancer stem cells by sustaining PI3K/AKT pathway activation. Int J Cancer 141:1402-1412
Araujo-Gomes N, Romero-Gavilan F, Sanchez-Perez AM, Gurruchaga M, Azkargorta M, Elortza F, Martinez-Ibanez M, Iloro I, Suay J, Goni I (2017) Characterization of serum proteins attached to distinct sol-gel hybrid surfaces. J Biomed Mater Res B Appl Biomater (In press)
Arbelaiz A, Azkargorta M, Krawczyk M, Santos-Laso A, Lapitz A, Perugorria MJ, Erice O, Gonzalez E, Jimenez-Aguero R, Lacasta A, Ibarra C, Sanchez-Campos A, Jimeno JP, Lammert F, Milkiewicz P, Marzioni M, Macias RIR, Marin JJG, Patel T, Gores GJ, Martinez I, Elortza F, Falcon-Perez JM, Bujanda L, Banales JM (2017) Serum extracellular vesicles contain protein biomarkers for primary sclerosing cholangitis and cholangiocarcinoma. Hepatology 66:1125-1143
Asimakopoulou A, Fulop A, Borkham-Kamphorst E, de Leur EV, Gassler N, Berger T, Beine B, Meyer HE, Mak TW, Hopf C, Henkel C, Weiskirchen R (2017) Altered mitochondrial and peroxisomal integrity in lipocalin-2-deficient mice with hepatic steatosis. Biochim Biophys Acta 1863:2093-2110
Bailey J, Shaw A, Fischer R, Ryan BJ, Kessler BM, McCullagh J, Wade-Martins R, Channon KM, Crabtree MJ (2017) A novel role for endothelial tetrahydrobiopterin in mitochondrial redox balance. Free Radic Biol Med 104:214-225
Bao K, Bostanci N, Thurnheer T, Belibasakis GN (2017) Proteomic shifts in multi-species oral biofilms caused by Anaeroglobus geminatus. Sci Rep 7:4409
Beker MC, Caglayan B, Yalcin E, Caglayan AB, Turkseven S, Gurel B, Kelestemur T, Sertel E, Sahin Z, Kutlu S, Kilic U, Baykal AT, Kilic E (2017) Time-of-Day Dependent Neuronal Injury After Ischemic Stroke: Implication of Circadian Clock Transcriptional Factor Bmal1 and Survival Kinase AKT. Mol Neurobiol (In press)
Bi B, Li F, Guo J, Li C, Jing R, Lv X, Chen X, Wang F, Azadzoi KM, Wang L, Liu Y, Yang JH (2017) Label-free quantitative proteomics unravels the importance of RNA processing in glioma malignancy. Neuroscience 351:84-95
Birot A, Eguienta K, Vazquez S, Claverol S, Bonneu M, Ekwall K, Javerzat JP, Vaur S (2017) A second Wpl1 anti-cohesion pathway requires dephosphorylation of fission yeast kleisin Rad21 by PP4. Embo J 36:1364-1378
Bjorkum AA, Oveland E, Stuhr L, Havnes MB, Berven F, Gronning M, Hope A (2017) Fast hyperbaric decompression after heliox saturation altered the brain proteome in rats. PLoS One 12:e0185765
Bottini S, Hamouda-Tekaya N, Mategot R, Zaragosi LE, Audebert S, Pisano S, Grandjean V, Mauduit C, Benahmed M, Barbry P, Repetto E, Trabucchi M (2017) Post-transcriptional gene silencing mediated by microRNAs is controlled by nucleoplasmic Sfpq. Nat Commun 8:1189
Boukpessi T, Hoac B, Coyac BR, Leger T, Garcia C, Wicart P, Whyte MP, Glorieux FH, Linglart A, Chaussain C, McKee MD (2017) Osteopontin and the dento-osseous pathobiology of X-linked hypophosphatemia. Bone 95:151-161
Brackmann M, Wang J, Basler M (2017) Type VI secretion system sheath inter-subunit interactions modulate its contraction. EMBO Rep (In press)
Brocard L, Immel F, Coulon D, Esnay N, Tuphile K, Pascal S, Claverol S, Fouillen L, Bessoule JJ, Brehelin C (2017) Proteomic Analysis of Lipid Droplets from Arabidopsis Aging Leaves Brings New Insight into Their Biogenesis and Functions. Front Plant Sci 8:894
Bromilow S, Gethings LA, Buckley M, Bromley M, Shewry PR, Langridge JI, Clare Mills EN (2017) A curated gluten protein sequence database to support development of proteomics methods for determination of gluten in gluten-free foods. J Proteomics 163:67-75
Bromilow SN, Gethings LA, Langridge JI, Shewry PR, Buckley M, Bromley MJ, Mills EN (2017) Comprehensive Proteomic Profiling of Wheat Gluten Using a Combination of Data-Independent and Data-Dependent Acquisition. Front Plant Sci 7:2020
Caillot N, Bouley J, Jain K, Mariano S, Luce S, Horiot S, Airouche S, Beuraud C, Beauvallet C, Devillier P, Chollet-Martin S, Kellenberger C, Mascarell L, Chabre H, Batard T, Nony E, Lombardi V, Baron-Bodo V, Moingeon P (2017) Sialylated Fetuin-A as a candidate predictive biomarker for successful grass pollen allergen immunotherapy. J Allergy Clin Immunol 140:759-770 e713
Caraveo G, Soste M, Cappelleti V, Fanning S, van Rossum DB, Whitesell L, Huang Y, Chung CY, Baru V, Zaichick S, Picotti P, Lindquist S (2017) FKBP12 contributes to alpha-synuclein toxicity by regulating the calcineurin-dependent phosphoproteome. Proc Natl Acad Sci U S A 114:E11313-E11322
Casanovas A, Pinto-Llorente R, Carrascal M, Abian J (2017) Large-Scale Filter-Aided Sample Preparation Method for the Analysis of the Ubiquitinome. Anal Chem 89:3840-3846
Cases O, Obry A, Ben-Yacoub S, Augustin S, Joseph A, Toutirais G, Simonutti M, Christ A, Cosette P, Kozyraki R (2017) Impaired vitreous composition and retinal pigment epithelium function in the FoxG1::LRP2 myopic mice. Biochim Biophys Acta 1863:1242-1254
Catenaccio A, Llavero Hurtado M, Diaz P, Lamont DJ, Wishart TM, Court FA (2017) Molecular analysis of axonal-intrinsic and glial-associated co-regulation of axon degeneration. Cell Death Dis 8:e3166
Chang L, Ni J, Beretov J, Wasinger VC, Hao J, Bucci J, Malouf D, Gillatt D, Graham PH, Li Y (2017) Identification of protein biomarkers and signaling pathways associated with prostate cancer radioresistance using label-free LC-MS/MS proteomic approach. Sci Rep 7:41834
Chen H, Zhang Q, Qiao L, Fan X, Zhang W, Zhao W, Chen JJ (2017) Cdc6 contributes to abrogating the G1 checkpoint under hypoxic conditions in HPV E7 expressing cells. Sci Rep 7:2927
Clamens T, Rosay T, Crepin A, Grandjean T, Kentache T, Hardouin J, Bortolotti P, Neidig A, Mooij M, Hillion M, Vieillard J, Cosette P, Overhage J, O'Gara F, Bouffartigues E, Dufour A, Chevalier S, Guery B, Cornelis P, Feuilloley MG, Lesouhaitier O (2017) The aliphatic amidase AmiE is involved in regulation of Pseudomonas aeruginosa virulence. Sci Rep 7:41178
Correia S, Hebraud M, Chafsey I, Chambon C, Viala D, Saenz Y, Capelo JL, Poeta P, Igrejas G (2017) Comparative subproteomic analysis of clinically acquired fluoroquinolone resistance and ciprofloxacin stress in Salmonella Typhimurium DT104B. Proteomics Clin Appl 11(In press)
Crouzet M, Claverol S, Lomenech AM, Le Senechal C, Costaglioli P, Barthe C, Garbay B, Bonneu M, Vilain S (2017) Pseudomonas aeruginosa cells attached to a surface display a typical proteome early as 20 minutes of incubation. PLoS One 12:e0180341
Dakic V, Minardi Nascimento J, Costa Sartore R, Maciel RM, de Araujo DB, Ribeiro S, Martins-de-Souza D, Rehen SK (2017) Short term changes in the proteome of human cerebral organoids induced by 5-MeO-DMT. Sci Rep 7:12863
Davidson R, Gardner S, Jupp O, Bullough A, Butters S, Watts L, Donell S, Traka M, Saha S, Mithen R, Peffers M, Clegg P, Bao Y, Cassidy A, Clark I (2017) Isothiocyanates are detected in human synovial fluid following broccoli consumption and can affect the tissues of the knee joint. Sci Rep 7:3398
De Munter S, Gornemann J, Derua R, Lesage B, Qian J, Heroes E, Waelkens E, Van Eynde A, Beullens M, Bollen M (2017) Split-BioID: a proximity biotinylation assay for dimerization-dependent protein interactions. FEBS Lett 591:415-424
Demircan T, Keskin I, Dumlu SN, Ayturk N, Avsaroglu ME, Akgun E, Ozturk G, Baykal AT (2017) Detailed tail proteomic analysis of axolotl (Ambystoma mexicanum) using an mRNA-seq reference database. Proteomics 17 (In press)
Dezonne RS, Sartore RC, Nascimento JM, Saia-Cereda VM, Romao LF, Alves-Leon SV, de Souza JM, Martins-de-Souza D, Rehen SK, Gomes FC (2017) Derivation of Functional Human Astrocytes from Cerebral Organoids. Sci Rep 7:45091
Dhawi F, Datta R, Ramakrishna W (2017) Proteomics provides insights into biological pathways altered by plant growth promoting bacteria and arbuscular mycorrhiza in sorghum grown in marginal soil. Biochim Biophys Acta 1865:243-251
Dominguez-Martin MA, Gomez-Baena G, Diez J, Lopez-Grueso MJ, Beynon RJ, Garcia-Fernandez JM (2017) Quantitative Proteomics Shows Extensive Remodeling Induced by Nitrogen Limitation in Prochlorococcusmarinus SS120. mSystems 2
Dozio V, Sanchez JC (2017) Characterisation of extracellular vesicle-subsets derived from brain endothelial cells and analysis of their protein cargo modulation after TNF exposure. J Extracell Vesicles 6:1302705
Du Z, Chen Y, Li X (2017) Quantitative proteomic analyses of the microbial degradation of estrone under various background nitrogen and carbon conditions. Water Res 123:361-368
Eastlake K, Heywood WE, Tracey-White D, Aquino E, Bliss E, Vasta GR, Mills K, Khaw PT, Moosajee M, Limb GA (2017) Comparison of proteomic profiles in the zebrafish retina during experimental degeneration and regeneration. Sci Rep 7:44601
Einer C, Hohenester S, Wimmer R, Wottke L, Artmann R, Schulz S, Gosmann C, Simmons A, Leitzinger C, Eberhagen C, Borchard S, Schmitt S, Hauck SM, von Toerne C, Jastroch M, Walheim E, Rust C, Gerbes AL, Popper B, Mayr D, Schnurr M, Vollmar AM, Denk G, Zischka H (2017) Data on chow, liver tissue and mitochondrial fatty acid compositions as well as mitochondrial proteome changes after feeding mice a western diet for 6-24 weeks. Data Brief 15:163-169
El-Rami F, Nelson K, Xu P (2017) Proteomic Approach for Extracting Cytoplasmic Proteins from Streptococcus sanguinis using Mass Spectrometry. J Mol Biol Res 7:50-57
Emmens JE, Jones DJL, Cao TH, Chan DCS, Romaine SPR, Quinn PA, Anker SD, Cleland JG, Dickstein K, Filippatos G, Hillege HL, Lang CC, Ponikowski P, Samani NJ, van Veldhuisen DJ, Zannad F, Zwinderman AH, Metra M, de Boer RA, Voors AA, Ng LL (2017) Proteomic diversity of high-density lipoprotein explains its association with clinical outcome in patients with heart failure. Eur J Heart Fail (In press)
Fella E, Sokratous K, Papacharalambous R, Kyriacou K, Phillips J, Sanderson S, Panayiotou E, Kyriakides T (2017) Pharmacological Stimulation of Phagocytosis Enhances Amyloid Plaque Clearance; Evidence from a Transgenic Mouse Model of ATTR Neuropathy. Front Mol Neurosci 10:138
Fernando DM, Chong P, Singh M, Spicer V, Unger M, Loewen PC, Westmacott G, Kumar A (2017) Multi-omics approach to study global changes in a triclosan-resistant mutant strain of Acinetobacter baumannii ATCC 17978. Int J Antimicrob Agents 49:74-80
Fisunov GY, Evsyutina DV, Garanina IA, Arzamasov AA, Butenko IO, Altukhov IA, Nikitina AS, Govorun VM (2017) Ribosome profiling reveals an adaptation strategy of reduced bacterium to acute stress. Biochimie 132:66-74
Furrer R, Eisele PS, Schmidt A, Beer M, Handschin C (2017) Paracrine cross-talk between skeletal muscle and macrophages in exercise by PGC-1alpha-controlled BNP. Sci Rep 7:40789
Garcez PP, Nascimento JM, de Vasconcelos JM, Madeiro da Costa R, Delvecchio R, Trindade P, Loiola EC, Higa LM, Cassoli JS, Vitoria G, Sequeira PC, Sochacki J, Aguiar RS, Fuzii HT, de Filippis AM, da Silva Goncalves Vianez Junior JL, Tanuri A, Martins-de-Souza D, Rehen SK (2017) Zika virus disrupts molecular fingerprinting of human neurospheres. Sci Rep 7:40780
Gardiner M, Bournazos AM, Maturana-Martinez C, Zhong L, Egan S (2017) Exoproteome Analysis of the Seaweed Pathogen Nautella italica R11 Reveals Temperature-Dependent Regulation of RTX-Like Proteins. Front Microbiol 8:1203
Glass LN, Swapna G, Chavadi SS, Tufariello JM, Mi K, Drumm JE, Lam TT, Zhu G, Zhan C, Vilcheze C, Arcos J, Chen Y, Bi L, Mehta S, Porcelli SA, Almo SC, Yeh SR, Jacobs WR, Jr., Torrelles JB, Chan J (2017) Mycobacterium tuberculosis universal stress protein Rv2623 interacts with the putative ATP binding cassette (APC) transporter Rv1747 to regulate mycobacterial growth. PLoS Pathog 13:e1006515
Gombar R, Pitcher TE, Lewis JA, Auld J, Vacratsis PO (2017) Proteomic characterization of seminal plasma from alternative reproductive tactics of Chinook salmon (Oncorhynchus tswatchysha). J Proteomics 157:1-9
Gomez-Baena G, Bennett RJ, Martinez-Rodriguez C, Wnek M, Laing G, Hickey G, McLean L, Beynon RJ, Carrol ED (2017) Quantitative Proteomics of Cerebrospinal Fluid in Paediatric Pneumococcal Meningitis. Sci Rep 7:7042
Goss R, Greifenhagen A, Bergner J, Volke D, Hoffmann R, Wilhelm C, Schaller-Laudel S (2017) Direct isolation of a functional violaxanthin cycle domain from thylakoid membranes of higher plants. Planta 245:793-806
Govaert E, Van Steendam K, Willems S, Vossaert L, Dhaenens M, Deforce D (2017) Comparison of fractionation proteomics for local SWATH library building. Proteomics (In press)
Graham LC, Eaton SL, Brunton PJ, Atrih A, Smith C, Lamont DJ, Gillingwater TH, Pennetta G, Skehel P, Wishart TM (2017) Proteomic profiling of neuronal mitochondria reveals modulators of synaptic architecture. Mol Neurodegener 12:77
Grosserueschkamp F, Bracht T, Diehl HC, Kuepper C, Ahrens M, Kallenbach-Thieltges A, Mosig A, Eisenacher M, Marcus K, Behrens T, Bruning T, Theegarten D, Sitek B, Gerwert K (2017) Spatial and molecular resolution of diffuse malignant mesothelioma heterogeneity by integrating label-free FTIR imaging, laser capture microdissection and proteomics. Sci Rep 7:44829
Guerreiro JF, Mira NP, Santos AXS, Riezman H, Sa-Correia I (2017) Membrane Phosphoproteomics of Yeast Early Response to Acetic Acid: Role of Hrk1 Kinase and Lipid Biosynthetic Pathways, in Particular Sphingolipids. Front Microbiol 8:1302
Guo Z, Cheng J, Sun H, Sun W (2017) A qualitative and quantitative evaluation of the peptide characteristics of microwave- and ultrasound-assisted digestion in discovery and targeted proteomic analyses. Rapid Commun Mass Spectrom 31:1353-1362
Hakimi O, Ternette N, Murphy R, Kessler BM, Carr A (2017) A quantitative label-free analysis of the extracellular proteome of human supraspinatus tendon reveals damage to the pericellular and elastic fibre niches in torn and aged tissue. PLoS One 12:e0177656
Hammam K, Saez-Ayala M, Rebuffet E, Gros L, Lopez S, Hajem B, Humbert M, Baudelet E, Audebert S, Betzi S, Lugari A, Combes S, Letard S, Casteran N, Mansfield C, Moussy A, De Sepulveda P, Morelli X, Dubreuil P (2017) Dual protein kinase and nucleoside kinase modulators for rationally designed polypharmacology. Nat Commun 8:1420
Handke J, Procopio N, Buckley M, van der Meer D, Williams G, Carr M, Williams A (2017) Successive bacterial colonisation of pork and its implications for forensic investigations. Forensic Sci Int 281:1-8
Hanhart P, Thiess M, Amari K, Bajdzienko K, Giavalisco P, Heinlein M, Kehr J (2017) Bioinformatic and expression analysis of the Brassica napus L. cyclophilins. Sci Rep 7:1514
Hauck SM, Lepper MF, Hertl M, Sekundo W, Deeg CA (2017) Proteome dynamics in biobanked horse peripheral blood derived lymphocytes (PBL) with induced autoimmune uveitis. Proteomics (In press)
Hendgen-Cotta UB, Esfeld S, Coman C, Ahrends R, Klein-Hitpass L, Flogel U, Rassaf T, Totzeck M (2017) A novel physiological role for cardiac myoglobin in lipid metabolism. Sci Rep 7:43219
Henry M, Power M, Kaushik P, Coleman O, Clynes M, Meleady P (2017) Differential Phosphoproteomic Analysis of Recombinant Chinese Hamster Ovary Cells Following Temperature Shift. J Proteome Res 16:2339-2358
Heyer R, Schallert K, Zoun R, Becher B, Saake G, Benndorf D (2017) Challenges and perspectives of metaproteomic data analysis. J Biotechnol 261:24-36
Hmmier A, O'Brien ME, Lynch V, Clynes M, Morgan R, Dowling P (2017) Proteomic analysis of bronchoalveolar lavage fluid (BALF) from lung cancer patients using label-free mass spectrometry. BBA Clin 7:97-104
Horcajo P, Xia D, Randle N, Collantes-Fernandez E, Wastling J, Ortega-Mora LM, Regidor-Cerrillo J (2017) Integrative transcriptome and proteome analyses define marked differences between Neospora caninum isolates throughout the tachyzoite lytic cycle. J Proteomics (In press)
Huang BX, Newcomer K, Kevala K, Barnaeva E, Zheng W, Hu X, Patnaik S, Southall N, Marugan J, Ferrer M, Kim HY (2017) Identification of 4-phenylquinolin-2(1H)-one as a specific allosteric inhibitor of Akt. Sci Rep 7:11673
Ibanez MI, Cabello P, Luque-Almagro VM, Saez LP, Olaya A, Sanchez de Medina V, Luque de Castro MD, Moreno-Vivian C, Roldan MD (2017) Quantitative proteomic analysis of Pseudomonas pseudoalcaligenes CECT5344 in response to industrial cyanide-containing wastewaters using Liquid Chromatography-Mass Spectrometry/Mass Spectrometry (LC-MS/MS). PLoS One 12:e0172908
Ilies M, Iuga CA, Loghin F, Dhople VM, Thiele T, Volker U, Hammer E (2017) Impact of blood sample collection methods on blood protein profiling studies. Clin Chim Acta 471:128-134
Iruarrizaga-Lejarreta M, Varela-Rey M, Fernandez-Ramos D, Martinez-Arranz I, Delgado TC, Simon J, Juan VG, et al. (2017) Role of Aramchol in steatohepatitis and fibrosis in mice. Hepatol Commun 1:911-927
Jozefowicz AM, Hartmann A, Matros A, Schum A, Mock HP (2017) Nitrogen Deficiency Induced Alterations in the Root Proteome of a Pair of Potato (Solanum tuberosum L.) Varieties Contrasting for their Response to Low N. Proteomics 17 (In press)
Kalsch J, Pott LL, Takeda A, Kumamoto H, Mollmann D, Canbay A, Sitek B, Baba HA (2017) Bathing in carbon dioxide-enriched water alters protein expression in keratinocytes of skin tissue in rats. Int J Biometeorol 61:739-746
Kavanagh EL, Lindsay S, Halasz M, Gubbins LC, Weiner-Gorzel K, Guang MHZ, McGoldrick A, Collins E, Henry M, Blanco-Fernandez A, Gorman PO, Fitzpatrick P, Higgins MJ, Dowling P, McCann A (2017) Protein and chemotherapy profiling of extracellular vesicles harvested from therapeutic induced senescent triple negative breast cancer cells. Oncogenesis 6:e388
Keck M, Androsova G, Gualtieri F, Walker A, von Ruden EL, Russmann V, Deeg CA, Hauck SM, Krause R, Potschka H (2017) A systems level analysis of epileptogenesis-associated proteome alterations. Neurobiol Dis 105:164-178
Keys TG, Wetter M, Hang I, Rutschmann C, Russo S, Mally M, Steffen M, Zuppiger M, Muller F, Schneider J, Faridmoayer A, Lin CW, Aebi M (2017) A biosynthetic route for polysialylating proteins in Escherichia coli. Metab Eng 44:293-301
Kimura Y, Yanagimachi M, Ino Y, Aketagawa M, Matsuo M, Okayama A, Shimizu H, Oba K, Morioka I, Imagawa T, Kaneko T, Yokota S, Hirano H, Mori M (2017) Identification of candidate diagnostic serum biomarkers for Kawasaki disease using proteomic analysis. Sci Rep 7:43732
Klingeborn M, Dismuke WM, Skiba NP, Kelly U, Stamer WD, Bowes Rickman C (2017) Directional Exosome Proteomes Reflect Polarity-Specific Functions in Retinal Pigmented Epithelium Monolayers. Sci Rep 7:4901
Kneeshaw S, Keyani R, Delorme-Hinoux V, Imrie L, Loake GJ, Le Bihan T, Reichheld JP, Spoel SH (2017) Nucleoredoxin guards against oxidative stress by protecting antioxidant enzymes. Proc Natl Acad Sci USA (In press)
Kopczynski D, Sickmann A, Ahrends R (2017) Computational proteomics tools for identification and quality control. J Biotechnol 261:126-130
Kuusela P, Saraswat M, Joenvaara S, Kaartinen J, Jarvinen A, Renkonen R (2017) Changes in plasma protein levels as an early indication of a bloodstream infection. PLoS One 12:e0172987
Laakkonen EK, Soliymani R, Karvinen S, Kaprio J, Kujala UM, Baumann M, Sipila S, Kovanen V, Lalowski M (2017) Estrogenic regulation of skeletal muscle proteome: a study of premenopausal women and postmenopausal MZ cotwins discordant for hormonal therapy. Aging Cell 16:1276-1287
Labisch T, Buchkremer S, Phan V, Kollipara L, Gatz C, Lentz C, Nolte K, Vervoorts J, Coraspe JA, Sickmann A, Carr S, Zahedi RP, Weis J, Roos A (2017) Tracking Effects of SIL1 Increase: Taking a Closer Look Beyond the Consequences of Elevated Expression Level. Mol Neurobiol (In press)
Lachen-Montes M, Gonzalez-Morales A, Zelaya MV, Perez-Valderrama E, Ausin K, Ferrer I, Fernandez-Irigoyen J, Santamaria E (2017) Olfactory bulb neuroproteomics reveals a chronological perturbation of survival routes and a disruption of prohibitin complex during Alzheimer's disease progression. Sci Rep 7:9115
Lai MC, Bechy AL, Denk F, Collins E, Gavriliouk M, Zaugg JB, Ryan BJ, Wade-Martins R, Caffrey TM (2017) Haplotype-specific MAPT exon 3 expression regulated by common intronic polymorphisms associated with Parkinsonian disorders. Mol Neurodegener 12:79
Lamont L, Baumert M, Ogrinc Potocnik N, Allen M, Vreeken R, Heeren RMA, Porta T (2017) Integration of Ion Mobility MS(E) after Fully Automated, Online, High-Resolution Liquid Extraction Surface Analysis Micro-Liquid Chromatography. Anal Chem 89:11143-11150
Landolt L, Eikrem O, Strauss P, Scherer A, Lovett DH, Beisland C, Finne K, Osman T, Ibrahim MM, Gausdal G, Ahmed L, Lorens JB, Thiery JP, Tan TZ, Sekulic M, Marti HP (2017) Clear Cell Renal Cell Carcinoma is linked to Epithelial-to-Mesenchymal Transition and to Fibrosis. Physiol Rep 5 (In press)
Lee R, Fischer R, Charles PD, Adlam D, Valli A, Di Gleria K, Kharbanda RK, Choudhury RP, Antoniades C, Kessler BM, Channon KM (2017) A novel workflow combining plaque imaging, plaque and plasma proteomics identifies biomarkers of human coronary atherosclerotic plaque disruption. Clin Proteomics 14:22
Li SY, Oyala PH, Britt RD, Weintraub ST, Hunsicker-Wang LM (2017) Reactive sites and course of reduction in the Rieske protein. J Biol Inorg Chem 22:545-557
Liljedahl L, Norlin J, McGuire JN, James P (2017) Effects of insulin and the glucagon-like peptide 1 receptor agonist liraglutide on the kidney proteome in db/db mice. Physiol Rep 5 (In press)
Lin F, Williams BJ, Thangella PAV, Ladak A, Schepmoes AA, Olivos HJ, Zhao K, Callister SJ, Bartley LE (2017) Proteomics Coupled with Metabolite and Cell Wall Profiling Reveal Metabolic Processes of a Developing Rice Stem Internode. Front Plant Sci 8:1134
Lindner S, Gruhle K, Schmidt R, Garamus VM, Ramsbeck D, Hause G, Meister A, Sinz A, Drescher S (2017) Azide-Modified Membrane Lipids: Synthesis, Properties, and Reactivity. Langmuir 33:4960-4973
Liu X, Guo Z, Sun H, Li W, Sun W (2017) Comprehensive Map and Functional Annotation of Human Pituitary and Thyroid Proteome. J Proteome Res 16:2680-2691
Lopes-Rodrigues V, Di Luca A, Mleczko J, Meleady P, Henry M, Pesic M, Cabrera D, van Liempd S, Lima RT, O'Connor R, Falcon-Perez JM, Vasconcelos MH (2017) Identification of the metabolic alterations associated with the multidrug resistant phenotype in cancer and their intercellular transfer mediated by extracellular vesicles. Sci Rep 7:44541
Lopez JE, Haynes SE, Majmudar JD, Martin BR, Fierke CA (2017) HDAC8 Substrates Identified by Genetically Encoded Active Site Photocrosslinking. J Am Chem Soc 139:16222-16227
Lumpkin RJ, Gu H, Zhu Y, Leonard M, Ahmad AS, Clauser KR, Meyer JG, Bennett EJ, Komives EA (2017) Site-specific identification and quantitation of endogenous SUMO modifications under native conditions. Nat Commun 8:1171
Maddala R, Skiba NP, Rao PV (2017) Vertebrate Lonesome Kinase Regulated Extracellular Matrix Protein Phosphorylation, Cell Shape, and Adhesion in Trabecular Meshwork Cells. J Cell Physiol 232:2447-2460
McLeod A, Mosleth EF, Rud I, Branco Dos Santos F, Snipen L, Liland KH, Axelsson L (2017) Effects of glucose availability in Lactobacillus sakei; metabolic change and regulation of the proteome and transcriptome. PLoS One 12:e0187542
Megger DA, Rosowski K, Radunsky C, Kosters J, Sitek B, Muller J (2017) Structurally related hydrazone-based metal complexes with different antitumor activities variably induce apoptotic cell death. Dalton Trans 46:4759-4767
Mezni A, Khazri A, Khazri O, Limam F, Cosette P, Aouani E (2017) Neuroprotective Activity of Grape Seed and Skin Extract Against Lithium Exposure Using Proteomic Research. Mol Neurobiol 54:2720-2730
Migani D, Smales CM, Bracewell DG (2017) Effects of lysosomal biotherapeutic recombinant protein expression on cell stress and protease and general host cell protein release in Chinese hamster ovary cells. Biotechnol Prog 33:666-676
Miikkulainen P, Hogel H, Rantanen K, Suomi T, Kouvonen P, Elo LL, Jaakkola PM (2017) HIF prolyl hydroxylase PHD3 regulates translational machinery and glucose metabolism in clear cell renal cell carcinoma. Cancer Metab 5:5
Miller MAE, O'Cualain R, Selley J, Knight D, Karim MF, Hubbard SJ, Johnson GN (2017) Dynamic Acclimation to High Light in Arabidopsis thaliana Involves Widespread Reengineering of the Leaf Proteome. Front Plant Sci 8:1239
Miyake K, Mori R, Homma Y, Matsuyama R, Okayama A, Murakami T, Hirano H, Endo I (2017) MZB1 in borderline resectable pancreatic cancer resected after neoadjuvant chemoradiotherapy. J Surg Res 220:391-401
Mossina A, Lukas C, Merl-Pham J, Uhl FE, Mutze K, Schamberger A, Staab-Weijnitz C, Jia J, Yildirim AO, Konigshoff M, Hauck SM, Eickelberg O, Meiners S (2017) Cigarette smoke alters the secretome of lung epithelial cells. Proteomics 17 (In press)
Moussa E, Huang H, Ahras M, Lall A, Thezenas ML, Fischer R, Kessler BM, Pain A, Billker O, Casals-Pascual C (2017) Proteomic profiling of the brain of mice with experimental cerebral malaria. J Proteomics (In press)
Mulvihill ED, Moloney NM, Owens RA, Dolan SK, Russell L, Doyle S (2017) Functional Investigation of Iron-Responsive Microsomal Proteins, including MirC, in Aspergillus fumigatus. Front Microbiol 8:418
Munoz-Marin MD, Gomez-Baena G, Diez J, Beynon RJ, Gonzalez-Ballester D, Zubkov MV, Garcia-Fernandez JM (2017) Glucose Uptake in Prochlorococcus: Diversity of Kinetics and Effects on the Metabolism. Front Microbiol 8:327
Murphy S, Brinkmeier H, Krautwald M, Henry M, Meleady P, Ohlendieck K (2017) Proteomic profiling of the dystrophin complex and membrane fraction from dystrophic mdx muscle reveals decreases in the cytolinker desmoglein and increases in the extracellular matrix stabilizers biglycan and fibronectin. J Muscle Res Cell Motil 38:251-268
Myers OD, Sumner SJ, Li S, Barnes S, Du X (2017) One Step Forward for Reducing False Positive and False Negative Compound Identifications from Mass Spectrometry Metabolomics Data: New Algorithms for Constructing Extracted Ion Chromatograms and Detecting Chromatographic Peaks. Anal Chem 89:8696-8703
Nawrot R, Lippmann R, Matros A, Musidlak O, Nowicki G, Mock HP (2017) Proteomic comparison of Chelidonium majus L. latex in different phases of plant development. Plant Physiol Biochem 112:312-325
Neves GW, Curty NA, Kubitschek-Barreira PH, Fontaine T, Souza GH, Cunha ML, Goldman GH, Beauvais A, Latge JP, Lopes-Bezerra LM (2017) Modifications to the composition of the hyphal outer layer of Aspergillus fumigatus modulates HUVEC proteins related to inflammatory and stress responses. J Proteomics 151:83-96
Nguyen EV, Huhtinen K, Goo YA, Kaipio K, Andersson N, Rantanen V, Hynninen J, Lahesmaa R, Carpen O, Goodlett DR (2017) Hyper-phosphorylation of Sequestosome-1 Distinguishes Resistance to Cisplatin in Patient Derived High Grade Serous Ovarian Cancer Cells. Mol Cell Proteomics 16:1377-1392
Niksirat H, Andersson L, Golpour A, Chupani L, James P (2017) Quantification of egg proteome changes during fertilization in sterlet Acipenser ruthenus. Biochem Biophys Res Commun 490:189-193
Nygaard G, Herfindal L, Asrud KS, Bjornstad R, Kopperud RK, Oveland E, Berven FS, Myhren L, Hoivik EA, Lunde THF, Bakke M, Doskeland SO, Selheim F (2017) Epac1-deficient mice have bleeding phenotype and thrombocytes with decreased GPIbbeta expression. Sci Rep 7:8725
Obermann J, Priglinger CS, Merl-Pham J, Geerlof A, Priglinger S, Gotz M, Hauck SM (2017) Proteome-wide Identification of Glycosylation-dependent Interactors of Galectin-1 and Galectin-3 on Mesenchymal Retinal Pigment Epithelial (RPE) Cells. Mol Cell Proteomics 16:1528-1546
Ochsner AM, Christen M, Hemmerle L, Peyraud R, Christen B, Vorholt JA (2017) Transposon Sequencing Uncovers an Essential Regulatory Function of Phosphoribulokinase for Methylotrophy. Curr Biol 27:2579-2588 e2576
Oehmcke-Hecht S, Nass LE, Wichura JB, Mikkat S, Kreikemeyer B, Fiedler T (2017) Deletion of the L-Lactate Dehydrogenase Gene ldh in Streptococcus pyogenes Leads to a Loss of SpeB Activity and a Hypovirulent Phenotype. Front Microbiol 8:1841
O'Sullivan F, Keenan J, Aherne S, O'Neill F, Clarke C, Henry M, Meleady P, Breen L, Barron N, Clynes M, Horgan K, Doolan P, Murphy R (2017) Parallel mRNA, proteomics and miRNA expression analysis in cell line models of the intestine. World J Gastroenterol 23:7369-7386
Ozsvari B, Sotgia F, Lisanti MP (2017) A new mutation-independent approach to cancer therapy: Inhibiting oncogenic RAS and MYC, by targeting mitochondrial biogenesis. Aging (Albany NY) 9:2098-2116
Paban V, Loriod B, Villard C, Buee L, Blum D, Pietropaolo S, Cho YH, Gory-Faure S, Mansour E, Gharbi A, Alescio-Lautier B (2017) Omics analysis of mouse brain models of human diseases. Gene 600:90-100
Pappesch R, Warnke P, Mikkat S, Normann J, Wisniewska-Kucper A, Huschka F, Wittmann M, Khani A, Schwengers O, Oehmcke-Hecht S, Hain T, Kreikemeyer B, Patenge N (2017) The Regulatory Small RNA MarS Supports Virulence of Streptococcus pyogenes. Sci Rep 7:12241
Passamani LZ, Barbosa RR, Reis RS, Heringer AS, Rangel PL, Santa-Catarina C, Grativol C, Veiga CFM, Souza-Filho GA, Silveira V (2017) Salt stress induces changes in the proteomic profile of micropropagated sugarcane shoots. PLoS One 12:e0176076
Paunas TIF, Finne K, Leh S, Marti HP, Mollnes TE, Berven F, Vikse BE (2017) Glomerular abundance of complement proteins characterized by proteomic analysis of laser-captured microdissected glomeruli associates with progressive disease in IgA nephropathy. Clin Proteomics 14:30
Peffers M, Jones AR, McCabe A, Anderson J (2017) Neopeptide Analyser: A software tool for neopeptide discovery in proteomics data. Wellcome Open Res 2:24
Petushkova NA, Zgoda VG, Pyatnitskiy MA, Larina OV, Teryaeva NB, Potapov AA, Lisitsa AV (2017) Post-translational modifications of FDA-approved plasma biomarkers in glioblastoma samples. PLoS One 12:e0177427
Philipp J, Azimzadeh O, Subramanian V, Merl-Pham J, Lowe D, Hladik D, Erbeldinger N, Ktitareva S, Fournier C, Atkinson MJ, Raj K, Tapio S (2017) Radiation-Induced Endothelial Inflammation Is Transferred via the Secretome to Recipient Cells in a STAT-Mediated Process. J Proteome Res 16:3903-3916
Porte B, Chatelain C, Hardouin J, Derambure C, Zerdoumi Y, Hauchecorne M, Dupre N, Bekri S, Gonzalez B, Marret S, Cosette P, Leroux P (2017) Proteomic and transcriptomic study of brain microvessels in neonatal and adult mice. PLoS One 12:e0171048
Potu H, Peterson LF, Kandarpa M, Pal A, Sun H, Durham A, Harms PW, Hollenhorst PC, Eskiocak U, Talpaz M, Donato NJ (2017) Usp9x regulates Ets-1 ubiquitination and stability to control NRAS expression and tumorigenicity in melanoma. Nat Commun 8:14449
Procopio N, Chamberlain AT, Buckley M (2017) Intra- and Interskeletal Proteome Variations in Fresh and Buried Bones. J Proteome Res 16:2016-2029
Quesada-Calvo F, Massot C, Bertrand V, Longuespee R, Bletard N, Somja J, Mazzucchelli G, Smargiasso N, Baiwir D, De Pauw-Gillet MC, Delvenne P, Malaise M, Coimbra Marques C, Polus M, De Pauw E, Meuwis MA, Louis E (2017) OLFM4, KNG1 and Sec24C identified by proteomics and immunohistochemistry as potential markers of early colorectal cancer stages. Clin Proteomics 14:9
Ross KC, Andrews AJ, Marion CD, Yen TJ, Bhattacharjee V (2017) Identification of the Serine Biosynthesis Pathway as a Critical Component of BRAF Inhibitor Resistance of Melanoma, Pancreatic, and Non-Small Cell Lung Cancer Cells. Mol Cancer Ther 16:1596-1609
Salah A, Yoshifuji H, Ito S, Kitagori K, Kiso K, Yamada N, Nakajima T, Haga H, Tsuruyama T, Miyagawa-Hayashino A (2017) High Expression of Galectin-3 in Patients with IgG4-Related Disease: A Proteomic Approach. Patholog Res Int 2017:9312142
Salazar C, Jones MD, Sturtevant D, Horn PJ, Crossley J, Zaman K, Chapman KD, Wrona M, Isaac G, Smith NW, Shulaev V (2017) Development and application of sub-2-mum particle CO2 -based chromatography coupled to mass spectrometry for comprehensive analysis of lipids in cottonseed extracts. Rapid Commun Mass Spectrom 31:591-605
Saraswat M, Joenvaara S, Seppanen H, Mustonen H, Haglund C, Renkonen R (2017) Comparative proteomic profiling of the serum differentiates pancreatic cancer from chronic pancreatitis. Cancer Med 6:1738-1751
Saraswat M, Makitie A, Agarwal R, Joenvaara S, Renkonen S (2017) Oral squamous cell carcinoma patients can be differentiated from healthy individuals with label-free serum proteomics. Br J Cancer 117:376-384
Sarver DC, Kharaz YA, Sugg KB, Gumucio JP, Comerford E, Mendias CL (2017) Sex differences in tendon structure and function. J Orthop Res 35:2117-2126
Schopper S, Kahraman A, Leuenberger P, Feng Y, Piazza I, Muller O, Boersema PJ, Picotti P (2017) Measuring protein structural changes on a proteome-wide scale using limited proteolysis-coupled mass spectrometry. Nat Protoc 12:2391-2410
Schwenzer A, Quirke AM, Marzeda AM, Wong A, Montgomery AB, Sayles HR, Eick S, Gawron K, Chomyszyn-Gajewska M, Lazarz-Bartyzel K, Davis S, Potempa J, Kessler BM, Fischer R, Venables PJ, Payne JB, Mikuls TR, Midwood KS (2017) Association of Distinct Fine Specificities of Anti-Citrullinated Peptide Antibodies With Elevated Immune Responses to Prevotella intermedia in a Subgroup of Patients With Rheumatoid Arthritis and Periodontitis. Arthritis Rheumatol 69:2303-2313
Shevchenko A, Yang Y, Knaust A, Verbavatz JM, Mai H, Wang B, Wang C (2017) Open sesame: Identification of sesame oil and oil soot ink in organic deposits of Tang Dynasty lamps from Astana necropolis in China. PLoS One 12:e0158636
Sivaramakrishnan M, McCarthy KD, Campagne S, Huber S, Meier S, Augustin A, Heckel T, et al. (2017) Binding to SMN2 pre-mRNA-protein complex elicits specificity for small molecule splicing modifiers. Nat Commun 8:1476
Smine S, Obry A, Kadri S, Hardouin J, Freret M, Amri M, Jouenne T, Limam F, Cosette P, Aouani E (2017) Brain proteomic modifications associated to protective effect of grape extract in a murine model of obesity. Biochim Biophys Acta 1865:578-588
Soares EA, Werth EG, Madronero LJ, Ventura JA, Rodrigues SP, Hicks LM, Fernandes PM (2017) Label-free quantitative proteomic analysis of pre-flowering PMeV-infected Carica papaya L. J Proteomics 151:275-283
Sollanek KJ, Burniston JG, Kavazis AN, Morton AB, Wiggs MP, Ahn B, Smuder AJ, Powers SK (2017) Global Proteome Changes in the Rat Diaphragm Induced by Endurance Exercise Training. PLoS One 12:e0171007
Soman KV, Stafford SJ, Pazdrak K, Wu Z, Luo X, White WI, Wiktorowicz JE, Calhoun WJ, Kurosky A (2017) Activation of Human Peripheral Blood Eosinophils by Cytokines in a Comparative Time-Course Proteomic/Phosphoproteomic Study. J Proteome Res 16:2663-2679
Song L, Xue R, Ge P, Li M, Wang L, Zheng F, Zhao L, Wang Z, Wang Q, Liu N, Sun X (2017) Identification of post-translational modifications of Abeta peptide in platelet membranes from patients with cerebral amyloid angiopathy. J Neurol Sci 383:11-17
Stoop MP, Runia TF, Stingl C, van der Vuurst de Vries RM, Luider TM, Hintzen RQ (2017) Decreased Neuro-Axonal Proteins in CSF at First Attack of Suspected Multiple Sclerosis. Proteomics Clin Appl 11 (In press)
Sydor S, Sowa JP, Megger DA, Schlattjan M, Jafoui S, Wingerter L, Carpinteiro A, Baba HA, Bechmann LP, Sitek B, Gerken G, Gulbins E, Canbay A (2017) Acid sphingomyelinase deficiency in Western diet-fed mice protects against adipocyte hypertrophy and diet-induced liver steatosis. Mol Metab 6:416-427
Taleb RSZ, Moez P, Younan D, Eisenacher M, Tenbusch M, Sitek B, Bracht T (2017) Quantitative proteome analysis of plasma microparticles for the characterization of HCV-induced hepatic cirrhosis and hepatocellular carcinoma. Proteomics Clin Appl (In press)
Tay AP, Geoghegan V, Yagoub D, Wilkins MR, Hart-Smith G (2017) MethylQuant: A Tool for Sensitive Validation of Enzyme-Mediated Protein Methylation Sites from Heavy-Methyl SILAC Data. J Proteome Res 17:359-373
Ting KR, Henry M, Meiller J, Larkin A, Clynes M, Meleady P, Bazou D, Dowling P, O'Gorman P (2017) Novel panel of protein biomarkers to predict response to bortezomib-containing induction regimens in multiple myeloma patients. BBA Clin 8:28-34
Valikangas T, Suomi T, Elo LL (2017) A comprehensive evaluation of popular proteomics software workflows for label-free proteome quantification and imputation. Brief Bioinform (In press)
Venkatesh HS, Tam LT, Woo PJ, Lennon J, Nagaraja S, Gillespie SM, Ni J, Duveau DY, Morris PJ, Zhao JJ, Thomas CJ, Monje M (2017) Targeting neuronal activity-regulated neuroligin-3 dependency in high-grade glioma. Nature 549:533-537
Victorio VCM, Souza G, Santos MCB, Vega AR, Cameron LC, Ferreira MSL (2017) Differential expression of albumins and globulins of wheat flours of different technological qualities revealed by nanoUPLC-UDMS(E). Food Chem 239:1027-1036
Vizcaino JA, Mayer G, Perkins S, Barsnes H, Vaudel M, Perez-Riverol Y, Ternent T, Uszkoreit J, Eisenacher M, Fischer L, Rappsilber J, Netz E, Walzer M, Kohlbacher O, Leitner A, Chalkley RJ, Ghali F, Martinez-Bartolome S, Deutsch EW, Jones AR (2017) The mzIdentML Data Standard Version 1.2, Supporting Advances in Proteome Informatics. Mol Cell Proteomics 16:1275-1285
von Toerne C, Laimighofer M, Achenbach P, Beyerlein A, de Las Heras Gala T, Krumsiek J, Theis FJ, Ziegler AG, Hauck SM (2017) Peptide serum markers in islet autoantibody-positive children. Diabetologia 60:287-295
Wadsworth C, Procopio N, Anderung C, Carretero JM, Iriarte E, Valdiosera C, Elburg R, Penkman K, Buckley M (2017) Comparing ancient DNA survival and proteome content in 69 archaeological cattle tooth and bone samples from multiple European sites. J Proteomics 158:1-8
Wen M, Jin Y, Manabe T, Chen S, Tan W (2017) A comparative analysis of human plasma and serum proteins by combining native PAGE, whole-gel slicing and quantitative LC-MS/MS: Utilizing native MS-electropherograms in proteomic analysis for discovering structure and interaction-correlated differences. Electrophoresis 38:3111-3123
Werth EG, McConnell EW, Gilbert TS, Couso Lianez I, Perez CA, Manley CK, Graves LM, Umen JG, Hicks LM (2017) Probing the global kinome and phosphoproteome in Chlamydomonas reinhardtii via sequential enrichment and quantitative proteomics. Plant J 89:416-426
Willems S, Bouyssie D, David M, Locard-Paulet M, Mechtler K, Schwammle V, Uszkoreit J, Vaudel M, Dorfer V (2017) Proceedings of the EuBIC Winter School 2017. J Proteomics 161:78-80
Wu CY, Hsu WL, Tsai MH, Liang JL, Lu JH, Yen CJ, Yu HS, Noda M, Lu CY, Chen CH, Yan SJ, Yoshioka T (2017) Hydrogen gas protects IP3Rs by reducing disulfide bridges in human keratinocytes under oxidative stress. Sci Rep 7:3606
Yadetie F, Oveland E, Doskeland A, Berven F, Goksoyr A, Karlsen OA (2017) Quantitative proteomics analysis reveals perturbation of lipid metabolic pathways in the liver of Atlantic cod (Gadus morhua) treated with PCB 153. Aquat Toxicol 185:19-28
Yang K, Adin C, Shen Q, Lee LJ, Yu L, Fadda P, Samogyi A, Ham K, Xu L, Gilor C, Ziouzenkova O (2017) Aldehyde dehydrogenase 1 a1 regulates energy metabolism in adipocytes from different species. Xenotransplantation 24 (In press)
Yang X, Feng L, Zhao L, Liu X, Hassani D, Huang D (2017) Effect of glycine nitrogen on lettuce growth under soilless culture: a metabolomics approach to identify the main changes occurred in plant primary and secondary metabolism. J Sci Food Agric 98:467-477
Yau YY, Leong RWL, Pudipeddi A, Redmond D, Wasinger VC (2017) Serological Epithelial Component Proteins Identify Intestinal Complications in Crohn's Disease. Mol Cell Proteomics 16:1244-1257
Yuan ET, Ino Y, Kawaguchi M, Kimura Y, Hirano H, Kinoshita-Kikuta E, Kinoshita E, Koike T (2017) A Phos-tag-based micropipette-tip method for rapid and selective enrichment of phosphopeptides. Electrophoresis 38:2447-2455
Zhang Y, Beard KFM, Swart C, Bergmann S, Krahnert I, Nikoloski Z, Graf A, Ratcliffe RG, Sweetlove LJ, Fernie AR, Obata T (2017) Protein-protein interactions and metabolite channelling in the plant tricarboxylic acid cycle. Nat Commun 8:15212
Zhao M, Li M, Yang Y, Guo Z, Sun Y, Shao C, Sun W, Gao Y (2017) A comprehensive analysis and annotation of human normal urinary proteome. Sci Rep 7:3024
Zhou X, Xing X, Hou J, Liu J (2017) Quantitative proteomics analysis of proteins involved in alkane uptake comparing the profiling of Pseudomonas aeruginosa SJTD-1 in response to n-octadecane and n-hexadecane. PLoS One 12:e0179842
2016
Acioglu C, Mirabelli E, Baykal AT, Ni L, Ratnayake A, Heary RF, Elkabes S (2016) Toll like receptor 9 antagonism modulates spinal cord neuronal function and survival: Direct versus astrocyte-mediated mechanisms. Brain Behav Immun 56:310-324
Ademowo OS, Hernandez B, Collins E, Rooney C, Fearon U, van Kuijk AW, Tak PP, Gerlag DM, FitzGerald O, Pennington SR (2016) Discovery and confirmation of a protein biomarker panel with potential to predict response to biological therapy in psoriatic arthritis. Ann Rheum Dis 75:234-241
Ahrne E, Glatter T, Vigano C, Schubert C, Nigg EA, Schmidt A (2016) Evaluation and Improvement of Quantification Accuracy in Isobaric Mass Tag-Based Protein Quantification Experiments. J Proteome Res 15:2537-2547
Akhtar MZ, Huang H, Kaisar M, Lo Faro ML, Rebolledo R, Morten K, Heather LC, Dona A, Leuvenink HG, Fuggle SV, Kessler BM, Pugh CW, Ploeg RJ (2016) Using an Integrated -Omics Approach to Identify Key Cellular Processes That Are Disturbed in the Kidney After Brain Death. Am J Transplant 16:1421-1440
Almeida AM, Nanni P, Ferreira AM, Fortes C, Grossmann J, Bessa RJ, Costa P (2016) The longissimus thoracis muscle proteome in Alentejana bulls as affected by growth path. J Proteomics 152:206-215
Angi M, Kalirai H, Prendergast S, Simpson D, Hammond DE, Madigan MC, Beynon RJ, Coupland SE (2016) In-depth proteomic profiling of the uveal melanoma secretome. Oncotarget 7:49623-49635
Annibal A, Riemer T, Jovanovic O, Westphal D, Griesser E, Pohl EE, Schiller J, Hoffmann R, Fedorova M (2016) Structural, biological and biophysical properties of glycated and glycoxidized phosphatidylethanolamines. Free Radic Biol Med 95:293-307
Armstrong SD, Xia D, Bah GS, Krishna R, Ngangyung HF, LaCourse EJ, McSorley HJ, Kengne-Ouafo JA, Chounna-Ndongmo PW, Wanji S, Enyong PA, Taylor DW, Blaxter ML, Wastling JM, Tanya VN, Makepeace BL (2016) Stage-specific Proteomes from Onchocerca ochengi, Sister Species of the Human River Blindness Parasite, Uncover Adaptations to a Nodular Lifestyle. Mol Cell Proteomics 15:2554-2575
Bauer M, Cubizolles F, Schmidt A, Nigg EA (2016) Quantitative analysis of human centrosome architecture by targeted proteomics and fluorescence imaging. Embo J 35:2152-2166
Bespyatykh J, Shitikov E, Butenko I, Altukhov I, Alexeev D, Mokrousov I, Dogonadze M, Zhuravlev V, Yablonsky P, Ilina E, Govorun V (2016) Proteome analysis of the Mycobacterium tuberculosis Beijing B0/W148 cluster. Sci Rep 6:28985
Betz BB, Jenks SJ, Cronshaw AD, Lamont DJ, Cairns C, Manning JR, Goddard J, Webb DJ, Mullins JJ, Hughes J, McLachlan S, Strachan MW, Price JF, Conway BR (2016) Urinary peptidomics in a rodent model of diabetic nephropathy highlights epidermal growth factor as a biomarker for renal deterioration in patients with type 2 diabetes. Kidney Int 89:1125-1135
Bivol LM, Iversen BM, Hultstrom M, Wallace PW, Reed RK, Wiig H, Tenstad O (2016) Unilateral renal ischaemia in rats induces a rapid secretion of inflammatory markers to renal lymph and increased capillary permeability. J Physiol 594:1709-1726
Blein-Nicolas M, Zivy M (2016) Thousand and one ways to quantify and compare protein abundances in label-free bottom-up proteomics. Biochim Biophys Acta 1864:883-895
Bonde MT, Pedersen M, Klausen MS, Jensen SI, Wulff T, Harrison S, Nielsen AT, Herrgard MJ, Sommer MO (2016) Predictable tuning of protein expression in bacteria. Nat Methods 13:233-236
Borgdorff H, Armstrong SD, Tytgat HL, Xia D, Ndayisaba GF, Wastling JM, van de Wijgert JH (2016) Unique Insights in the Cervicovaginal Lactobacillus iners and L. crispatus Proteomes and Their Associations with Microbiota Dysbiosis. PLoS One 11:e0150767
Branson OE, Freitas MA (2016) Tag-Count Analysis of Large-Scale Proteomic Data. J Proteome Res 15:4742-4746
Brauers E, Roos A, Kollipara L, Zahedi RP, Beckmann A, Mohanadas N, Bauer H, Hausler M, Thoma S, Kress W, Senderek J, Weis J (2016) The Caveolin-3 G56S sequence variant of unknown significance: Muscle biopsy findings and functional cell biological analysis. Proteomics Clin Appl (In press)
Broeckx V, Boonen K, Pringels L, Sagaert X, Prenen H, Landuyt B, Schoofs L, Maes E (2016) Comparison of multiple protein extraction buffers for GeLC-MS/MS proteomic analysis of liver and colon formalin-fixed, paraffin-embedded tissues. Mol Biosyst 12:553-565
Broering R, Trippler M, Werner M, Real CI, Megger DA, Bracht T, Schweinsberg V, Sitek B, Eisenacher M, Meyer HE, Baba HA, Weber F, Hoffmann AC, Gerken G, Schlaak JF (2016) Hepatic expression of proteasome subunit alpha type-6 is upregulated during viral hepatitis and putatively regulates the expression of ISG15 ubiquitin-like modifier, a proviral host gene in hepatitis C virus infection. J Viral Hepat 23:375-386
Buts K, Hertog ML, Ho QT, America AH, Cordewener J, Vercammen J, Carpentier SC, Nicolai B (2016) Influence of pre-harvest calcium, potassium and triazole application on the proteome of apple at harvest. J Sci Food Agric 96:4984-4993
Cappelle K, Smagghe G, Dhaenens M, Meeus I (2016) Israeli Acute Paralysis Virus Infection Leads to an Enhanced RNA Interference Response and Not Its Suppression in the Bumblebee Bombus terrestris. Viruses 8 of apple at harvest. J Sci Food Agric 96:4984-4993
Carella P, Merl-Pham J, Wilson DC, Dey S, Hauck SM, Vlot AC, Cameron RK (2016) Comparative Proteomics Analysis of Phloem Exudates Collected during the Induction of Systemic Acquired Resistance. Plant Physiol 171:1495-1510
Casadei N, Sood P, Ulrich T, Fallier-Becker P, Kieper N, Helling S, May C, Glaab E, Chen J, Nuber S, Marcus K, Rapaport D, Ott T, Riess O, Kruger R, Fitzgerald JC (2016) Mitochondrial defects and neurodegeneration in mice overexpressing wild-type or G399S mutant HtrA2. Hum Mol Genet 25:459-471
Cevik O, Baykal AT, Sener A (2016) Platelets Proteomic Profiles of Acute Ischemic Stroke Patients. PLoS One 11:e0158287
Chen F, Spiessens C, D'Hooghe T, Peeraer K, Carpentier S (2016) Follicular fluid biomarkers for human in vitro fertilization outcome: Proof of principle. Proteome Sci 14:17
Cholet C, Claverol S, Claisse O, Rabot A, Osowsky A, Dumot V, Ferrari G, Geny L (2016) Tartaric acid pathways in Vitis vinifera L. (cv. Ugni blanc): a comparative study of two vintages with contrasted climatic conditions. BMC Plant Biol 16:144
Ciregia F, Kollipara L, Giusti L, Zahedi RP, Giacomelli C, Mazzoni MR, Giannaccini G, Scarpellini P, Urbani A, Sickmann A, Lucacchini A, Bazzichi L (2016) Bottom-up proteomics suggests an association between differential expression of mitochondrial proteins and chronic fatigue syndrome. Transl Psychiatry 6:e904
Colak D, Alaiya AA, Kaya N, Muiya NP, AlHarazi O, Shinwari Z, Andres E, Dzimiri N (2016) Integrated Left Ventricular Global Transcriptome and Proteome Profiling in Human End-Stage Dilated Cardiomyopathy. PLoS One 11:e0162669
Correia S, Hebraud M, Chafsey I, Chambon C, Viala D, Torres C, de Toro M, Capelo JL, Poeta P, Igrejas G (2016) Impacts of experimentally induced and clinically acquired quinolone resistance on the membrane and intracellular subproteomes of Salmonella Typhimurium DT104B. J Proteomics 145:46-59
Coskun A, Baykal AT, Kazan D, Akgoz M, Senal MO, Berber I, Titiz I, Bilsel G, Kilercik H, Karaosmanoglu K, Cicek M, Yurtsever I, Yazici C (2016) Proteomic Analysis of Kidney Preservation Solutions Prior to Renal Transplantation. PLoS One 11:e0168755
Cowan E, Kumar P, Burch KJ, Grieve DJ, Green BD, Graham SF (2016) Treatment of lean and diet-induced obesity (DIO) mice with a novel stable obestatin analogue alters plasma metabolite levels as detected by untargeted LC-MS metabolomics. Metabolomics 12:124
Del Boccio P, Rossi C, di Ioia M, Cicalini I, Sacchetta P, Pieragostino D (2016) Integration of metabolomics and proteomics in multiple sclerosis: From biomarkers discovery to personalized medicine. Proteomics Clin Appl 10:470-484
Demarchi B, Hall S, Roncal-Herrero T, Freeman CL, Woolley J, Crisp MK, Wilson J, et al. (2016) Protein sequences bound to mineral surfaces persist into deep time. Elife 5
Denis S, Sayd T, Georges A, Chambon C, Chalancon S, Sante-Lhoutellier V, Blanquet-Diot S (2016) Digestion of cooked meat proteins is slightly affected by age as assessed using the dynamic gastrointestinal TIM model and mass spectrometry. Food Funct 7:2682-2691
Dhawi F, Datta R, Ramakrishna W (2016) Proteomics provides insights into biological pathways altered by plant growth promoting bacteria and arbuscular mycorrhiza in sorghum grown in marginal soil. Biochim Biophys Acta 1865:243-251
Dionisio G, Kryger P, Steenberg T (2016) Label-Free Differential Proteomics and Quantification of Exoenzymes from Isolates of the Entomopathogenic Fungus Beauveria bassiana. Insects 7 (in press)
Distler U, Kuharev J, Navarro P, Tenzer S (2016) Label-free quantification in ion mobility-enhanced data-independent acquisition proteomics. Nat Protoc 11:795-812
Dong S, Liu L, Wu W, Armstrong SD, Xia D, Nan H, Hiscox JA, Chen H (2016) Determination of the interactome of non-structural protein12 from highly pathogenic porcine reproductive and respiratory syndrome virus with host cellular proteins using high throughput proteomics and identification of HSP70 as a cellular factor for virus replication. J Proteomics 146:58-69
Dowle AA, Wilson J, Thomas JR (2016) Comparing the Diagnostic Classification Accuracy of iTRAQ, Peak-Area, Spectral-Counting, and emPAI Methods for Relative Quantification in Expression Proteomics. J Proteome Res 15:3550-3562
Downs ML, Baumert JL, Taylor SL, Mills EN (2016) Mass spectrometric analysis of allergens in roasted walnuts. J Proteomics 142:62-69
Dudekula K, Le Bihan T (2016) Data from quantitative label free proteomics analysis of rat spleen. Data Brief 8:494-500
Duressa TF, Boonen K, Huybrechts R (2016) A quantitative peptidomics approach to unravel immunological functions of angiotensin converting enzyme in Locusta migratoria. Gen Comp Endocrinol 235:120-129
Fascellaro G, Petrera A, Lai ZW, Nanni P, Grossmann J, Burger S, Biniossek ML, Gomez-Auli A, Schilling O, Imkamp F (2016) Comprehensive Proteomic Analysis of Nitrogen-Starved Mycobacterium smegmatis Deltapup Reveals the Impact of Pupylation on Nitrogen Stress Response. J Proteome Res 15:2812-2825
Frolov A, Bilova T, Paudel G, Berger R, Balcke GU, Birkemeyer C, Wessjohann LA (2016) Early responses of mature Arabidopsis thaliana plants to reduced water potential in the agar-based polyethylene glycol infusion drought model. J Plant Physiol 208:70-83
Gasser V, Baco E, Cunrath O, August PS, Perraud Q, Zill N, Schleberger C, Schmidt A, Paulen A, Bumann D, Mislin GL, Schalk IJ (2016) Catechol siderophores repress the pyochelin pathway and activate the enterobactin pathway in Pseudomonas aeruginosa: an opportunity for siderophore-antibiotic conjugates development. Environ Microbiol 18:819-832
Geiger R, Rieckmann JC, Wolf T, Basso C, Feng Y, Fuhrer T, Kogadeeva M, Picotti P, Meissner F, Mann M, Zamboni N, Sallusto F, Lanzavecchia A (2016) L-Arginine Modulates T Cell Metabolism and Enhances Survival and Anti-tumor Activity. Cell 167:829-842 e813
Gergondey R, Garcia C, Serre V, Camadro JM, Auchere F (2016) The adaptive metabolic response involves specific protein glutathionylation during the filamentation process in the pathogen Candida albicans. Biochim Biophys Acta 1862:1309-1323
Govaert E, Van Steendam K, Scheerlinck E, Vossaert L, Meert P, Stella M, Willems S, De Clerck L, Dhaenens M, Deforce D (2016) Extracting histones for the specific purpose of label-free MS. Proteomics 16:2937-2944
Grandl G, Muller S, Moest H, Moser C, Wollscheid B, Wolfrum C (2016) Depot specific differences in the adipogenic potential of precursors are mediated by collagenous extracellular matrix and Flotillin 2 dependent signaling. Mol Metab 5:937-947
Grosche A, Hauser A, Lepper MF, Mayo R, von Toerne C, Merl-Pham J, Hauck SM (2016) The Proteome of Native Adult Muller Glial Cells From Murine Retina. Mol Cell Proteomics 15:462-480
Grouneva I, Muth-Pawlak D, Battchikova N, Aro EM (2016) Changes in Relative Thylakoid Protein Abundance Induced by Fluctuating Light in the Diatom Thalassiosira pseudonana. J Proteome Res 15:1649-1658
Gu H, Ren JM, Jia X, Levy T, Rikova K, Yang V, Lee KA, Stokes MP, Silva JC (2016) Quantitative Profiling of Post-translational Modifications by Immunoaffinity Enrichment and LC-MS/MS in Cancer Serum without Immunodepletion. Mol Cell Proteomics 15:692-702
Gutierrez-Carbonell E, Takahashi D, Luthje S, Gonzalez-Reyes JA, Mongrand S, Contreras-Moreira B, Abadia A, Uemura M, Abadia J, Lopez-Millan AF (2016) A Shotgun Proteomic Approach Reveals That Fe Deficiency Causes Marked Changes in the Protein Profiles of Plasma Membrane and Detergent-Resistant Microdomain Preparations from Beta vulgaris Roots. J Proteome Res 15:2510-2524
Hammond DE, Claydon AJ, Simpson DM, Edward D, Stockley P, Hurst JL, Beynon RJ (2016) Proteome Dynamics: Tissue Variation in the Kinetics of Proteostasis in Intact Animals. Mol Cell Proteomics 15:1204-1219
Heinz A, Huertas AC, Schrader CU, Pankau R, Gosch A, Schmelzer CE (2016) Elastins from patients with Williams-Beuren syndrome and healthy individuals differ on the molecular level. Am J Med Genet A 170:1832-1842
Heinzelmann K, Noskovicova N, Merl-Pham J, Preissler G, Winter H, Lindner M, Hatz R, Hauck SM, Behr J, Eickelberg O (2016) Surface proteome analysis identifies platelet derived growth factor receptor-alpha as a critical mediator of transforming growth factor-beta-induced collagen secretion. Int J Biochem Cell Biol 74:44-59
Henderson MX, Wirak GS, Zhang YQ, Dai F, Ginsberg SD, Dolzhanskaya N, Staropoli JF, Nijssen PC, Lam TT, Roth AF, Davis NG, Dawson G, Velinov M, Chandra SS (2016) Neuronal ceroid lipofuscinosis with DNAJC5/CSPalpha mutation has PPT1 pathology and exhibit aberrant protein palmitoylation. Acta Neuropathol 131:621-637
Hernandez-Castellano LE, Ferreira AM, Nanni P, Grossmann J, Arguello A, Capote J, Cai G, Lippolis J, Castro N, de Almeida AM (2016) The goat (Capra hircus) mammary gland secretory tissue proteome as influenced by weight loss: A study using label free proteomics. J Proteomics 145:60-69
Hoehenwarter W, Monchgesang S, Neumann S, Majovsky P, Abel S, Muller J (2016) Comparative expression profiling reveals a role of the root apoplast in local phosphate response. BMC Plant Biol 16:106
Holman SW, Hammond DE, Simpson DM, Waters J, Hurst JL, Beynon RJ (2016) Protein turnover measurement using selected reaction monitoring-mass spectrometry (SRM-MS). Philos Trans A Math Phys Eng Sci 374
Horton ER, Humphries JD, Stutchbury B, Jacquemet G, Ballestrem C, Barry ST, Humphries MJ (2016) Modulation of FAK and Src adhesion signaling occurs independently of adhesion complex composition. J Cell Biol 212:349-364
Hunt VL, Tsai IJ, Coghlan A, Reid AJ, Holroyd N, Foth BJ, Tracey A, et al. (2016) The genomic basis of parasitism in the Strongyloides clade of nematodes. Nat Genet 48:299-307
Ino Y, Arakawa N, Ishiguro H, Uemura H, Kubota Y, Hirano H, Toda T (2016) Phosphoproteome analysis demonstrates the potential role of THRAP3 phosphorylation in androgen-independent prostate cancer cell growth. Proteomics 16:1069-1078
Johnson PE, Sayers RL, Gethings LA, Balasundaram A, Marsh JT, Langridge JI, Mills EN (2016) Quantitative Proteomic Profiling of Peanut Allergens in Food Ingredients Used for Oral Food Challenges. Anal Chem 88:5689-5695
Kahne T, Richter S, Kolodziej A, Smalla KH, Pielot R, Engler A, Ohl FW, Dieterich DC, Seidenbecher C, Tischmeyer W, Naumann M, Gundelfinger ED (2016) Proteome rearrangements after auditory learning: high-resolution profiling of synapse-enriched protein fractions from mouse brain. J Neurochem 138:124-138
Kaisar M, van Dullemen LF, Thezenas ML, Charles PD, Ploeg RJ, Kessler BM (2016) Plasma Biomarker Profile Alterations during Variable Blood Storage. Clin Chem 62:1272-1274
Kaisar M, van Dullemen LF, Thezenas ML, Zeeshan Akhtar M, Huang H, Rendel S, Charles PD, Fischer R, Ploeg RJ, Kessler BM (2016) Plasma degradome affected by variable storage of human blood. Clin Proteomics 13:26
Kanonidis EI, Roy MM, Deighton RF, Le Bihan T (2016) Protein Co-Expression Analysis as a Strategy to Complement a Standard Quantitative Proteomics Approach: Case of a Glioblastoma Multiforme Study. PLoS One 11:e0161828
Kelly PS, McSweeney S, Coleman O, Carillo S, Henry M, Chandran D, Kellett A, Bones J, Clynes M, Meleady P, Barron N (2016) Process-relevant concentrations of the leachable bDtBPP impact negatively on CHO cell production characteristics. Biotechnol Prog 32:1547-1558
Kharaz YA, Tew SR, Peffers M, Canty-Laird EG, Comerford E (2016) Proteomic differences between native and tissue-engineered tendon and ligament. Proteomics 16:1547-1556
Klobucar M, Sedic M, Gehrig P, Grossmann J, Bilic M, Kovac-Bilic L, Pavelic K, Kraljevic Pavelic S (2016) Basement membrane protein ladinin-1 and the MIF-CD44-beta1 integrin signaling axis are implicated in laryngeal cancer metastasis. Biochim Biophys Acta 1862:1938-1954
Kohtala S, Theilmann W, Suomi T, Wigren HK, Porkka-Heiskanen T, Elo LL, Rokka A, Rantamaki T (2016) Brief Isoflurane Anesthesia Produces Prominent Phosphoproteomic Changes in the Adult Mouse Hippocampus. ACS Chem Neurosci 7:749-756
Konig T, Troder SE, Bakka K, Korwitz A, Richter-Dennerlein R, Lampe PA, Patron M, Muhlmeister M, Guerrero-Castillo S, Brandt U, Decker T, Lauria I, Paggio A, Rizzuto R, Rugarli EI, De Stefani D, Langer T (2016) The m-AAA Protease Associated with Neurodegeneration Limits MCU Activity in Mitochondria. Mol Cell 64:148-162
Kras KA, Willis WT, Barker N, Czyzyk T, Langlais PR, Katsanos CS (2016) Subsarcolemmal mitochondria isolated with the proteolytic enzyme nagarse exhibit greater protein specific activities and functional coupling. Biochem Biophys Rep 6:101-107
Kumar Y, Zhang L, Panigrahi P, Dholakia BB, Dewangan V, Chavan SG, Kunjir SM, Wu X, Li N, Rajmohanan PR, Kadoo NY, Giri AP, Tang H, Gupta VS (2016) Fusarium oxysporum mediates systems metabolic reprogramming of chickpea roots as revealed by a combination of proteomics and metabolomics. Plant Biotechnol J 14:1589-1603
Lachen-Montes M, Gonzalez-Morales A, de Morentin XM, Perez-Valderrama E, Ausin K, Zelaya MV, Serna A, Aso E, Ferrer I, Fernandez-Irigoyen J, Santamaria E (2016) An early dysregulation of FAK and MEK/ERK signaling pathways precedes the beta-amyloid deposition in the olfactory bulb of APP/PS1 mouse model of Alzheimer's disease. J Proteomics 148:149-158
Lereim RR, Oveland E, Xiao Y, Torkildsen O, Wergeland S, Myhr KM, Sun SC, Berven FS (2016) The Brain Proteome of the Ubiquitin Ligase Peli1 Knock-Out Mouse during Experimental Autoimmune Encephalomyelitis. J Proteomics Bioinform 9:209-219
Leroy C, Ramos P, Cornille K, Bonenfant D, Fritsch C, Voshol H, Bentires-Alj M (2016) Activation of IGF1R/p110beta/AKT/mTOR confers resistance to alpha-specific PI3K inhibition. Breast Cancer Res 18:41
Le-Trilling VT, Megger DA, Katschinski B, Landsberg CD, Ruckborn MU, Tao S, Krawczyk A, Bayer W, Drexler I, Tenbusch M, Sitek B, Trilling M (2016) Broad and potent antiviral activity of the NAE inhibitor MLN4924. Sci Rep 6:19977
Li X, Gao Y (2016) Potential urinary aging markers of 20-month-old rats. PeerJ 4:e2058
Liljedahl L, Pedersen MH, Norlin J, McGuire JN, James P (2016) N-glycosylation proteome enrichment analysis in kidney reveals differences between diabetic mouse models. Clin Proteomics 13:22
Limonier F, Van Steendam K, Waeterloos G, Brusselmans K, Sneyers M, Deforce D (2016) An application of mass spectrometry for quality control of biologicals: Highly sensitive profiling of plasma residuals in human plasma-derived immunoglobulin. J Proteomics 152:312-320
Lin CY, Chen CY, Yu CH, Yu IS, Lin SR, Wu JT, Lin YH, Kuo PL, Wu JC, Lin SW (2016) Human X-linked Intellectual Disability Factor CUL4B Is Required for Post-meiotic Sperm Development and Male Fertility. Sci Rep 6:20227
Lin PC, Yang YF, Tyan YC, Hsiao ES, Chu PC, Lee CT, Lee JC, Chen YM, Liao PC (2016) Identification of Phosphorylated Cyclin-Dependent Kinase 1 Associated with Colorectal Cancer Survival Using Label-Free Quantitative Analyses. PLoS One 11:e0158844
Liu F, Bai X, Yang FQ, Zhang XJ, Hu Y, Li P, Wan JB (2016) Discriminating from species of Curcumae Radix (Yujin) by a UHPLC/Q-TOFMS-based metabolomics approach. Chin Med 11:21
Lochmatter C, Fischer R, Charles PD, Yu Z, Powrie F, Kessler BM (2016) Integrative Phosphoproteomics Links IL-23R Signaling with Metabolic Adaptation in Lymphocytes. Sci Rep 6:24491
Lockman KA, Htun V, Sinha R, Treskes P, Nelson LJ, Martin SF, Rogers SM, Le Bihan T, Hayes PC, Plevris JN (2016) Proteomic profiling of cellular steatosis with concomitant oxidative stress in vitro. Lipids Health Dis 15:114
Lohr K, Pachl F, Moghaddas Gholami A, Geillinger KE, Daniel H, Kuster B, Klingenspor M (2016) Reduced mitochondrial mass and function add to age-related susceptibility toward diet-induced fatty liver in C57BL/6J mice. Physiol Rep 4 (In press)
Ly A, Merl-Pham J, Priller M, Gruhn F, Senninger N, Ueffing M, Hauck SM (2016) Proteomic Profiling Suggests Central Role Of STAT Signaling during Retinal Degeneration in the rd10 Mouse Model. J Proteome Res 15:1350-1359
McClure-Begley TD, Esterlis I, Stone KL, Lam TT, Grady SR, Colangelo CM, Lindstrom JM, Marks MJ, Picciotto MR (2016) Evaluation of the Nicotinic Acetylcholine Receptor-Associated Proteome at Baseline and Following Nicotine Exposure in Human and Mouse Cortex. eNeuro 3
McConnell JC, O'Connell OV, Brennan K, Weiping L, Howe M, Joseph L, Knight D, O'Cualain R, Lim Y, Leek A, Waddington R, Rogan J, Astley SM, Gandhi A, Kirwan CC, Sherratt MJ, Streuli CH (2016) Increased peri-ductal collagen micro-organization may contribute to raised mammographic density. Breast Cancer Res 18:5
Moloney NM, Owens RA, Meleady P, Henry M, Dolan SK, Mulvihill E, Clynes M, Doyle S (2016) The iron-responsive microsomal proteome of Aspergillus fumigatus. J Proteomics 136:99-111
Moskovets EV, Ivanov AR (2016) Comparative studies of peak intensities and chromatographic separation of proteolytic digests, PTMs, and intact proteins obtained by nanoLC-ESI MS analysis at room and elevated temperatures. Anal Bioanal Chem 408:3953-3968
Mulay A, Akram KM, Williams D, Armes H, Russell C, Hood D, Armstrong S, Stewart JP, Brown SD, Bingle L, Bingle CD (2016) An in vitro model of murine middle ear epithelium. Dis Model Mech 9:1405-1417
Murphy S, Dowling P, Zweyer M, Mundegar RR, Henry M, Meleady P, Swandulla D, Ohlendieck K (2016) Proteomic analysis of dystrophin deficiency and associated changes in the aged mdx-4cv heart model of dystrophinopathy-related cardiomyopathy. J Proteomics 145:24-36
Naboulsi W, Bracht T, Megger DA, Reis H, Ahrens M, Turewicz M, Eisenacher M, Tautges S, Canbay AE, Meyer HE, Weber F, Baba HA, Sitek B (2016) Quantitative proteome analysis reveals the correlation between endocytosis-associated proteins and hepatocellular carcinoma dedifferentiation. Biochim Biophys Acta 1864:1579-1585
Nakamura H, Yamashita N, Kimura A, Kimura Y, Hirano H, Makihara H, Kawamoto Y, Jitsuki-Takahashi A, Yonezaki K, Takase K, Miyazaki T, Nakamura F, Tanaka F, Goshima Y (2016) Comprehensive behavioral study and proteomic analyses of CRMP2-deficient mice. Genes Cells 21:1059-1079
Nassar AF, Williams BJ, Yaworksy DC, Patel V, Rusling JF (2016) Rapid label-free profiling of oral cancer biomarker proteins using nano-UPLC-Q-TOF ion mobility mass spectrometry. Proteomics Clin Appl 10:280-289
Okayama A, Kimura Y, Miyagi Y, Oshima T, Oshita F, Ito H, Nakayama H, Nagashima T, Rino Y, Masuda M, Ryo A, Hirano H (2016) Relationship between phosphorylation of sperm-specific antigen and prognosis of lung adenocarcinoma. J Proteomics 139:60-66
Opsahl JA, Vaudel M, Guldbrandsen A, Aasebo E, Van Pesch V, Franciotta D, Myhr KM, Barsnes H, Berle M, Torkildsen O, Kroksveen AC, Berven FS (2016) Label-free analysis of human cerebrospinal fluid addressing various normalization strategies and revealing protein groups affected by multiple sclerosis. Proteomics 16:1154-1165
Palfi A, Hokamp K, Hauck SM, Vencken S, Millington-Ward S, Chadderton N, Carrigan M, Kortvely E, Greene CM, Kenna PF, Farrar GJ (2016) microRNA regulatory circuits in a mouse model of inherited retinal degeneration. Sci Rep 6:31431
Paudel G, Bilova T, Schmidt R, Greifenhagen U, Berger R, Tarakhovskaya E, Stockhardt S, Balcke GU, Humbeck K, Brandt W, Sinz A, Vogt T, Birkemeyer C, Wessjohann L, Frolov A (2016) Osmotic stress is accompanied by protein glycation in Arabidopsis thaliana. J Exp Bot 67:6283-6295
Peters M, Guidato PM, Peters K, Megger DA, Sitek B, Classen B, Heise EM, Bufe A (2016) Allergy-Protective Arabinogalactan Modulates Human Dendritic Cells via C-Type Lectins and Inhibition of NF-kappaB. J Immunol 196:1626-1635
Pu Z, Ino Y, Kimura Y, Tago A, Shimizu M, Natsume S, Sano Y, Fujimoto R, Kaneko K, Shea DJ, Fukai E, Fuji S, Hirano H, Okazaki K (2016) Changes in the Proteome of Xylem Sap in Brassica oleracea in Response to Fusarium oxysporum Stress. Front Plant Sci 7:31
Rabbani N, Ashour A, Thornalley PJ (2016) Mass spectrometric determination of early and advanced glycation in biology. Glycoconj J 33:553-568
Radzikowski JL, Vedelaar S, Siegel D, Ortega AD, Schmidt A, Heinemann M (2016) Bacterial persistence is an active sigmaS stress response to metabolic flux limitation. Mol Syst Biol 12:882
Richards SL, Cawley AT, Cavicchioli R, Suann CJ, Pickford R, Raftery MJ (2016) Aptamer based peptide enrichment for quantitative analysis of gonadotropin-releasing hormone by LC-MS/MS. Talanta 150:671-680
Riis S, Stensballe A, Emmersen J, Pennisi CP, Birkelund S, Zachar V, Fink T (2016) Mass spectrometry analysis of adipose-derived stem cells reveals a significant effect of hypoxia on pathways regulating extracellular matrix. Stem Cell Res Ther 7:52
Romas L, Birse K, Mayer KH, Abou M, Westmacott G, Giguere R, Febo I, Cranston RD, Carballo-Dieguez A, McGowan I, Burgener A (2016) Rectal 1% Tenofovir Gel Use Associates with Altered Epidermal Protein Expression. AIDS Res Hum Retroviruses 32:1005-1015
Rombouts I, Lambrecht MA, Carpentier SC, Delcour JA (2016) Identification of lanthionine and lysinoalanine in heat-treated wheat gliadin and bovine serum albumin using tandem mass spectrometry with higher-energy collisional dissociation. Amino Acids 48:959-971
Sabbagh B, Mindt S, Neumaier M, Findeisen P (2016) Clinical applications of MS-based protein quantification. Proteomics Clin Appl 10:323-345
Saia-Cereda VM, Cassoli JS, Schmitt A, Falkai P, Martins-de-Souza D (2016) Differential proteome and phosphoproteome may impact cell signaling in the corpus callosum of schizophrenia patients. Schizophr Res 177:70-77
Sayd T, Chambon C, Sante-Lhoutellier V (2016) Quantification of peptides released during in vitro digestion of cooked meat. Food Chem 197 Pt B:1311-1323
Schafer N, Grosche A, Reinders J, Hauck SM, Pouw RB, Kuijpers TW, Wouters D, Ehrenstein B, Enzmann V, Zipfel PF, Skerka C, Pauly D (2016) Complement Regulator FHR-3 Is Elevated either Locally or Systemically in a Selection of Autoimmune Diseases. Front Immunol 7:542
Scheerlinck E, Van Steendam K, Daled S, Govaert E, Vossaert L, Meert P, Van Nieuwerburgh F, Van Soom A, Peelman L, De Sutter P, Heindryckx B, Dhaenens M, Deforce D (2016) Assessing the impact of minimizing arginine conversion in fully defined SILAC culture medium in human embryonic stem cells. Proteomics 16:2605-2614
Schmidt R, Krizsan A, Volke D, Knappe D, Hoffmann R (2016) Identification of New Resistance Mechanisms in Escherichia coli against Apidaecin 1b Using Quantitative Gel- and LC-MS-Based Proteomics. J Proteome Res 15:2607-2617
Schmidt R, Ostorhazi E, Wende E, Knappe D, Hoffmann R (2016) Pharmacokinetics and in vivo efficacy of optimized oncocin derivatives. J Antimicrob Chemother 71:1003-1011
Schottler S, Klein K, Landfester K, Mailander V (2016) Protein source and choice of anticoagulant decisively affect nanoparticle protein corona and cellular uptake. Nanoscale 8:5526-5536
Sebe-Pedros A, Pena MI, Capella-Gutierrez S, Anto M, Gabaldon T, Ruiz-Trillo I, Sabido E (2016) High-Throughput Proteomics Reveals the Unicellular Roots of Animal Phosphosignaling and Cell Differentiation. Dev Cell 39:186-197
Sidhu VK, Huang BX, Desai A, Kevala K, Kim HY (2016) Role of DHA in aging-related changes in mouse brain synaptic plasma membrane proteome. Neurobiol Aging 41:73-85
Singh S, Dubey VK (2016) Quantitative Proteome Analysis of Leishmania donovani under Spermidine Starvation. PLoS One 11:e0154262
Solari FA, Mattheij NJ, Burkhart JM, Swieringa F, Collins PW, Cosemans JM, Sickmann A, Heemskerk JW, Zahedi RP (2016) Combined Quantification of the Global Proteome, Phosphoproteome, and Proteolytic Cleavage to Characterize Altered Platelet Functions in the Human Scott Syndrome. Mol Cell Proteomics 15:3154-3169
Spencer WJ, Pearring JN, Salinas RY, Loiselle DR, Skiba NP, Arshavsky VY (2016) Progressive Rod-Cone Degeneration (PRCD) Protein Requires N-Terminal S-Acylation and Rhodopsin Binding for Photoreceptor Outer Segment Localization and Maintaining Intracellular Stability. Biochemistry 55:5028-5037
Standish AJ, Teh MY, Tran EN, Doyle MT, Baker PJ, Morona R (2016) Unprecedented Abundance of Protein Tyrosine Phosphorylation Modulates Shigella flexneri Virulence. J Mol Biol 428:4197-4208
Takahashi D, Kawamura Y, Uemura M (2016) Cold acclimation is accompanied by complex responses of glycosylphosphatidylinositol (GPI)-anchored proteins in Arabidopsis. J Exp Bot 67:5203-5215
Thorpe CT, Peffers MJ, Simpson D, Halliwell E, Screen HR, Clegg PD (2016) Anatomical heterogeneity of tendon: Fascicular and interfascicular tendon compartments have distinct proteomic composition. Sci Rep 6:20455
Tikka S, Monogioudi E, Gotsopoulos A, Soliymani R, Pezzini F, Scifo E, Uusi-Rauva K, Tyynela J, Baumann M, Jalanko A, Simonati A, Lalowski M (2016) Proteomic Profiling in the Brain of CLN1 Disease Model Reveals Affected Functional Modules. Neuromolecular Med 18:109-133
Treindl F, Ruprecht B, Beiter Y, Schultz S, Dottinger A, Staebler A, Joos TO, Kling S, Poetz O, Fehm T, Neubauer H, Kuster B, Templin MF (2016) A bead-based western for high-throughput cellular signal transduction analyses. Nat Commun 7:12852
Tyleckova J, Valekova I, Zizkova M, Rakocyova M, Marsala S, Marsala M, Gadher SJ, Kovarova H (2016) Surface N-glycoproteome patterns reveal key proteins of neuronal differentiation. J Proteomics 132:13-20
Vaidya B, Cho SY, Oh KS, Kim SH, Kim YO, Jeong EH, Nguyen TT, Kim IS, Kwon J, Kim D (2016) Effectiveness of Periodic Treatment of Quercetin against Influenza A Virus H1N1 through Modulation of Protein Expression. J Agric Food Chem 64:4416-4425
Valikangas T, Suomi T, Elo LL (2016) A systematic evaluation of normalization methods in quantitative label-free proteomics. Brief Bioinform (In press)
van den Berg CB, Duvekot JJ, Guzel C, Hansson SR, de Leeuw TG, Steegers EA, Versendaal J, Luider TM, Stoop MP (2016) Elevated levels of protein AMBP in cerebrospinal fluid of women with preeclampsia compared to normotensive pregnant women. Proteomics Clin Appl (In press)
Vanzo E, Merl-Pham J, Velikova V, Ghirardo A, Lindermayr C, Hauck SM, Bernhardt J, Riedel K, Durner J, Schnitzler JP (2016) Modulation of Protein S-Nitrosylation by Isoprene Emission in Poplar. Plant Physiol 170:1945-1961
Veit J, Sachsenberg T, Chernev A, Aicheler F, Urlaub H, Kohlbacher O (2016) LFQProfiler and RNP(xl): Open-Source Tools for Label-Free Quantification and Protein-RNA Cross-Linking Integrated into Proteome Discoverer. J Proteome Res 15:3441-3448
Verdonck R, De Haes W, Cardoen D, Menschaert G, Huhn T, Landuyt B, Baggerman G, Boonen K, Wenseleers T, Schoofs L (2016) Fast and Reliable Quantitative Peptidomics with labelpepmatch. J Proteome Res 15:1080-1089
Verrastro I, Tveen-Jensen K, Woscholski R, Spickett CM, Pitt AR (2016) Reversible oxidation of phosphatase and tensin homolog (PTEN) alters its interactions with signaling and regulatory proteins. Free Radic Biol Med 90:24-34
Walker A, Russmann V, Deeg CA, von Toerne C, Kleinwort KJ, Szober C, Rettenbeck ML, von Ruden EL, Goc J, Ongerth T, Boes K, Salvamoser JD, Vezzani A, Hauck SM, Potschka H (2016) Proteomic profiling of epileptogenesis in a rat model: Focus on inflammation. Brain Behav Immun 53:138-158
Wang X, Pan T (2016) Methionine Mistranslation Bypasses the Restraint of the Genetic Code to Generate Mutant Proteins with Distinct Activities. PLoS Genet 12:e1005832
Welinder KG, Hansen R, Overgaard MT, Brohus M, Sonderkaer M, von Bergen M, Rolle-Kampczyk U, Otto W, Lindahl TL, Arinell K, Evans AL, Swenson JE, Revsbech IG, Frobert O (2016) Biochemical Foundations of Health and Energy Conservation in Hibernating Free-ranging Subadult Brown Bear Ursus arctos. J Biol Chem 291:22509-22523
Winter L, Wittig I, Peeva V, Eggers B, Heidler J, Chevessier F, Kley RA, Barkovits K, Strecker V, Berwanger C, Herrmann H, Marcus K, Kornblum C, Kunz WS, Schroder R, Clemen CS (2016) Mutant desmin substantially perturbs mitochondrial morphology, function and maintenance in skeletal muscle tissue. Acta Neuropathol 132:453-473
Yadetie F, Bjorneklett S, Garberg HK, Oveland E, Berven F, Goksoyr A, Karlsen OA (2016) Quantitative analyses of the hepatic proteome of methylmercury-exposed Atlantic cod (Gadus morhua) suggest oxidative stress-mediated effects on cellular energy metabolism. BMC Genomics 17:554
Ye J, Zhang H, He W, Zhu B, Zhou D, Chen Z, Ashraf U, Wei Y, Liu Z, Fu ZF, Chen H, Cao S (2016) Quantitative phosphoproteomic analysis identifies the critical role of JNK1 in neuroinflammation induced by Japanese encephalitis virus. Sci Signal 9:ra98
Yilmaz Z, Eralp Inan O, Kocaturk M, Baykal AT, Hacariz O, Hatipoglu I, Tvarijonaviciute A, Cansev M, Ceron J, Ulus IH (2016) Changes in serum proteins after endotoxin administration in healthy and choline-treated calves. BMC Vet Res 12:210
Zhang Y, Crofton EJ, Fan X, Li D, Kong F, Sinha M, Luxon BA, Spratt HM, Lichti CF, Green TA (2016) Convergent transcriptomics and proteomics of environmental enrichment and cocaine identifies novel therapeutic strategies for addiction. Neuroscience 339:254-266
Zhang Z, Boonen K, Ferrari P, Schoofs L, Janssens E, van Noort V, Rolland F, Geuten K (2016) UV crosslinked mRNA-binding proteins captured from leaf mesophyll protoplasts. Plant Methods 12:42
Zhao F, Elkelish A, Durner J, Lindermayr C, Winkler JB, Rusmall io RF, Behrendt H, Traidl-Hoffmann C, Holzinger A, Kofler W, Braun P, von Toerne C, Hauck SM, Ernst D, Frank U (2016) Common ragweed (Ambrosia artemisiifolia L.): allergenicity and molecular characterization of pollen after plant exposure to elevated NO2. Plant Cell Environ 39:147-164
2015
Aird SD, Aggarwal S, Villar-Briones A, Tin MM, Terada K, Mikheyev AS (2015) Snake venoms are integrated systems, but abundant venom proteins evolve more rapidly. BMC Genomics 16:647
Akhtar MZ, Huang H, Kaisar M, Lo Faro ML, Robolledo R, Morten K, Heather LC, Dona A, Leuvenink HG, Fuggle SV, Kessler BM, Pugh CW, Ploeg RJ (2015) Using an integrated -omics approach to identify key cellular processes that are disturbed in the kidney following brain death. Am J Transplant (In press)
Almaiman_A, Abdullah_R, Abdul_AB, Allauddin_ZN, Alaiya_AA, Eid_EEM, Shinwari_Z, Juhani_GA, Rasheed_W, Bakshi_N, Owaidah_T, Ahmed_SO, Aljurf_M (2015) The Potential of Protein Expression Profiles in Categorization Risks for Acute Myeloid Leukemia: A Pilot Study. Int Journal of Hematology and Oncology 25:205-214
Antberg_L (2015) Pathway-centric analysis of the DNA damageresponse to chemotherapeutic agents in two breast cell lines. eupa open proteomics (In press)
Bao K, Bostanci N, Selevsek N, Thurnheer T, Belibasakis GN (2015) Quantitative Proteomics Reveal Distinct Protein Regulations Caused by Aggregatibacter actinomycetemcomitans within Subgingival Biofilms. PLoS One 10:e0119222
Barjaktarovic Z, Kempf SJ, Sriharshan A, Merl-Pham J, Atkinson MJ, Tapio S (2015) Ionizing radiation induces immediate protein acetylation changes in human cardiac microvascular endothelial cells. J Radiat Res (In press)
Behmoaras J, Diaz AG, Venda L, Ko JH, Srivastava P, Montoya A, Faull P, Webster Z, Moyon B, Pusey CD, Abraham DJ, Petretto E, Cook TH, Aitman TJ (2015) Macrophage epoxygenase determines a profibrotic transcriptome signature. J Immunol 194:4705-4716
Benevento M, Tonge PD, Puri MC, Nagy A, Heck AJ, Munoz J (2015) Fluctuations in histone H4 isoforms during cellular reprogramming monitored by middle-down proteomics. Proteomics 15:3219-3231
Beretov J, Wasinger VC, Millar EK, Schwartz P, Graham PH, Li Y (2015) Proteomic Analysis of Urine to Identify Breast Cancer Biomarker Candidates Using a Label-Free LC-MS/MS Approach. PLoS One 10:e0141876
Bhandari S, Gupta P, Quinn P, Sandhu J, Hakimi A, Jones D, Ng L (2015) Pleiotropic effects of statins in hypercholesterolaemia: a prospective observational study using a lipoproteomic based approach. Lancet 385 Suppl 1:S21
Bi Z, Merl-Pham J, Uehlein N, Zimmer I, Muhlhans S, Aichler M, Walch AK, Kaldenhoff R, Palme K, Schnitzler JP, Block K (2015) RNAi-mediated downregulation of poplar plasma membrane intrinsic proteins (PIPs) changes plasma membrane proteome composition and affects leaf physiology. J Proteomics 128:321-332
Birse K, Arnold KB, Novak RM, McCorrister S, Shaw S, Westmacott GR, Ball TB, Lauffenburger DA, Burgener A (2015) Molecular Signatures of Immune Activation and Epithelial Barrier Remodeling Are Enhanced during the Luteal Phase of the Menstrual Cycle: Implications for HIV Susceptibility. J Virol 89:8793-8805
Bivol LM, Iversen BM, Hultstrom M, Wallace PW, Reed RK, Wiig H, Tenstad O (2015) Unilateral renal ischemia in rats induces a rapid secretion of inflammatory markers to renal lymph and increased peritubular capillary permeability. J Physiol (In press)
Bostanci N, Bao K, Wahlander A, Grossmann J, Thurnheer T, Belibasakis GN (2015) Secretome of gingival epithelium in response to subgingival biofilms. Mol Oral Microbiol 30:323-335
Bracht T, Schweinsberg V, Trippler M, Kohl M, Ahrens M, Padden J, Naboulsi W, Barkovits K, Megger DA, Eisenacher M, Borchers CH, Schlaak JF, Hoffmann AC, Weber F, Baba HA, Meyer HE, Sitek B (2015) Analysis of disease-associated protein expression using quantitative proteomics-fibulin-5 is expressed in association with hepatic fibrosis. J Proteome Res 14:2278-2286
Burch TC, Isaac G, Booher CL, Rhim JS, Rainville P, Langridge J, Baker A, Nyalwidhe JO (2015) Comparative Metabolomic and Lipidomic Analysis of Phenotype Stratified Prostate Cells. PLoS One 10:e0134206
Burch TC, Morris MA, Campbell-Thompson M, Pugliese A, Nadler JL, Nyalwidhe JO (2015) Proteomic Analysis of Disease Stratified Human Pancreas Tissue Indicates Unique Signature of Type 1 Diabetes. PLoS One 10:e0135663
Butt AQ, McArdle A, Gibson DS, FitzGerald O, Pennington SR (2015) Psoriatic arthritis under a proteomic spotlight: application of novel technologies to advance diagnosis and management. Curr Rheumatol Rep 17:35
Cantley LG, Colangelo CM, Stone KL, Chung L, Belcher J, Abbott T, Cantley JL, Williams KR, Parikh CR (2015) Development of a Targeted Urine Proteome Assay for Kidney Diseases. Proteomics Clin Appl (in press)
Casadei N, Sood P, Ulrich T, Kieper N, Helling S, May C, Glaab E, Chen J, Nuber S, Marcus K, Rapaport D, Ott T, Riess O, Kruger R, Fitzgerald JC (2015) Mitochondrial defects and neurodegeneration in mice overexpressing wild-type or G399S mutant HtrA2. Hum Mol Genet (In press)
Casanovas A, Sprenger RR, Tarasov K, Ruckerbauer DE, Hannibal-Bach HK, Zanghellini J, Jensen ON, Ejsing CS (2015) Quantitative analysis of proteome and lipidome dynamics reveals functional regulation of global lipid metabolism. Chem Biol 22:412-425
Castelli LM, Talavera D, Kershaw CJ, Mohammad-Qureshi SS, Costello JL, Rowe W, Sims PF, Grant CM, Hubbard SJ, Ashe MP, Pavitt GD (2015) The 4E-BP Caf20p Mediates Both eIF4E-Dependent and Independent Repression of Translation. PLoS Genet 11:e1005233
Clabaut A, Grare C, Leger T, Hardouin P, Broux O (2015) Variations of secretome profiles according to conditioned medium preparation: The example of human mesenchymal stem cell-derived adipocytes. Electrophoresis 36:2587-2593
Colangelo CM, Shifman M, Cheung KH, Stone KL, Carriero NJ, Gulcicek EE, Lam TT, Wu T, Bjornson RD, Bruce C, Nairn AC, Rinehart J, Miller PL, Williams KR (2015) YPED: An Integrated Bioinformatics Suite and Database for Mass Spectrometry-based Proteomics Research. Genomics Proteomics Bioinformatics 13:25-35
Dhaenens M, Glibert P, Meert P, Vossaert L, Deforce D (2015) Histone proteolysis: a proposal for categorization into 'clipping' and 'degradation'. Bioessays 37:70-79
Di Luca A, Henry M, Meleady P, O'Connor R (2015) Label-free LC-MS analysis of HER2+ breast cancer cell line response to HER2 inhibitor treatment. Daru 23:40
Dorosz SA, Ginolhac A, Kahne T, Naumann M, Sauter T, Salsmann A, Bueb JL (2015) Role of Calprotectin as a Modulator of the IL27-Mediated Proinflammatory Effect on Endothelial Cells. Mediators Inflamm 2015:737310
Dowling P, Hughes DJ, Larkin AM, Meiller J, Henry M, Meleady P, Lynch V, Pardini B, Naccarati A, Levy M, Vodicka P, Neary P, Clynes M (2015) Elevated levels of 14-3-3 proteins, serotonin, gamma enolase and pyruvate kinase identified in clinical samples from patients diagnosed with colorectal cancer. Clin Chim Acta 441:133-141
Espino JA, Mali VS, Jones LM (2015) In Cell Footprinting Coupled with Mass Spectrometry for the Structural Analysis of Proteins in Live Cells. Anal Chem 87:7971-7978
Fesenko IA, Arapidi GP, Skripnikov AY, Alexeev DG, Kostryukova ES, Manolov AI, Altukhov IA, Khazigaleeva RA, Seredina AV, Kovalchuk SI, Ziganshin RH, Zgoda VG, Novikova SE, Semashko TA, Slizhikova DK, Ptushenko VV, Gorbachev AY, Govorun VM, Ivanov VT (2015) Specific pools of endogenous peptides are present in gametophore, protonema, and protoplast cells of the moss Physcomitrella patens. BMC Plant Biol 15:87
Finamore F, Priego-Capote F, Nolli S, Zufferey A, Fontana P, Sanchez JC (2015) Characterisation of the influences of aspirin-acetylation and glycation on human plasma proteins. J Proteomics 114:125-135
Foe IT, Child MA, Majmudar JD, Krishnamurthy S, van der Linden WA, Ward GE, Martin BR, Bogyo M (2015) Global Analysis of Palmitoylated Proteins in Toxoplasma gondii. Cell Host Microbe 18:501-511
Garelli A, Heredia F, Casimiro AP, Macedo A, Nunes C, Garcez M, Dias AR, Volonte YA, Uhlmann T, Caparros E, Koyama T, Gontijo AM (2015) Dilp8 requires the neuronal relaxin receptor Lgr3 to couple growth to developmental timing. Nat Commun 6:8732
Gentzel M, Schille C, Rauschenberger V, Schambony A (2015) Distinct functionality of dishevelled isoforms on Ca2+/calmodulin-dependent protein kinase 2 (CamKII) in Xenopus gastrulation. Mol Biol Cell 26:966-977
Glatter T, Ahrne E, Schmidt A (2015) Comparison of Different Sample Preparation Protocols Reveals Lysis Buffer-Specific Extraction Biases in Gram-Negative Bacteria and Human Cells. J Proteome Res 14:4472-4485
Glibert P, Meert P, Van Steendam K, Van Nieuwerburgh F, De Coninck D, Martens L, Dhaenens M, Deforce D (2015) Phospho-iTRAQ: assessing isobaric labels for the large-scale study of phosphopeptide stoichiometry. J Proteome Res 14:839-849
Gomez-Sanchez JA, Carty L, Iruarrizaga-Lejarreta M, Palomo-Irigoyen M, Varela-Rey M, Griffith M, Hantke J, Macias-Camara N, Azkargorta M, Aurrekoetxea I, De Juan VG, Jefferies HB, Aspichueta P, Elortza F, Aransay AM, Martinez-Chantar ML, Baas F, Mato JM, Mirsky R, Woodhoo A, Jessen KR (2015) Schwann cell autophagy, myelinophagy, initiates myelin clearance from injured nerves. J Cell Biol 210:153-168
Graessel A, Hauck SM, von Toerne C, Kloppmann E, Goldberg T, Koppensteiner H, Schindler M, Knapp B, Krause L, Dietz K, Schmidt-Weber CB, Suttner K (2015) A Combined Omics Approach to Generate the Surface Atlas of Human Naive CD4+ T Cells during Early T-Cell Receptor Activation. Mol Cell Proteomics 14:2085-2102
Graham DB, Becker CE, Doan A, Goel G, Villablanca EJ, Knights D, Mok A, Ng AC, Doench JG, Root DE, Clish CB, Xavier RJ (2015) Functional genomics identifies negative regulatory nodes controlling phagocyte oxidative burst. Nat Commun 6:7838
Grison MS, Brocard L, Fouillen L, Nicolas W, Wewer V, Dormann P, Nacir H, Benitez-Alfonso Y, Claverol S, Germain V, Boutte Y, Mongrand S, Bayer EM (2015) Specific membrane lipid composition is important for plasmodesmata function in Arabidopsis. Plant Cell 27:1228-1250
Grosche A, Hauser A, Lepper MF, Mayo R, von Toerne C, Merl-Pham J, Hauck SM (2015) The proteome of native adult Muller glial cells from murine retina. Mol Cell Proteomics (in press)
Harrington TD, Mohamed A, Tran VN, Biria S, Gargouri M, Park JJ, Gang DR, Beyenal H (2015) Neutral red-mediated microbial electrosynthesis by Escherichia coli, Klebsiella pneumoniae, and Zymomonas mobilis. Bioresour Technol 195:57-65
Haslene-Hox H, Oveland E, Woie K, Salvesen HB, Tenstad O, Wiig H (2015) Distribution volumes of macromolecules in human ovarian and endometrial cancers--effects of extracellular matrix structure. Am J Physiol Heart Circ Physiol 308:H18-28
Heywood WE, Camuzeaux S, Doykov I, Patel N, Preece RL, Footitt E, Cleary M, Clayton P, Grunewald S, Abulhoul L, Chakrapani A, Sebire NJ, Hindmarsh P, de Koning TJ, Heales S, Burke D, Gissen P, Mills K (2015) Proteomic Discovery and Development of a Multiplexed Targeted MRM-LC-MS/MS Assay for Urine Biomarkers of Extracellular Matrix Disruption in Mucopolysaccharidoses I, II, and VI. Anal Chem 87:12238-12244
Heywood WE, Galimberti D, Bliss E, Sirka E, Paterson RW, Magdalinou NK, Carecchio M, Reid E, Heslegrave A, Fenoglio C, Scarpini E, Schott JM, Fox NC, Hardy J, Bahtia K, Heales S, Sebire NJ, Zetterburg H, Mills K (2015) Identification of novel CSF biomarkers for neurodegeneration and their validation by a high-throughput multiplexed targeted proteomic assay. Mol Neurodegener 10:64
Hofener M, Pachl F, Kuster B, Sewald N (2015) Inhibitor-based affinity probes for the investigation of JAK signaling pathways. Proteomics 15:3066-3074
Holland A, Dowling P, Meleady P, Henry M, Zweyer M, Mundegar RR, Swandulla D, Ohlendieck K (2015) Label-free mass spectrometric analysis of the mdx-4cv diaphragm identifies the matricellular protein periostin as a potential factor involved in dystrophinopathy-related fibrosis. Proteomics 15:2318-2331
Hyder CL, Kemppainen K, Isoniemi KO, Imanishi SY, Goto H, Inagaki M, Fazeli E, Eriksson JE, Tornquist K (2015) Sphingolipids inhibit vimentin-dependent cell migration. J Cell Sci 128:2057-2069
Ikoma-Seki K, Nakamura K, Morishita S, Ono T, Sugiyama K, Nishino H, Hirano H, Murakoshi M (2015) Role of LRP1 and ERK and cAMP Signaling Pathways in Lactoferrin-Induced Lipolysis in Mature Rat Adipocytes. PLoS One 10:e0141378
Inagaki T, Iwasaki S, Matsumura Y, Kawamura T, Tanaka T, Abe Y, Yamasaki A, Tsurutani Y, Yoshida A, Chikaoka Y, Nakamura K, Magoori K, Nakaki R, Osborne TF, Fukami K, Aburatani H, Kodama T, Sakai J (2015) The FBXL10/KDM2B scaffolding protein associates with novel polycomb repressive complex-1 to regulate adipogenesis. J Biol Chem 290:4163-4177
Ittig SJ, Schmutz C, Kasper CA, Amstutz M, Schmidt A, Sauteur L, Vigano MA, Low SH, Affolter M, Cornelis GR, Nigg EA, Arrieumerlou C (2015) A bacterial type III secretion-based protein delivery tool for broad applications in cell biology. J Cell Biol 211:913-931
Jaafar SN, Coelho AV, Sheehan D (2015) Redox proteomic analysis of mytilus edulis gills: effects of the pharmaceutical diclofenac on a non-target organism. Drug Test Anal 7:957-966
Jackson AP, Goyard S, Xia D, Foth BJ, Sanders M, Wastling JM, Minoprio P, Berriman M (2015) Global Gene Expression Profiling through the Complete Life Cycle of Trypanosoma vivax. PLoS Negl Trop Dis 9:e0003975
Jeong EH, Vaidya B, Cho SY, Park MA, Kaewintajuk K, Kim SR, Oh MJ, Choi JS, Kwon J, Kim D (2015) Identification of regulators of the early stage of viral hemorrhagic septicemia virus infection during curcumin treatment. Fish Shellfish Immunol 45:184-193
Kauko O, Laajala TD, Jumppanen M, Hintsanen P, Suni V, Haapaniemi P, Corthals G, Aittokallio T, Westermarck J, Imanishi SY (2015) Label-free quantitative phosphoproteomics with novel pairwise abundance normalization reveals synergistic RAS and CIP2A signaling. Sci Rep 5:13099
Kellett-Clarke H, Stegmann M, Barclay AN, Metcalfe C (2015) CD44 Binding to Hyaluronic Acid Is Redox Regulated by a Labile Disulfide Bond in the Hyaluronic Acid Binding Site. PLoS One 10:e0138137
Kershaw CJ, Costello JL, Castelli LM, Talavera D, Rowe W, Sims PF, Ashe MP, Hubbard SJ, Pavitt GD, Grant CM (2015) The yeast La related protein Slf1p is a key activator of translation during the oxidative stress response. PLoS Genet 11:e1004903
Kershaw CJ, Costello JL, Talavera D, Rowe W, Castelli LM, Sims PF, Grant CM, Ashe MP, Hubbard SJ, Pavitt GD (2015) Integrated multi-omics analyses reveal the pleiotropic nature of the control of gene expression by Puf3p. Sci Rep 5:15518
Kimura A, Kurata Y, Nakabayashi J, Kagawa H, Hirano H (2015) N-Myristoylation of the Rpt2 subunit of the yeast 26S proteasome is implicated in the subcellular compartment-specific protein quality control system. J Proteomics 130:33-41
Koeck DE, Koellmeier T, Zverlov VV, Liebl W, Schwarz WH (2015) Differences in biomass degradation between newly isolated environmental strains of Clostridium thermocellum and heterogeneity in the size of the cellulosomal scaffoldin. Syst Appl Microbiol 38:424-432
Kraljevic Pavelic S, Klobucar M, Sedic M, Micek V, Gehrig P, Grossman J, Pavelic K, Vojnikovic B (2015) UV-induced retinal proteome changes in the rat model of age-related macular degeneration. Biochim Biophys Acta 1852:1833-1845
Kulej K, Avgousti DC, Weitzman MD, Garcia BA (2015) Characterization of histone post-translational modifications during virus infection using mass spectrometry-based proteomics. Methods 90:8-20
Kurbasic E, Sjostrom M, Krogh M, Folkesson E, Grabau D, Hansson K, Ryden L, Waldemarson S, James P, Nimeus E (2015) Changes in glycoprotein expression between primary breast tumour and synchronous lymph node metastases or asynchronous distant metastases. Clin Proteomics 12:13
Law KP, Han TL, Tong C, Baker PN (2015) Mass spectrometry-based proteomics for pre-eclampsia and preterm birth. Int J Mol Sci 16:10952-10985
Leger T, Garcia C, Ounissi M, Lelandais G, Camadro JM (2015) The metacaspase (Mca1p) has a dual role in farnesol-induced apoptosis in Candida albicans. Mol Cell Proteomics 14:93-108
Lemeer S, Gholami AM, Wu Z, Kuster B (2015) Quantitative proteome profiling of human myoma and myometrium tissue reveals kinase expression signatures with potential for therapeutic intervention. Proteomics 15:356-364
Lento S, Brioschi M, Barcella S, Nasim MT, Ghilardi S, Barbieri SS, Tremoli E, Banfi C (2015) Proteomics of tissue factor silencing in cardiomyocytic cells reveals a new role for this coagulation factor in splicing machinery control. J Proteomics 119:75-89
Leroy C, Shen Q, Strande V, Meyer R, McLaughlin ME, Lezan E, Bentires-Alj M, Voshol H, Bonenfant D, Alex Gaither L (2015) CUB-domain-containing protein 1 overexpression in solid cancers promotes cancer cell growth by activating Src family kinases. Oncogene 34:5593-5598
Li M, Zhao M, Gao Y (2015) Effect of transient blood glucose increases after oral glucose intake on the human urinary proteome. Proteomics Clin Appl 9:618-622
Lichti_CF, Wildburger_NC, Shavkunov_AS, Mostovenko_E, Liu_H, Sulman_EP, Nilsson_CL (2015) The proteomic landscape of glioma stem-like cells. EuPA Open Proteomics (in press)
Ligat L, Saint-Laurent N, El-Mrani A, Gigoux V, Al Saati T, Tomasini R, Nigri J, Dejean S, Pont F, Baer R, Guillermet-Guibert J, Cordelier P, Lopez F, Dufresne M (2015) Pancreatic preneoplastic lesions plasma signatures and biomarkers based on proteome profiling of mouse models. Br J Cancer 113:1590-1598
Lindqvist J, Imanishi SY, Torvaldson E, Malinen M, Remes M, Orn F, Palvimo JJ, Eriksson JE (2015) Cyclin-dependent kinase 5 acts as a critical determinant of AKT-dependent proliferation and regulates differential gene expression by the androgen receptor in prostate cancer cells. Mol Biol Cell 26:1971-1984
Liu H, Zhang L, Zhao B, Zhang Z, Qin L, Zhang Q, Wang Q, Lu Z, Gao X (2015) Hypothalamus metabolomic profiling to elucidate the tissue-targeted biochemical basis of febrile response in yeast-induced pyrexia rats. Chem Biol Interact 231:61-70
Lorber B, Chew DJ, Hauck SM, Chong RS, Fawcett JW, Martin KR (2015) Retinal glia promote dorsal root ganglion axon regeneration. PLoS One 10:e0115996
Loukovaara S, Nurkkala H, Tamene F, Gucciardo E, Liu X, Repo P, Lehti K, Varjosalo M (2015) Quantitative Proteomics Analysis of Vitreous Humor from Diabetic Retinopathy Patients. J Proteome Res 14:5131-5143
Luan R, Cheng H, Li L, Zhao Q, Liu H, Wu Z, Zhao L, Yang J, Hao J, Yin Z (2015) Maternal Lipopolysaccharide Exposure Promotes Immunological Functional Changes in Adult Offspring CD4+ T Cells. Am J Reprod Immunol 73:522-535
Mackay GM, Zheng L, van den Broek NJ, Gottlieb E (2015) Analysis of Cell Metabolism Using LC-MS and Isotope Tracers. Methods Enzymol 561:171-196
Meert P, Govaert E, Scheerlinck E, Dhaenens M, Deforce D (2015) Pitfalls in histone propionylation during bottom-up mass spectrometry analysis. Proteomics 15:2966-2971
Milleret V, Buzzi S, Gehrig P, Ziogas A, Grossmann J, Schilcher K, Zinkernagel AS, Zucker A, Ehrbar M (2015) Protein adsorption steers blood contact activation on engineered cobalt chromium alloy oxide layers. Acta Biomater 24:343-351
Millet C, Ausiannikava D, Le Bihan T, Granneman S, Makovets S (2015) Cell populations can use aneuploidy to survive telomerase insufficiency. Nat Commun 6:8664
Miroshnichenko Iu V, Petushkova NA, Moskaleva NE, Teryaeva NB, Zgoda VG, Ilgisonis EV, Belyaev AY (2015) [The possibility of using PlasmaDeepDive MRM panel in clinical diagnostics]. Biomed Khim 61:272-278
Molin S, Merl J, Dietrich KA, Regauer M, Flaig M, Letule V, Saucke T, Herzinger T, Ruzicka T, Hauck SM (2015) The hand eczema proteome: imbalance of epidermal barrier proteins. Br J Dermatol 172:994-1001
Moloney NM, Owens RA, Meleady P, Henry M, Dolan S, Mulvihill E, Clynes M, Doyle S (2015) The iron-responsive microsomal proteome of Aspergillus fumigatus. J Proteomics (In press)
Moulder R, Bhosale SD, Erkkila T, Laajala E, Salmi J, Nguyen EV, Kallionpaa H, Mykkanen J, Vaha-Makila M, Hyoty H, Veijola R, Ilonen J, Simell T, Toppari J, Knip M, Goodlett DR, Lahdesmaki H, Simell O, Lahesmaa R (2015) Serum proteomes distinguish children developing type 1 diabetes in a cohort with HLA-conferred susceptibility. Diabetes 64:2265-2278
Mulek M, Fekete A, Wiest J, Holzgrabe U, Mueller MJ, Hogger P (2015) Profiling a gut microbiota-generated catechin metabolite's fate in human blood cells using a metabolomic approach. J Pharm Biomed Anal 114:71-81
Munday DC, Wu W, Smith N, Fix J, Noton SL, Galloux M, Touzelet O, Armstrong SD, Dawson JM, Aljabr W, Easton AJ, Rameix-Welti MA, de Oliveira AP, Simabuco FM, Ventura AM, Hughes DJ, Barr JN, Fearns R, Digard P, Eleouet JF, Hiscox JA (2015) Interactome analysis of the human respiratory syncytial virus RNA polymerase complex identifies protein chaperones as important cofactors that promote L-protein stability and RNA synthesis. J Virol 89:917-930
Murphy S, Henry M, Meleady P, Zweyer M, Mundegar RR, Swandulla D, Ohlendieck K (2015) Simultaneous Pathoproteomic Evaluation of the Dystrophin-Glycoprotein Complex and Secondary Changes in the mdx-4cv Mouse Model of Duchenne Muscular Dystrophy. Biology (Basel) 4:397-423
Murphy S, Ohlendieck K (2015) Proteomic pro/ling of the HSPB chaperonome: Mass spectrometricidenti/cation of small heat shock proteins in stressed skeletal muscles. JOURNAL OF INTEGRATED OMICS 5:17-29
Naboulsi W, Megger DA, Bracht T, Kohl M, Turewicz M, Eisenacher M, Voss DM, Schlaak JF, Hoffmann AC, Weber F, Baba HA, Meyer HE, Sitek B (2015) Quantitative Tissue Proteomics Analysis Reveals Versican as Potential Biomarker for Early-Stage Hepatocellular Carcinoma. J Proteome Res 15:38-47
Nassar AF, Williams BJ, Yaworksy DC, Patel V, Rusling JF (2015) Rapid label-free profiling of oral cancer biomarker proteins using nanoUPLC-Q-TOF ion mobility mass spectrometry. Proteomics Clin Appl (In press)
Nguyen EV, Imanishi SY, Haapaniemi P, Yadav A, Saloheimo M, Corthals GL, Pakula T (2015) Quantitative site-specific phosphoproteomics of Trichoderma reesei signaling pathways upon induction of hydrolytic enzyme production. J Proteome Res (In press)
Niksirat H, James P, Andersson L, Kouba A, Kozak P (2015) Label-free protein quantification in freshly ejaculated versus post-mating spermatophores of the noble crayfish Astacus astacus. J Proteomics 123:70-77
Nishikawa K, Iwaya K, Kinoshita M, Fujiwara Y, Akao M, Sonoda M, Thiruppathi S, Suzuki T, Hiroi S, Seki S, Sakamoto T (2015) Resveratrol increases CD68(+) Kupffer cells colocalized with adipose differentiation-related protein and ameliorates high-fat-diet-induced fatty liver in mice. Mol Nutr Food Res 59:1155-1170
O'Callaghan PM, Berthelot ME, Young RJ, Graham JW, Racher AJ, Aldana D (2015) Diversity in host clone performance within a Chinese hamster ovary cell line. Biotechnol Prog 31:1187-1200Oveland E, Muth T, Rapp E, Martens L, Berven FS, Barsnes H (2015) Viewing the proteome: How to visualize proteomics data? Proteomics 15:1341-1355
Oveland E, Muth T, Rapp E, Martens L, Berven FS, Barsnes H (2015) Viewing the proteome: How to visualize proteomics data? Proteomics 15:1341-1355
Padden J, Ahrens M, Kalsch J, Bertram S, Megger DA, Bracht T, Eisenacher M, Kocabayoglu P, Meyer HE, Sipos B, Baba HA, Sitek B (2015) Immunohistochemical Markers Distinguishing Cholangiocellular Carcinoma from Pancreatic Ductal Adenocarcinoma Discovered by Proteomic Analysis of Microdissected Cells. Mol Cell Proteomics (In press)
Peffers MJ, McDermott B, Clegg PD, Riggs CM (2015) Comprehensive protein profiling of synovial fluid in osteoarthritis following protein equalization. Osteoarthritis Cartilage 23:1204-1213
Ramm S, Morissey B, Hernandez B, Rooney C, Pennington SR, Mally A (2015) Application of a discovery to targeted LC-MS proteomics approach to identify deregulated proteins associated with idiosyncratic liver toxicity in a rat model of LPS/diclofenac co-administration. Toxicology 331:100-111
Ramm SA, Edward DA, Claydon AJ, Hammond DE, Brownridge P, Hurst JL, Beynon RJ, Stockley P (2015) Sperm competition risk drives plasticity in seminal fluid composition. BMC Biol 13:87
Rani ST, Takahashi D, Uemura M, Podile AR (2015) Proteins Associated with Oxidative Burst and Cell Wall Strengthening Accumulate During Citrus-Xanthomonas Non-Host Interaction. Plant Mol Biol Rep 33:1349–1360
Rani TS, Durgeshwar P, Podile AR (2015) Accumulation of transcription factors and cell signaling-related proteins in the nucleus during citrus-Xanthomonas interaction. J Plant Physiol 184:20-27
Rattay S, Trilling M, Megger DA, Sitek B, Meyer HE, Hengel H, Le-Trilling VT (2015) The Canonical Immediate Early 3 Gene Product pIE611 of Mouse Cytomegalovirus Is Dispensable for Viral Replication but Mediates Transcriptional and Posttranscriptional Regulation of Viral Gene Products. J Virol 89:8590-8598
Reis H, Putter C, Megger DA, Bracht T, Weber F, Hoffmann AC, Bertram S, Wohlschlager J, Hagemann S, Eisenacher M, Scherag A, Schlaak JF, Canbay A, Meyer HE, Sitek B, Baba HA (2015) A structured proteomic approach identifies 14-3-3Sigma as a novel and reliable protein biomarker in panel based differential diagnostics of liver tumors. Biochim Biophys Acta 1854:641-650
Richarme G, Mihoub M, Dairou J, Bui LC, Leger T, Lamouri A (2015) Parkinsonism-associated protein DJ-1/Park7 is a major protein deglycase that repairs methylglyoxal- and glyoxal-glycated cysteine, arginine, and lysine residues. J Biol Chem 290:1885-1897
Rombouts I, Lambrecht MA, Carpentier SC, Delcour JA (2015) Identification of lanthionine and lysinoalanine in heat-treated wheat gliadin and bovine serum albumin using tandem mass spectrometry with higher-energy collisional dissociation. Amino Acids (In press)
Rotival M, Ko JH, Srivastava PK, Kerloc'h A, Montoya A, Mauro C, Faull P, Cutillas PR, Petretto E, Behmoaras J (2015) Integrating phosphoproteome and transcriptome reveals new determinants of macrophage multinucleation. Mol Cell Proteomics 14:484-498
Rouillon J, Poupiot J, Zocevic A, Amor F, Leger T, Garcia C, Camadro JM, Wong B, Pinilla R, Cosette J, Coenen-Stass AM, McClorey G, Roberts TC, Wood MJ, Servais L, Udd B, Voit T, Richard I, Svinartchouk F (2015) Serum proteomic profiling reveals fragments of MYOM3 as potential biomarkers for monitoring the outcome of therapeutic interventions in muscular dystrophies. Hum Mol Genet 24:4916-4932
Ruprecht B, Koch H, Medard G, Mundt M, Kuster B, Lemeer S (2015) Comprehensive and reproducible phosphopeptide enrichment using iron immobilized metal ion affinity chromatography (Fe-IMAC) columns. Mol Cell Proteomics 14:205-215
Schaefer A, Neschen S, Kahle M, Sarioglu H, Gaisbauer T, Imhof A, Adamski J, Hauck SM, Ueffing M (2015) The Epoxyeicosatrienoic Acid Pathway Enhances Hepatic Insulin Signaling and Is Repressed in Insulin-Resistant Mouse Liver. Mol Cell Proteomics (In press)
Scheerlinck E, Dhaenens M, Van Soom A, Peelman L, De Sutter P, Van Steendam K, Deforce D (2015) Minimizing technical variation during sample preparation prior to label-free quantitative mass spectrometry. Anal Biochem 490:14-19
Schira J, Falkenberg H, Hendricks M, Waldera-Lupa DM, Kogler G, Meyer HE, Muller HW, Stuhler K (2015) Characterization of Regenerative Phenotype of Unrestricted Somatic Stem Cells (USSC) from Human Umbilical Cord Blood (hUCB) by Functional Secretome Analysis. Mol Cell Proteomics 14:2630-2643
Schmidt A, Kochanowski K, Vedelaar S, Ahrne E, Volkmer B, Callipo L, Knoops K, Bauer M, Aebersold R, Heinemann M (2015) The quantitative and condition-dependent Escherichia coli proteome. Nat Biotechnol 34:104-110
Shearer JJ, Wold EA, Umbaugh CS, Lichti CF, Nilsson CL, Figueiredo ML (2015) Inorganic Arsenic Related Changes in the Stromal Tumor Microenvironment in a Prostate Cancer Cell-Conditioned Media Model. Environ Health Perspect (In press)
Shen K, Sun J, Cao X, Zhou D, Li J (2015) Comparison of Different Buffers for Protein Extraction from Formalin-Fixed and Paraffin-Embedded Tissue Specimens. PLoS One 10:e0142650
Singh V, Stingl C, Stoop MP, Zeneyedpour L, Neuteboom RF, Smitt PS, Hintzen RQ, Luider TM (2015) Proteomics Urine Analysis of Pregnant Women Suffering from Multiple Sclerosis. J Proteome Res (In press)
Slade WO, Werth EG, McConnell EW, Alvarez S, Hicks LM (2015) Quantifying reversible oxidation of protein thiols in photosynthetic organisms. J Am Soc Mass Spectrom 26:631-640
Sznajder A, Pfeiffer D, Jendrossek D (2015) Comparative proteome analysis reveals four novel polyhydroxybutyrate (PHB) granule-associated proteins in Ralstonia eutropha H16. Appl Environ Microbiol 81:1847-1858
Ternette N, Block PD, Sanchez-Bernabeu A, Borthwick N, Pappalardo E, Abdul-Jawad S, Ondondo B, Charles PD, Dorrell L, Kessler BM, Hanke T (2015) Early Kinetics of the HLA Class I-Associated Peptidome of MVA.HIVconsv-Infected Cells. J Virol 89:5760-5771
Tikka S, Monogioudi E, Gotsopoulos A, Soliymani R, Pezzini F, Scifo E, Uusi-Rauva K, Tyynela J, Baumann M, Jalanko A, Simonati A, Lalowski M (2015) Proteomic Profiling in the Brain of CLN1 Disease Model Reveals Affected Functional Modules. Neuromolecular Med (In press)
Tomar D, Prajapati P, Lavie J, Singh K, Lakshmi S, Bhatelia K, Roy M, Singh R, Benard G (2015) TRIM4; a novel mitochondrial interacting RING E3 ligase, sensitizes the cells to hydrogen peroxide (H2O2) induced cell death. Free Radic Biol Med 89:1036-1048
Vallon_T (2015) Applying systems biology tools to study n-butanol degradation in Pseudomonas putida KT2440. Engineering in Life Sciences (In press)
Vanhove AC, Vermaelen W, Swennen R, Carpentier SC (2015) A look behind the screens: Characterization of the HSP70 family during osmotic stress in a non-model crop. J Proteomics 119:10-20
Vaudel M, Burkhart JM, Zahedi RP, Oveland E, Berven FS, Sickmann A, Martens L, Barsnes H (2015) PeptideShaker enables reanalysis of MS-derived proteomics data sets. Nat Biotechnol 33:22-24
Vehmas AP, Adam M, Laajala TD, Kastenmuller G, Prehn C, Rozman J, Ohlsson C, Fuchs H, Hrabe de Angelis M, Gailus-Durner V, Elo LL, Aittokallio T, Adamski J, Corthals G, Poutanen M, Strauss L (2015) Liver lipid metabolism is altered by increased circulating estrogen to androgen ratio in male mouse. J Proteomics 133:66-75
Verrastro I, Tveen-Jensen K, Woscholski R, Spickett CM, Pitt AR (2015) Reversible oxidation of phosphatase and tensin homolog (PTEN) alters its interactions with signaling and regulatory proteins. Free Radic Biol Med 90:24-34
Wang F, Liu K, Han L, Jiang B, Wang M, Fang X (2015) Function of a p24 Heterodimer in Morphogenesis and Protein Transport in Penicillium oxalicum. Sci Rep 5:11875
Wildburger NC, Ali SR, Hsu WC, Shavkunov AS, Nenov MN, Lichti CF, LeDuc RD, Mostovenko E, Panova-Elektronova NI, Emmett MR, Nilsson CL, Laezza F (2015) Quantitative proteomics reveals protein-protein interactions with fibroblast growth factor 12 as a component of the voltage-gated sodium channel 1.2 (nav1.2) macromolecular complex in Mammalian brain. Mol Cell Proteomics 14:1288-1300
Wojdyla K, Williamson J, Roepstorff P, Rogowska-Wrzesinska A (2015) The SNO/SOH TMT strategy for combinatorial analysis of reversible cysteine oxidations. J Proteomics 113:415-434
Xiao Y, Wu W, Gao J, Smith N, Burkard C, Xia D, Zhang M, Wang C, Archibald A, Digard P, Zhou EM, Hiscox JA (2015) Characterization of the interactome of the porcine reproductive and respiratory syndrome virus (PRRSV) NSP2 protein reveals the hyper variable region as a binding platform for association with 14-3-3 proteins. J Proteome Res (In press)
Yang L, Lian X, Zhang W, Guo J, Wang Q, Li Y, Chen Y, Yin X, Yang P, Lan F, He QY, Zhang G, Wang T (2015) Finding Missing Proteins from the Epigenetically Manipulated Human Cell with Stringent Quality Criteria. J Proteome Res 14:3645-3657
Ye F, Xin Z, Han W, Fan J, Yin B, Wu S, Yang W, Yuan J, Qiang B, Sun W, Peng X (2015) Quantitative Proteomics Analysis of the Hepatitis C Virus Replicon High-Permissive and Low-Permissive Cell Lines. PLoS One 10:e0142082
Yerlikaya A, Okur E, Baykal AT, Acilan C, Boyaci I, Ulukaya E (2015) A proteomic analysis of p53-independent induction of apoptosis by bortezomib in 4T1 breast cancer cell line. J Proteomics 113:315-325
Yeung EN, Treskes P, Martin SF, Manning JR, Dunbar DR, Rogers SM, Le Bihan T, Lockman KA, Morley SD, Hayes PC, Nelson LJ, Plevris JN (2015) Fibrinogen production is enhanced in an in-vitro model of non-alcoholic fatty liver disease: an isolated risk factor for cardiovascular events? Lipids Health Dis 14:86
Yin X, Zhang Y, Guo S, Jin H, Wang W, Yang P (2015) Large scale systematic proteomic quantification from non-metastatic to metastatic colorectal cancer. Sci Rep 5:12120
Yoo M, Bestel-Corre G, Croux C, Riviere A, Meynial-Salles I, Soucaille P (2015) A Quantitative System-Scale Characterization of the Metabolism of Clostridium acetobutylicum. MBio 6 (In press)
Yoon EJ, Nait Chabane Y, Goussard S, Snesrud E, Courvalin P, De E, Grillot-Courvalin C (2015) Contribution of Resistance-Nodulation-Cell Division Efflux Systems to Antibiotic Resistance and Biofilm Formation in Acinetobacter baumannii. MBio 6 (In press)
Zaarur N, Xu X, Lestienne P, Meriin AB, McComb M, Costello CE, Newnam GP, Ganti R, Romanova NV, Shanmugasundaram M, Silva ST, Bandeiras TM, Matias PM, Lobachev KS, Lednev IK, Chernoff YO, Sherman MY (2015) RuvbL1 and RuvbL2 enhance aggresome formation and disaggregate amyloid fibrils. Embo J 34:2363-2382
Zhang L, Li X, Hill RC, Qiu Y, Zhang W, Hansen KC, Zhao R (2015) Brr2 plays a role in spliceosomal activation in addition to U4/U6 unwinding. Nucleic Acids Res 43:3286-3297
Zhang Y, Bhamber R, Riba-Garcia I, Liao H, Unwin RD, Dowsey AW (2015) Streaming visualisation of quantitative mass spectrometry data based on a novel raw signal decomposition method. Proteomics 15:1419-1427
Zhang Y, Guo Z, Zou L, Yang Y, Zhang L, Ji N, Shao C, Sun W, Wang Y (2015) A comprehensive map and functional annotation of the normal human cerebrospinal fluid proteome. J Proteomics 119:90-99
Zhao F, Elkelish A, Durner J, Lindermayr C, Winkler JB, Rusmall io RF, Behrendt H, Traidl-Hoffmann C, Holzinger A, Kofler W, Braun P, von Toerne C, Hauck SM, Ernst D, Frank U (2015) Common ragweed (Ambrosia artemisiifolia L.): Allergenicity and molecular characterisation of pollen after plant exposure to elevated NO. Plant Cell Environ (in press)
Zhao Z, Wu F, Ding S, Sun L, Liu Z, Ding K, Lu J (2015) Label-free quantitative proteomic analysis reveals potential biomarkers and pathways in renal cell carcinoma. Tumour Biol 36:939-951
Zigdon H, Savidor A, Levin Y, Meshcheriakova A, Schiffmann R, Futerman AH (2015) Identification of a biomarker in cerebrospinal fluid for neuronopathic forms of Gaucher disease. PLoS One 10:e0120194
2014
Aasebo E, Opsahl JA, Bjorlykke Y, Myhr KM, Kroksveen AC, Berven FS (2014) Effects of blood contamination and the rostro-caudal gradient on the human cerebrospinal fluid proteome. PLoS One 9:e90429
Ademowo OS, Hernandez B, Collins E, Rooney C, Fearon U, van Kuijk AW, Tak PP, Gerlag DM, FitzGerald O, Pennington SR (2014) Discovery and confirmation of a protein biomarker panel with potential to predict response to biological therapy in psoriatic arthritis. Ann Rheum Dis (In press)
Adiguzel Z, Baykal AT, Kacar O, Yilmaz VT, Ulukaya E, Acilan C (2014) Biochemical and Proteomic Analysis of a Potential Anticancer Agent: Palladium(II) Saccharinate Complex of Terpyridine Acting through Double Strand Break Formation. J Proteome Res 13:5240-5249
Al-Moghrabi N, Nofel A, Al-Yousef N, Madkhali S, Bin Amer SM, Alaiya A, Shinwari Z, Al-Tweigeri T, Karakas B, Tulbah A, Aboussekhra A (2014) The molecular significance of methylated BRCA1 promoter in white blood cells of cancer-free females. BMC Cancer 14:830
Ansari D, Andersson R, Bauden MP, Andersson B, Connolly JB, Welinder C, Sasor A, Marko-Varga G (2014) Protein deep sequencing applied to biobank samples from patients with pancreatic cancer. J Cancer Res Clin Oncol
Armstrong SD, Babayan SA, Lhermitte-Vallarino N, Gray N, Xia D, Martin C, Kumar S, Taylor DW, Blaxter ML, Wastling JM, Makepeace BL (2014) Comparative analysis of the secretome from a model filarial nematode (Litomosoides sigmodontis) reveals maximal diversity in gravid female parasites. Mol Cell Proteomics 13:2527-2544
Astarita G, McKenzie JH, Wang B, Strassburg K, Doneanu A, Johnson J, Baker A, Hankemeier T, Murphy J, Vreeken RJ, Langridge J, Kang JX (2014) A protective lipidomic biosignature associated with a balanced omega-6/omega-3 ratio in fat-1 transgenic mice. PLoS One 9:e96221
Atrih A, Mudaliar MA, Zakikhani P, Lamont DJ, Huang JT, Bray SE, Barton G, Fleming S, Nabi G (2014) Quantitative proteomics in resected renal cancer tissue for biomarker discovery and profiling. Br J Cancer 110:1622-1633
Avanesov AS, Ma S, Pierce KA, Yim SH, Lee BC, Clish CB, Gladyshev VN (2014) Age- and diet-associated metabolome remodeling characterizes the aging process driven by damage accumulation. Elife 3:e02077
Bauer M, Ahrne E, Baron AP, Glatter T, Fava LL, Santamaria A, Nigg EA, Schmidt A (2014) Evaluation of data-dependent and -independent mass spectrometric workflows for sensitive quantification of proteins and phosphorylation sites. J Proteome Res 13:5973-5988
Ben Mlouka MA, Khemiri A, Seyer D, Hardouin J, Chan Tchi Song P, De E, Jouenne T, Cosette P (2014) Characterization of new outer membrane proteins of Pseudomonas aeruginosa using a combinatorial peptide ligand library. Anal Bioanal Chem (In press)
Bonacci T, Audebert S, Camoin L, Baudelet E, Bidaut G, Garcia M, Witzel, II, Perkins ND, Borg JP, Iovanna JL, Soubeyran P (2014) Identification of new mechanisms of cellular response to chemotherapy by tracking changes in post-translational modifications by ubiquitin and ubiquitin-like proteins. J Proteome Res 13:2478-2494
Bracht T, Hagemann S, Loscha M, Megger DA, Padden J, Eisenacher M, Kuhlmann K, Meyer HE, Baba HA, Sitek B (2014) Proteome Analysis of a Hepatocyte-Specific BIRC5 (Survivin)-Knockout Mouse Model during Liver Regeneration. J Proteome Res 13:2771-2782
Brown LM (2014) Quantitative shotgun proteomics with data-independent acquisition and traveling wave ion mobility spectrometry: a versatile tool in the life sciences. Adv Exp Med Biol 806:79-91
Buck AH, Coakley G, Simbari F, McSorley HJ, Quintana JF, Le Bihan T, Kumar S, Abreu-Goodger C, Lear M, Harcus Y, Ceroni A, Babayan SA, Blaxter M, Ivens A, Maizels RM (2014) Exosomes secreted by nematode parasites transfer small RNAs to mammalian cells and modulate innate immunity. Nat Commun 5:5488
Bui-Nguyen TM, Dennis WE, Jackson DA, Stallings JD, Lewis JA (2014) Detection of dichlorvos adducts in a hepatocyte cell line. J Proteome Res 13:3583-3595
Burniston JG, Connolly J, Kainulainen H, Britton SL, Koch LG (2014) Label-free profiling of skeletal muscle using high-definition mass spectrometry. Proteomics 14:2339-2344
Buts K, Michielssens S, Hertog ML, Hayakawa E, Cordewener J, America AH, Nicolai BM, Carpentier SC (2014) Improving the identification rate of data independent label-free quantitative proteomics experiments on non-model crops: A case study on apple fruit. J Proteomics 105:31-45
Chawade A, Sandin M, Teleman J, Malmstrom J, Levander F (2014) Data processing has major impact on the outcome of quantitative label-free LC-MS analysis. J Proteome Res
Choi M, Chang CY, Clough T, Broudy D, Killeen T, MacLean B, Vitek O (2014) MSstats: an R package for statistical analysis of quantitative mass spectrometry-based proteomic experiments. Bioinformatics 30:2524-2526
Christensen JB, Dionisio G, Poulsen HD, Brinch-Pedersen H (2014) Effect of pH and recombinant barley (Hordeum vulgare L.) endoprotease B2 on degradation of proteins in soaked barley. J Agric Food Chem 62:8562-8570
Claussnitzer M, Dankel SN, Klocke B, Grallert H, Glunk V, Berulava T, Lee H, et al. (2014) Leveraging cross-species transcription factor binding site patterns: from diabetes risk loci to disease mechanisms. Cell 156:343-358
Colak E, Leslie A, Zausmer K, Khatamzas E, Kubarenko AV, Pichulik T, Klimosch SN, Mayer A, Siggs O, Hector A, Fischer R, Klesser B, Rautanen A, Frank M, Hill AV, Manoury B, Beutler B, Hartl D, Simmons A, Weber AN (2014) RNA and Imidazoquinolines Are Sensed by Distinct TLR7/8 Ectodomain Sites Resulting in Functionally Disparate Signaling Events. J Immunol 192:5963-5973
Coumans JV, Gau D, Poljak A, Wasinger V, Roy P, Moens P (2014) Green fluorescent protein expression triggers proteome changes in breast cancer cells. Exp Cell Res 320:33-45
Cruz DECR, Bernardes DASA, Soares R, Almeida AM, Coelho AV, Marques DASJ, Branquinho C (2014) Differential proteomics of dehydration and rehydration in bryophytes: evidence towards a common desiccation tolerance mechanism. Plant Cell Environ 37:1499-1515
Dagley LF, Croft NP, Isserlin R, Olsen JB, Fong V, Emili A, Purcell AW (2014) Discovery of novel disease-specific and membrane-associated candidate markers in a mouse model of multiple sclerosis. Mol Cell Proteomics 13:679-700
Daou P, Hasan S, Breitsprecher D, Baudelet E, Camoin L, Audebert S, Goode BL, Badache A (2014) Essential and nonredundant roles for Diaphanous formins in cortical microtubule capture and directed cell migration. Mol Biol Cell 25:658-668
Darby AC, Christina Gill A, Armstrong SD, Hartley CS, Xia D, Wastling JM, Makepeace BL (2014) Integrated transcriptomic and proteomic analysis of the global response of Wolbachia to doxycycline-induced stress. Isme J 8:925-937
de Costa D, Broodman I, Calame W, Stingl C, Dekker LJ, Vernhout RM, de Koning HJ, Hoogsteden HC, Sillevis Smitt PA, van Klaveren RJ, Luider TM, Vanduijn MM (2014) Peptides from the variable region of specific antibodies are shared among lung cancer patients. PLoS One 9:e96029
Deighton RF, Le Bihan T, Martin SF, Gerth AM, McCulloch M, Edgar JM, Kerr LE, Whittle IR, McCulloch J (2014) Interactions among mitochondrial proteins altered in glioblastoma. J Neurooncol 118:247-256
Deschoenmaeker F, Facchini R, Leroy B, Badri H, Zhang CC, Wattiez R (2014) Proteomic and cellular views of Arthrospira sp. PCC 8005 adaptation to nitrogen depletion. Microbiology 160:1224-1236
DeVeale B, Bausch-Fluck D, Seaberg R, Runciman S, Akbarian V, Karpowicz P, Yoon C, Song H, Leeder R, Zandstra PW, Wollscheid B, van der Kooy D (2014) Surfaceome profiling reveals regulators of neural stem cell function. Stem Cells 32:258-268
Diepenbroek M, Casadei N, Esmer H, Saido TC, Takano J, Kahle PJ, Nixon RA, Rao MV, Melki R, Pieri L, Helling S, Marcus K, Krueger R, Masliah E, Riess O, Nuber S (2014) Overexpression of the calpain-specific inhibitor calpastatin reduces human alpha-Synuclein processing, aggregation and synaptic impairment in [A30P]alphaSyn transgenic mice. Hum Mol Genet 23:3975-3989
Feng Y, De Franceschi G, Kahraman A, Soste M, Melnik A, Boersema PJ, de Laureto PP, Nikolaev Y, Oliveira AP, Picotti P (2014) Global analysis of protein structural changes in complex proteomes. Nat Biotechnol 32:1036-1044
Fengos G, Schmidt A, Martin K, Fluri E, Aebersold R, Iber D, Pertz O (2014) Spatial proteomic and phospho-proteomic organization in three prototypical cell migration modes. Proteome Sci 12:23
Fessart D, Martin-Negrier ML, Claverol S, Thiolat ML, Crevel H, Toussaint C, Bonneu M, Muller B, Savineau JP, Delom F (2014) Proteomic remodeling of proteasome in right heart failure. J Mol Cell Cardiol 66:41-52
Finamore F, Priego-Capote F, Nolli S, Zufferey A, Fontana P, Sanchez J (2014) Characterisation of the influences of aspirin-acetylation and glycation on human plasma proteins. J Proteomics 114C:125-135
Finne K, Vethe H, Skogstrand T, Leh S, Dahl TD, Tenstad O, Berven FS, Reed RK, Vikse BE (2014) Proteomic analysis of formalin-fixed paraffin-embedded glomeruli suggests depletion of glomerular filtration barrier proteins in two-kidney, one-clip hypertensive rats. Nephrol Dial Transplant 29:2217-2227
Flenkenthaler F, Windschuttl S, Frohlich T, Schwarzer JU, Mayerhofer A, Arnold GJ (2014) Secretome analysis of testicular peritubular cells: a window into the human testicular microenvironment and the spermatogonial stem cell niche in man. J Proteome Res 13:1259-1269
Folkard DL, Melchini A, Traka MH, Al-Bakheit A, Saha S, Mulholland F, Watson A, Mithen RF (2014) Suppression of LPS-induced transcription and cytokine secretion by the dietary isothiocyanate sulforaphane. Mol Nutr Food Res 58:2286-2296
Fujioka Y, Suzuki SW, Yamamoto H, Kondo-Kakuta C, Kimura Y, Hirano H, Akada R, Inagaki F, Ohsumi Y, Noda NN (2014) Structural basis of starvation-induced assembly of the autophagy initiation complex. Nat Struct Mol Biol 21:513-521
Fuzesi-Levi MG, Ben-Nissan G, Bianchi E, Zhou H, Deery MJ, Lilley KS, Levin Y, Sharon M (2014) Dynamic regulation of the COP9 signalosome in response to DNA damage. Mol Cell Biol 34:1066-1076
Gerster S, Kwon T, Ludwig C, Matondo M, Vogel C, Marcotte EM, Aebersold R, Buhlmann P (2014) Statistical approach to protein quantification. Mol Cell Proteomics 13:666-677
Gobert M, Sayd T, Gatellier P, Sante-Lhoutellier V (2014) Application to proteomics to understand and modify meat quality. Meat Sci 98:539-543
Griciuc A, Roux MJ, Merl J, Giangrande A, Hauck SM, Aron L, Ueffing M (2014) Proteomic survey reveals altered energetic patterns and metabolic failure prior to retinal degeneration. J Neurosci 34:2797-2812
Gu X, Hu Z, Ebrahem Q, Crabb JS, Mahfouz RZ, Radivoyevitch T, Crabb JW, Saunthararajah Y (2014) Runx1 regulation of Pu.1 corepressor/coactivator exchange identifies specific molecular targets for leukemia differentiation therapy. J Biol Chem 289:14881-14895
Guo M, Zhao B, Liu H, Zhang L, Peng L, Qin L, Zhang Z, Li J, Cai C, Gao X (2014) A Metabolomic Strategy to Screen the Prototype Components and Metabolites of Shuang-Huang-Lian Injection in Human Serum by Ultra Performance Liquid Chromatography Coupled with Quadrupole Time-of-Flight Mass Spectrometry. J Anal Methods Chem 2014:241505
Gutierrez-Carbonell E, Takahashi D, Lattanzio G, Rodriguez-Celma J, Kehr J, Soll J, Philippar K, Uemura M, Abadia J, Lopez-Millan AF (2014) The Distinct Functional Roles of the Inner and Outer Chloroplast Envelope of Pea (Pisum sativum) As Revealed by Proteomic Approaches. J Proteome Res 13:2941-2953
Hacariz O, Baykal AT, Akgun M, Kavak P, Sagiroglu MS, Sayers GP (2014) Generating a detailed protein profile of Fasciola hepatica during the chronic stage of infection in cattle. Proteomics 14:1519-1530
Haga SW, Wu HF (2014) Overview of software options for processing, analysis and interpretation of mass spectrometric proteomic data. J Mass Spectrom 49:959-969
Hatipoglu I, Ercan D, Acilan C, Basalp A, Durali D, Baykal AT (2014) Hepatitis B virus e antigen (HBeAg) may have a negative effect on dendritic cell generation. Immunobiology 219:944-949
Heim A, Grimm C, Muller U, Haussler S, Mackeen MM, Merl J, Hauck SM, Kessler BM, Schofield CJ, Wolf A, Bottger A (2014) Jumonji domain containing protein 6 (Jmjd6) modulates splicing and specifically interacts with arginine-serine-rich (RS) domains of SR- and SR-like proteins. Nucleic Acids Res 42:7833-7850
Helm D, Vissers JP, Hughes CJ, Hahne H, Ruprecht B, Pachl F, Grzyb A, Richardson K, Wildgoose J, Maier SK, Marx H, Wilhelm M, Becher I, Lemeer S, Bantscheff M, Langridge JI, Kuster B (2014) Ion Mobility Tandem Mass Spectrometry Enhances Performance of Bottom-up Proteomics. Mol Cell Proteomics 13:3709-3715
Heming JD, Huffman JB, Jones LM, Homa FL (2014) Isolation and characterization of the herpes simplex virus 1 terminase complex. J Virol 88:225-236
Hofener M, Pachl F, Take T, Fischer von Mollard G, Kuster B, Sewald N (2014) Investigating RET RTK signaling pathways using an IAP-based activity-profiling approach. J Proteome Res 13:3628-3634
Huang Y, Mao Y, Buczek-Thomas JA, Nugent MA, Zaia J (2014) Oligosaccharide substrate preferences of human extracellular sulfatase Sulf2 using liquid chromatography-mass spectrometry based glycomics approaches. PLoS One 9:e105143
Jackson AP, Otto TD, Darby A, Ramaprasad A, Xia D, Echaide IE, Farber M, et al. (2014) The evolutionary dynamics of variant antigen genes in Babesia reveal a history of genomic innovation underlying host-parasite interaction. Nucleic Acids Res 42:7113-7131
Kim PY, Tan O, Diakiw SM, Carter D, Sekerye EO, Wasinger VC, Liu T, Kavallaris M, Norris MD, Haber M, Chesler L, Dolnikov A, Trahair TN, Cheung NK, Marshall GM, Cheung BB (2014) Identification of plasma complement C3 as a potential biomarker for neuroblastoma using a quantitative proteomic approach. J Proteomics 96:1-12
Kochin V, Shimi T, Torvaldson E, Adam SA, Goldman A, Pack CG, Melo-Cardenas J, Imanishi SY, Goldman RD, Eriksson JE (2014) Interphase phosphorylation of lamin A. J Cell Sci 127:2683-2696
Kohl M, Megger DA, Trippler M, Meckel H, Ahrens M, Bracht T, Weber F, Hoffmann AC, Baba HA, Sitek B, Schlaak JF, Meyer HE, Stephan C, Eisenacher M (2014) A practical data processing workflow for multi-OMICS projects. Biochim Biophys Acta 1844:52-62
Kretschmer A, Moller G, Lee H, Laumen H, von Toerne C, Schramm K, Prokisch H, Eyerich S, Wahl S, Baurecht H, Franke A, Claussnitzer M, Eyerich K, Teumer A, Milani L, Klopp N, Hauck SM, Illig T, Peters A, Waldenberger M, Adamski J, Reischl E, Weidinger S (2014) A common atopy-associated variant in the Th2 cytokine locus control region impacts transcriptional regulation and alters SMAD3 and SP1 binding. Allergy 69:632-642
Ku X, Heinzlmeir S, Liu X, Medard G, Kuster B (2014) A new chemical probe for quantitative proteomic profiling of fibroblast growth factor receptor and its inhibitors. J Proteomics 96:44-55
Kuhn A, Nieters A, Kottgen A, Goek ON, Michels K, Nothlings U, Jacobs G, Meisinger C, Pessler F, Akmatov MF, Kuhnisch J, Moebus S, Glocker E, Naus S, Keimling M, Leitzmann M, Linseisen J, Sarioglu H, von Toerne C, Hauck SM, Wallaschofski H, Wichmann HE, Illig T (2014) Feasibility and quality development of biomaterials in the pretest studies of the German National Cohort. Bundesgesundheitsblatt Gesundheitsforschung Gesundheitsschutz 57:1255-1263
Laiakis EC, Strassburg K, Bogumil R, Lai S, Vreeken RJ, Hankemeier T, Langridge J, Plumb RS, Fornace AJ, Jr., Astarita G (2014) Metabolic phenotyping reveals a lipid mediator response to ionizing radiation. J Proteome Res 13:4143-4154
Lemeer S, Gholami AM, Wu Z, Kuster B (2014) Quantitative proteome profiling of human myoma and myometrium tissue reveals kinase expression signatures with potential for therapeutic intervention. Proteomics
Lennon R, Byron A, Humphries JD, Randles MJ, Carisey A, Murphy S, Knight D, Brenchley PE, Zent R, Humphries MJ (2014) Global analysis reveals the complexity of the human glomerular extracellular matrix. J Am Soc Nephrol 25:939-951
Li M, Zhao M, Gao Y (2014) Changes of proteins induced by anticoagulants can be more sensitively detected in urine than in plasma. Sci China Life Sci (In press)
Li X, Zhao M, Li M, Jia L, Gao Y (2014) Effects of Three Commonly-used Diuretics on the Urinary Proteome. Genomics Proteomics Bioinformatics (In press)
Linge A, Maurya P, Friedrich K, Baretton GB, Kelly S, Henry M, Clynes M, Larkin A, Meleady P (2014) Identification and Functional Validation of RAD23B as a Potential Protein in Human Breast Cancer Progression. J Proteome Res (In press)
Lukasse PN, America AH (2014) Protein Inference Using Peptide Quantification Patterns. J Proteome Res (In press)
Luo R, Fang L, Jin H, Wang D, An K, Xu N, Chen H, Xiao S (2014) Label-free quantitative phosphoproteomic analysis reveals differentially regulated proteins and pathway in PRRSV-infected pulmonary alveolar macrophages. J Proteome Res 13:1270-1280
Luxton HJ, Barnouin K, Kelly G, Hanrahan S, Totty N, Neal DE, Whitaker HC (2014) Regulation of the localisation and function of the oncogene LYRIC/AEG-1 by ubiquitination at K486 and K491. Mol Oncol 8:633-641
Ly A, Scheerer MF, Zukunft S, Muschet C, Merl J, Adamski J, de Angelis MH, Neschen S, Hauck SM, Ueffing M (2014) Retinal proteome alterations in a mouse model of type 2 diabetes. Diabetologia 57:192-203
MacDonald G, Nalvarte I, Smirnova T, Vecchi M, Aceto N, Dolemeyer A, Frei A, Lienhard S, Wyckoff J, Hess D, Seebacher J, Keusch JJ, Gut H, Salaun D, Mazzarol G, Disalvatore D, Bentires-Alj M, Di Fiore PP, Badache A, Hynes NE (2014) Memo is a copper-dependent redox protein with an essential role in migration and metastasis. Sci Signal 7:ra56
Majek P, Pecankova K, Dyr JE (2014) The effect of the biological variability of samples on Coomassie blue dye based fast staining for SDS-PAGE in nonfixed gels. Electrophoresis 35:3008-3011
Megger DA, Pott LL, Ahrens M, Padden J, Bracht T, Kuhlmann K, Eisenacher M, Meyer HE, Sitek B (2014) Comparison of label-free and label-based strategies for proteome analysis of hepatoma cell lines. Biochim Biophys Acta 1844:967-976
Miller MH, Walsh SV, Atrih A, Huang JT, Ferguson MA, Dillon JF (2014) The serum proteome of nonalcoholic fatty liver disease: a multimodal approach to discovery of biomarkers of nonalcoholic steatohepatitis. J Gastroenterol Hepatol 29:1839-1847
Molin S, Merl J, Dietrich KA, Regauer M, Flaig M, Letule V, Saucke T, Herzinger T, Ruzicka T, Hauck SM (2014) The hand eczema proteome: imbalance of epidermal barrier proteins. Br J Dermatol
Okayama A, Miyagi Y, Oshita F, Nishi M, Nakamura Y, Nagashima Y, Akimoto K, Ryo A, Hirano H (2014) Proteomic analysis of proteins related to prognosis of lung adenocarcinoma. J Proteome Res 13:4686-4694
Otto A, Becher D, Schmidt F (2014) Quantitative proteomics in the field of microbiology. Proteomics 14:547-565
Paardekooper Overman J, Yi JS, Bonetti M, Soulsby M, Preisinger C, Stokes MP, Hui L, Silva JC, Overvoorde J, Giansanti P, Heck AJ, Kontaridis MI, den Hertog J, Bennett AM (2014) PZR coordinates Shp2 Noonan and LEOPARD syndrome signaling in zebrafish and mice. Mol Cell Biol 34:2874-2889
Paglia G, Williams JP, Menikarachchi L, Thompson JW, Tyldesley-Worster R, Halldorsson S, Rolfsson O, Moseley A, Grant D, Langridge J, Palsson BO, Astarita G (2014) Ion mobility derived collision cross sections to support metabolomics applications. Anal Chem 86:3985-3993
Paglia G, Angel PM, Williams JP, Richardson K, Olivos HJ, Thompson JW, Menikarachchi LC, Lai S, Walsh C, Moseley MA, Plumb RS, Grant DF, Palsson BO, Langridge JI, Geromanos SJ, Astarita G (2014) Ion-Mobility-Derived Collision Cross Section as an Additional Measure for Lipid Fingerprinting and Identification. Anal Chem
Peffers MJ, Thorpe CT, Collins JA, Eong R, Wei TK, Screen HR, Clegg PD (2014) Proteomic analysis reveals age-related changes in tendon matrix composition, with age- and injury-specific matrix fragmentation. J Biol Chem 289:25867-25878
Pischedda F, Szczurkowska J, Cirnaru MD, Giesert F, Vezzoli E, Ueffing M, Sala C, Francolini M, Hauck SM, Cancedda L, Piccoli G (2014) A cell surface biotinylation assay to reveal membrane-associated neuronal cues: Negr1 regulates dendritic arborization. Mol Cell Proteomics 13:733-748
Plaza-Menacho I, Barnouin K, Goodman K, Martinez-Torres RJ, Borg A, Murray-Rust J, Mouilleron S, Knowles P, McDonald NQ (2014) Oncogenic RET kinase domain mutations perturb the autophosphorylation trajectory by enhancing substrate presentation in trans. Mol Cell 53:738-751
Politis A, Stengel F, Hall Z, Hernandez H, Leitner A, Walzthoeni T, Robinson CV, Aebersold R (2014) A mass spectrometry-based hybrid method for structural modeling of protein complexes. Nat Methods 11:403-406
Pournaras S, Poulou A, Dafopoulou K, Chabane YN, Kristo I, Makris D, Hardouin J, Cosette P, Tsakris A, De E (2014) Growth retardation, reduced invasiveness, and impaired colistin-mediated cell death associated with colistin resistance development in Acinetobacter baumannii. Antimicrob Agents Chemother 58:828-832
Qi D, Krishna R, Jones AR (2014) The jmzQuantML programming interface and validator for the mzQuantML data standard. Proteomics 14:685-688
Qu J, Young R, Page BJ, Shen X, Tata N, Li J, Duan X, Fallavollita JA, Canty JM, Jr. (2014) Reproducible Ion-Current-Based Approach for 24-Plex Comparison of the Tissue Proteomes of Hibernating versus Normal Myocardium in Swine Models. J Proteome Res (In press)
Ramasamy P, Murphy CC, Clynes M, Horgan N, Moriarty P, Tiernan D, Beatty S, Kennedy S, Meleady P (2014) Proteomics in uveal melanoma. Exp Eye Res 118:1-12
Ritamo I, Cloutier M, Valmu L, Neron S, Rabina J (2014) Comparison of the glycosylation of in vitro generated polyclonal human IgG and therapeutic immunoglobulins. Mol Immunol 57:255-262
Rocha B, Calamia V, Casas V, Carrascal M, Blanco FJ, Ruiz-Romero C (2014) Secretome analysis of human mesenchymal stem cells undergoing chondrogenic differentiation. J Proteome Res 13:1045-1054
Rogowska-Wrzesinska A, Wrzesinski K, Fey SJ (2014) Heteromer score-using internal standards to assess the quality of proteomic data. Proteomics 14:1042-1047
Romao JM, He ML, McAllister TA, Guan LL (2014) Effect of age on bovine subcutaneous fat proteome: molecular mechanisms of physiological variations during beef cattle growth. J Anim Sci 92:3316-3327
Rouillon J, Zocevic A, Leger T, Garcia C, Camadro JM, Udd B, Wong B, Servais L, Voit T, Svinartchouk F (2014) Proteomics profiling of urine reveals specific titin fragments as biomarkers of Duchenne muscular dystrophy. Neuromuscul Disord 24:563-573
Searcy JL, Le Bihan T, Salvadores N, McCulloch J, Horsburgh K (2014) Impact of age on the cerebrovascular proteomes of wild-type and Tg-SwDI mice. PLoS One 9:e89970
Sedic M, Gethings LA, Vissers JP, Shockcor JP, McDonald S, Vasieva O, Lemac M, Langridge JI, Batinic D, Pavelic SK (2014) Label-free mass spectrometric profiling of urinary proteins and metabolites from paediatric idiopathic nephrotic syndrome. Biochem Biophys Res Commun 452:21-26
Shevchenko A, Yang Y, Knaust A, Thomas H, Jiang H, Lu E, Wang C (2014) Proteomics identifies the composition and manufacturing recipe of the 2500-year old sourdough bread from Subeixi cemetery in China. J Proteomics 105:363-371
Skeffington AW, Graf A, Duxbury Z, Gruissem W, Smith AM (2014) Glucan, Water Dikinase Exerts Little Control over Starch Degradation in Arabidopsis Leaves at Night. Plant Physiol 165:866-879
Spieler D, Kaffe M, Knauf F, Bessa J, Tena JJ, Giesert F, Schormair B, et al. (2014) Restless legs syndrome-associated intronic common variant in Meis1 alters enhancer function in the developing telencephalon. Genome Res 24:592-603
Su MA, Huang YT, Chen IT, Lee DY, Hsieh YC, Li CY, Ng TH, Liang SY, Lin SY, Huang SW, Chiang YA, Yu HT, Khoo KH, Chang GD, Lo CF, Wang HC (2014) An Invertebrate Warburg Effect: A Shrimp Virus Achieves Successful Replication by Altering the Host Metabolome via the PI3K-Akt-mTOR Pathway. PLoS Pathog 10:e1004196
Suila H, Hirvonen T, Kotovuori A, Ritamo I, Kerkela E, Anderson H, Natunen S, Tuimala J, Laitinen S, Nystedt J, Rabina J, Valmu L (2014) Human umbilical cord blood-derived mesenchymal stromal cells display a novel interaction between P-selectin and galectin-1. Scand J Immunol 80:12-21
Sultana N, Florance HV, Johns A, Smirnoff N (2014) Ascorbate deficiency influences the leaf cell wall glycoproteome in Arabidopsis thaliana. Plant Cell Environ (In press)
Sundin L, Vanholme R, Geerinck J, Goeminne G, Hofer R, Kim H, Ralph J, Boerjan W (2014) Mutation of the Inducible ARABIDOPSIS THALIANA CYTOCHROME P450 REDUCTASE2 Alters Lignin Composition and Improves Saccharification. Plant Physiol 166:1956-1971
Talamantes T, Ughy B, Domonkos I, Kis M, Gombos Z, Prokai L (2014) Label-free LC-MS/MS identification of phosphatidylglycerol-regulated proteins in Synechocystis sp. PCC6803. Proteomics 14:1053-1057
Tang Z, Baykal AT, Gao H, Quezada HC, Zhang H, Bereczki E, Serhatli M, Baykal B, Acioglu C, Wang S, Ioja E, Ji X, Zhang Y, Guan Z, Winblad B, Pei JJ (2014) mTor Is a Signaling Hub in Cell Survival: A Mass-Spectrometry-Based Proteomics Investigation. J Proteome Res
Theron L, Gueugneau M, Coudy C, Viala D, Bijlsma A, Butler-Browne G, Maier A, Bechet D, Chambon C (2014) Label-free quantitative protein profiling of vastus lateralis muscle during human aging. Mol Cell Proteomics 13:283-294
Tu C, Sheng Q, Li J, Shen X, Zhang M, Shyr Y, Qu J (2014) ICan: An Optimized Ion-Current-Based Quantification Procedure with Enhanced Quantitative Accuracy and Sensitivity in Biomarker Discovery. J Proteome Res 13:5888-5897
Uhl PB, Szober CM, Amann B, Alge-Priglinger C, Ueffing M, Hauck SM, Deeg CA (2014) In situ cell surface proteomics reveals differentially expressed membrane proteins in retinal pigment epithelial cells during autoimmune uveitis. J Proteomics 109:50-62
Van Acker R, Leple JC, Aerts D, Storme V, Goeminne G, Ivens B, Legee F, Lapierre C, Piens K, Van Montagu MC, Santoro N, Foster CE, Ralph J, Soetaert W, Pilate G, Boerjan W (2014) Improved saccharification and ethanol yield from field-grown transgenic poplar deficient in cinnamoyl-CoA reductase. Proc Natl Acad Sci U S A 111:845-850
Vandermoten S, Harmel N, Mazzucchelli G, De Pauw E, Haubruge E, Francis F (2014) Comparative analyses of salivary proteins from three aphid species. Insect Mol Biol 23:67-77
Vanderschuren H, Nyaboga E, Poon JS, Baerenfaller K, Grossmann J, Hirsch-Hoffmann M, Kirchgessner N, Nanni P, Gruissem W (2014) Large-Scale Proteomics of the Cassava Storage Root and Identification of a Target Gene to Reduce Postharvest Deterioration. Plant Cell 26:1913-1924
Vehmas AP, Muth-Pawlak D, Huhtinen K, Saloniemi-Heinonen T, Jaakkola K, Laajala TD, Kaprio H, Suvitie PA, Aittokallio T, Siitari H, Perheentupa A, Poutanen M, Corthals GL (2014) Ovarian Endometriosis Signatures Established through Discovery and Directed Mass Spectrometry Analysis. J Proteome Res 13:4983-4994
Velikova V, Ghirardo A, Vanzo E, Merl J, Hauck SM, Schnitzler JP (2014) Genetic manipulation of isoprene emissions in poplar plants remodels the chloroplast proteome. J Proteome Res 13:2005-2018
Vranakis I, Goniotakis I, Psaroulaki A, Sandalakis V, Tselentis Y, Gevaert K, Tsiotis G (2014) Proteome studies of bacterial antibiotic resistance mechanisms. J Proteomics 97:88-99
Wiens O, Xia D, von Schubert C, Wastling JM, Dobbelaere DA, Heussler VT, Woods KL (2014) Cell cycle-dependent phosphorylation of Theileria annulata schizont surface proteins. PLoS One 9:e103821
Wilhelm T, Jones AM (2014) Identification of related peptides through the analysis of fragment ion mass shifts. J Proteome Res 13:4002-4011
Wojdyla K, Williamson J, Roepstorff P, Rogowska-Wrzesinska A (2014) The SNO/SOH TMT strategy for combinatorial analysis of reversible cysteine oxidations. J Proteomics 113:415-434
Xu J, Chatterjee M, Baguley TD, Brouillette J, Kurup P, Ghosh D, Kanyo J, Zhang Y, Seyb K, Ononenyi C, Foscue E, Anderson GM, Gresack J, Cuny GD, Glicksman MA, Greengard P, Lam TT, Tautz L, Nairn AC, Ellman JA, Lombroso PJ (2014) Inhibitor of the tyrosine phosphatase STEP reverses cognitive deficits in a mouse model of alzheimer's disease. PLoS Biol 12:e1001923
Yang WS, SriRamaratnam R, Welsch ME, Shimada K, Skouta R, Viswanathan VS, Cheah JH, Clemons PA, Shamji AF, Clish CB, Brown LM, Girotti AW, Cornish VW, Schreiber SL, Stockwell BR (2014) Regulation of ferroptotic cancer cell death by GPX4. Cell 156:317-331
Yerlikaya A, Okur E, Baykal AT, Acilan C, Boyaci I, Ulukaya E (2014) A proteomic analysis of p53-independent induction of apoptosis by bortezomib in 4T1 breast cancer cell line. J Proteomics 113:315-325
Yin X, Liu X, Zhang Y, Yan G, Wang F, Lu H, Shen H, Yang P (2014) Rapid and sensitive profiling and quantification of the human cell line proteome by LC-MS/MS without prefractionation. Proteomics 14:2008-2016
Zhang A, Zhou X, Zhao H, Guan Y, Zhou S, Yan GL, Ma Z, Liu Q, Wang X (2014) Rapidly improved determination of metabolites from biological data sets using the high-efficient TransOmics tool. Mol Biosyst 10:2160-2165
Zhao X, Sidoli S, Wang L, Wang W, Guo L, Jensen ON, Zheng L (2014) Comparative proteomic analysis of histone post-translational modifications upon ischemia/reperfusion-induced retinal injury. J Proteome Res 13:2175-21868
Zhao Z, Wu F, Ding S, Sun L, Liu Z, Ding K, Lu J (2014) Label-free quantitative proteomic analysis reveals potential biomarkers and pathways in renal cell carcinoma. Tumour Biol
2013
Ahmed S, Maratha A, Butt AQ, Shevlin E, Miggin SM. (2013) TRIF-mediated TLR3 and TLR4 signaling is negatively regulated by ADAM15. J Immunol 190:2217-2228
Ahrne E, Molzahn L, Glatter T, Schmidt A. (2013) Critical assessment of proteome-wide label-free absolute abundance estimation strategies. Proteomics 13:2567-2578
Alajati A, Sausgruber N, Aceto N, Duss S, Sarret S, Voshol H, Bonenfant D, Bentires-Alj M. (2013) Mammary tumor formation and metastasis evoked by a HER2 splice variant. Cancer Res 73:5320-5327
Babaei F, Ramalingam R, Tavendale A, Liang Y, Yan LS, Ajuh P, Cheng SH, Lam YW (2013) Novel blood collection method allows plasma proteome analysis from single zebrafish. J Proteome Res 12:1580-1590
Bager R, Johansen JS, Jensen JK, Stensballe A, Jendroszek A, Buxbom L, Sorensen HP, Andreasen PA. (2013) Protein conformational change delayed by steric hindrance from an N-linked glycan. J Mol Biol 425:2867-2877
Berle M, Kroksveen AC, Garberg H, Aarhus M, Haaland OA, Wester K, Ulvik RJ, Helland C, Berven F. (2013) Quantitative proteomics comparison of arachnoid cyst fluid and cerebrospinal fluid collected perioperatively from arachnoid cyst patients. Fluids Barriers CNS 10:17 (In press)
Bond NJ, Shliaha PV, Lilley KS, Gatto L. (2013) Improving qualitative and quantitative performance for MS(E)-based label-free proteomics. J Proteome Res 12:2340-2353
Bostanci N, Ramberg P, Wahlander A, Grossman J, Jonsson D, Barnes VM, Papapanou PN. (2013) Label-free quantitative proteomics reveals differentially regulated proteins in experimental gingivitis. J Proteome Res 12:657-678
Brownridge P, Lawless C, Payapilly AB, Lanthaler K, Holman SW, Harman VM, Grant CM, Beynon RJ, Hubbard SJ. (2013) Quantitative analysis of chaperone network throughput in budding yeast. Proteomics 13:1276-1291
Bruce C, Stone K, Gulcicek E, Williams K. (2013) Proteomics and the analysis of proteomic data: 2013 overview of current protein-profiling technologies. Curr Protoc Bioinformatics Chapter 13:Unit 13 21
Cerny AC, Oberacker T, Pfannstiel J, Weigold S, Will C, Huber A. (2013) Mutation of light-dependent phosphorylation sites of the Drosophila transient receptor potential-like (TRPL) ion channel affects its subcellular localization and stability. J Biol Chem 288:15600-15613
Collier TS, Diraviyam K, Monsey J, Shen W, Sept D, Bose R. (2013) Carboxyl group footprinting mass spectrometry and molecular dynamics identify key interactions in the HER2-HER3 receptor tyrosine kinase interface. J Biol Chem 288:25254-25264
Cruciani F, Wasinger V, Turroni S, Calanni F, Donders G, Brigidi P, Vitali B. (2013) Proteome profiles of vaginal fluids from women affected by bacterial vaginosis and healthy controls: outcomes of rifaximin treatment. J Antimicrob Chemother 68:2648-2659
Dagley LF, Emili A, Purcell AW. (2013) Application of quantitative proteomics technologies to the biomarker discovery pipeline for multiple sclerosis. Proteomics Clin Appl 7:91-108
Degroote RL, Hauck SM, Treutlein G, Amann B, Frohlich KJ, Kremmer E, Merl J, Stangassinger M, Ueffing M, Deeg CA. (2013) Expression changes and novel interaction partners of talin 1 in effector cells of autoimmune uveitis. J Proteome Res 12:5812-5819
Euler KN, Hauck SM, Ueffing M, Deeg CA. (2013) Bovine neonatal pancytopenia--comparative proteomic characterization of two BVD vaccines and the producer cell surface proteome (MDBK). BMC Vet Res 9:18 (in press)
Fischer R, Bowness P, Kessler BM. (2013) Two birds with one stone: Doing metabolomics with your proteomics kit. Proteomics 13:3371-3386
Franko A, Baris OR, Bergschneider E, von Toerne C, Hauck SM, Aichler M, Walch AK, Wurst W, Wiesner RJ, Johnston IC, de Angelis MH (2013) Efficient isolation of pure and functional mitochondria from mouse tissues using automated tissue disruption and enrichment with anti-TOM22 magnetic beads. PLoS One 8:e82392
Frei AP, Moest H, Novy K, Wollscheid B. (2013) Ligand-based receptor identification on living cells and tissues using TRICEPS. Nat Protoc 8:1321-1336
Freret M, Drouot L, Obry A, Ahmed-Lacheheb S, Dauly C, Adriouch S, Cosette P, Authier FJ, Boyer O. (2013) Overexpression of MHC class I in muscle of lymphocyte-deficient mice causes a severe myopathy with induction of the unfolded protein response. Am J Pathol 183:893-904
Frese CK, Boender AJ, Mohammed S, Heck AJ, Adan RA, Altelaar AF. (2013) Profiling of diet-induced neuropeptide changes in rat brain by quantitative mass spectrometry. Anal Chem 85:4594-4604
Gluck F, Hoogland C, Antinori P, Robin X, Nikitin F, Zufferey A, Pasquarello C, Fetaud V, Dayon L, Muller M, Lisacek F, Geiser L, Hochstrasser D, Sanchez JC, Scherl A. (2013) EasyProt--an easy-to-use graphical platform for proteomics data analysis. J Proteomics 79:146-160
Hahne H, Sobotzki N, Nyberg T, Helm D, Borodkin VS, van Aalten DM, Agnew B, Kuster B. (2013) Proteome wide purification and identification of O-GlcNAc-modified proteins using click chemistry and mass spectrometry. J Proteome Res 12:927-936
Hakenjos JP, Bejai S, Ranftl Q, Behringer C, Vlot AC, Absmanner B, Hammes U, Heinzlmeir S, Kuster B, Schwechheimer C. (2013) ML3 is a NEDD8- and ubiquitin-modified protein. Plant Physiol 163:135-149
Han B, Zhang L, Feng M, Fang Y, Li J (2013) An integrated proteomics reveals pathological mechanism of honeybee (Apis cerena) sacbrood disease. J Proteome Res 12:1881-1897
Haslene-Hox H, Oveland E, Woie K, Salvesen HB, Wiig H, Tenstad O. (2013) Increased WD-repeat containing protein 1 in interstitial fluid from ovarian carcinomas shown by comparative proteomic analysis of malignant and healthy gynecological tissue. Biochim Biophys Acta 1834:2347-2359
Herrmann AG, Deighton RF, Le Bihan T, McCulloch MC, Searcy JL, Kerr LE, McCulloch J. (2013) Adaptive changes in the neuronal proteome: mitochondrial energy production, endoplasmic reticulum stress, and ribosomal dysfunction in the cellular response to metabolic stress. J Cereb Blood Flow Metab 33:673-683
Hirayama M, Kobayashi D, Mizuguchi S, Morikawa T, Nagayama M, Midorikawa U, Wilson MM, Nambu AN, Yoshizawa AC, Kawano S, Araki N. (2013) Integrated proteomics identified novel activation of dynein IC2-GR-COX-1 signaling in neurofibromatosis type I (NF1) disease model cells. Mol Cell Proteomics 12:1377-1394
Holland A, Dowling P, Zweyer M, Swandulla D, Henry M, Clynes M, Ohlendieck K. (2013) Proteomic profiling of cardiomyopathic tissue from the aged mdx model of Duchenne muscular dystrophy reveals a drastic decrease in laminin, nidogen and annexin. Proteomics 13:2312-2323
Ijsselstijn L, Stoop MP, Stingl C, Sillevis Smitt PA, Luider TM, Dekker LJ. (2013) Comparative Study of Targeted and Label-free Mass Spectrometry Methods for Protein Quantification. J Proteome Res (in press)
Kehasse A, Rich CB, Lee A, McComb ME, Costello CE, Trinkaus-Randall V. (2013) Epithelial wounds induce differential phosphorylation changes in response to purinergic and EGF receptor activation. Am J Pathol 183:1841-1852
Kerkela E, Hakkarainen T, Makela T, Raki M, Kambur O, Kilpinen L, Nikkila J, Lehtonen S, Ritamo I, Pernu R, Pietila M, Takalo R, Juvonen T, Bergstrom K, Kalso E, Valmu L, Laitinen S, Lehenkari P, Nystedt J. (2013) Transient proteolytic modification of mesenchymal stromal cells increases lung clearance rate and targeting to injured tissue. Stem Cells Transl Med 2:510-520
Kim PY, Tan O, Diakiw SM, Carter D, Sekerye EO, Wasinger VC, Liu T, Kavallaris M, Norris MD, Haber M, Chesler L, Dolnikov A, Trahair TN, Cheung NK, Marshall GM, Cheung BB. (2013) Identification of plasma Complement C3 as a potential biomarker for neuroblastoma using a quantitative proteomic approach. J Proteomics 96C:1-12
Kohl M, Megger DA, Trippler M, Meckel H, Ahrens M, Bracht T, Weber F, Hoffmann AC, Baba HA, Sitek B, Schlaak JF, Meyer HE, Stephan C, Eisenacher M. (2013) A practical data processing workflow for multi-OMICS projects. Biochim Biophys Acta 1844:52-62
Ku X, Heinzlmeir S, Liu X, Medard G, Kuster B. (2013) A new chemical probe for quantitative proteomic profiling of fibroblast growth factor receptor and its inhibitors. J Proteomics 96C:44-55
Leon IR, Schwammle V, Jensen ON, Sprenger RR. (2013) Quantitative assessment of in-solution digestion efficiency identifies optimal protocols for unbiased protein analysis. Mol Cell Proteomics 12:2992-3005
Lichti CF, Liu H, Shavkunov AS, Mostovenko E, Sulman EP, Ezhilarasan R, Wang Q, Kroes RA, Moskal JC, Fenyo D, Oksuz BA, Conrad CA, Lang FF, Berven FS, Vegvari A, Rezeli M, Marko-Varga G, Hober S, Nilsson CL (2013) Integrated chromosome 19 transcriptomic and proteomic data sets derived from glioma cancer stem-cell lines. J Proteome Res 13:191-199
Lynskey NN, Goulding D, Gierula M, Turner CE, Dougan G, Edwards RJ, Sriskandan S (2013) RocA truncation underpins hyper-encapsulation, carriage longevity and transmissibility of serotype M18 group A streptococci. PLoS Pathog 9:e1003842
Maller O, Hansen KC, Lyons TR, Acerbi I, Weaver VM, Prekeris R, Tan AC, Schedin P. (2013) Collagen architecture in pregnancy-induced protection from breast cancer. J Cell Sci 126:4108-4110
Marcon C, Lamkemeyer T, Malik WA, Ungrue D, Piepho HP, Hochholdinger F. (2013) Heterosis-associated proteome analyses of maize (Zea mays L.) seminal roots by quantitative label-free LC-MS. J Proteomics 93:295-302
McGouran JF, Gaertner SR, Altun M, Kramer HB, Kessler BM. (2013) Deubiquitinating Enzyme Specificity for Ubiquitin Chain Topology Profiled by Di-Ubiquitin Activity Probes. Chem Biol (in press)
Megger DA, Bracht T, Meyer HE, Sitek B. (2013) Label-free quantification in clinical proteomics. Biochim Biophys Acta 1834:1581-1590
Mehlan H, Schmidt F, Weiss S, Schuler J, Fuchs S, Riedel K, Bernhardt J. (2013) Data visualization in environmental proteomics. Proteomics 13:2805-2821
Merl J, Deeg CA, Swadzba ME, Ueffing M, Hauck SM. (2013) Identification of autoantigens in body fluids by combining pull-downs and organic precipitations of intact immune complexes with quantitative label-free mass spectrometry. J Proteome Res 12:5656-5665
Morrissey B, O'Shea C, Armstrong J, Rooney C, Staunton L, Sheehan M, Shannon AM, Pennington SR. (2013) Development of a label-free LC-MS/MS strategy to approach the identification of candidate protein biomarkers of disease recurrence in prostate cancer patients in a clinical trial of combined hormone and radiation therapy. Proteomics Clin Appl 7:316-326
Mutsaers CA, Lamont DJ, Hunter G, Wishart TM, Gillingwater TH. (2013) Label-free proteomics identifies Calreticulin and GRP75/Mortalin as peripherally accessible protein biomarkers for spinal muscular atrophy. Genome Med 5:95
Nahnsen S, Bielow C, Reinert K, Kohlbacher O. (2013) Tools for label-free peptide quantification. Mol Cell Proteomics 12:549-556
Natunen S, Lampinen M, Suila H, Ritamo I, Pitkanen V, Nairn AV, Rabina J, Laitinen S, Moremen KW, Reutter W, Valmu L. (2013) Metabolic glycoengineering of mesenchymal stromal cells with N-propanoylmannosamine. Glycobiology 23:1004-1012
Olsson N, Carlsson P, James P, Hansson K, Waldemarson S, Malmstrom P, Ferno M, Ryden L, Wingren C, Borrebaeck CA. (2013) Grading breast cancer tissues using molecular portraits. Mol Cell Proteomics 12:3612-3623
Orchard S, Binz PA, Jones AR, Vizcaino JA, Deutsch EW, Hermjakob H. (2013) Preparing to work with big data in proteomics - a report on the HUPO-PSI Spring Workshop: April 15-17, 2013, Liverpool, UK. Proteomics 13:2931-2937
Peffers MJ, Beynon RJ, Clegg PD. (2013) Absolute quantification of selected proteins in the human osteoarthritic secretome. Int J Mol Sci 14:20658-20681
Peltoniemi H, Natunen S, Ritamo I, Valmu L, Rabina J. (2013) Novel data analysis tool for semiquantitative LC-MS-MS2 profiling of N-glycans. Glycoconj J 30:159-170
Quebatte M, Dick MS, Kaever V, Schmidt A, Dehio C. (2013) Dual input control: activation of the Bartonella henselae VirB/D4 type IV secretion system by the stringent sigma factor RpoH1 and the BatR/BatS two-component system. Mol Microbiol 90:756-775
Ramasamy P, Murphy CC, Clynes M, Horgan N, Moriarty P, Tiernan D, Beatty S, Kennedy S, Meleady P. (2013) Proteomics in uveal melanoma. Exp Eye Res 118C:1-12
Ritamo I, Rabina J, Natunen S, Valmu L. (2013) Nanoscale reversed-phase liquid chromatography-mass spectrometry of permethylated N-glycans. Anal Bioanal Chem 405:2469-2480
Rodriguez-Suarez E, Whetton AD. (2013) The application of quantification techniques in proteomics for biomedical research. Mass Spectrom Rev 32:1-26
Schmidt W, Rainville LC, McEneff G, Sheehan D, Quinn B. (2013) A proteomic evaluation of the effects of the pharmaceuticals diclofenac and gemfibrozil on marine mussels (Mytilus spp.): evidence for chronic sublethal effects on stress-response proteins. Drug Test Anal (in press)
Schmutz C, Ahrne E, Kasper CA, Tschon T, Sorg I, Dreier RF, Schmidt A, Arrieumerlou C. (2013) Systems-level overview of host protein phosphorylation during Shigella flexneri infection revealed by phosphoproteomics. Mol Cell Proteomics 12:2952-2968
Skiba NP, Spencer WJ, Salinas RY, Lieu EC, Thompson JW, Arshavsky VY. (2013) Proteomic identification of unique photoreceptor disc components reveals the presence of PRCD, a protein linked to retinal degeneration. J Proteome Res 12:3010-3018
Song W, Mentink RA, Henquet MG, Cordewener JH, van Dijk AD, Bosch D, America AH, van der Krol AR. (2013) N-glycan occupancy of Arabidopsis N-glycoproteins. J Proteomics 93:343-355
Stoop MP, Singh V, Stingl C, Martin R, Khademi M, Olsson T, Hintzen RQ, Luider TM. (2013) Effects of natalizumab treatment on the cerebrospinal fluid proteome of multiple sclerosis patients. J Proteome Res 12:1101-1107
Takahashi D, Kawamura Y, Uemura M. (2013) Changes of detergent-resistant plasma membrane proteins in oat and rye during cold acclimation: association with differential freezing tolerance. J Proteome Res 12:4998-5011
Ternette N, Yang M, Laroyia M, Kitagawa M, O'Flaherty L, Wolhulter K, Igarashi K, Saito K, Kato K, Fischer R, Berquand A, Kessler BM, Lappin T, Frizzell N, Soga T, Adam J, Pollard PJ. (2013) Inhibition of mitochondrial aconitase by succination in fumarate hydratase deficiency. Cell Rep 3:689-700
Trinh HV, Grossmann J, Gehrig P, Roschitzki B, Schlapbach R, Greber UF, Hemmi S. (2013) iTRAQ-Based and Label-Free Proteomics Approaches for Studies of Human Adenovirus Infections. Int J Proteomics 2013:581862
van Ooijen G, Martin SF, Barrios-Llerena ME, Hindle M, Le Bihan T, O'Neill JS, Millar AJ. (2013) Functional analysis of the rodent CK1tau mutation in the circadian clock of a marine unicellular alga. BMC Cell Biol 14:46
von Toerne C, Kahle M, Schafer A, Ispiryan R, Blindert M, Hrabe De Angelis M, Neschen S, Ueffing M, Hauck SM. (2013) Apoe, Mbl2, and Psp plasma protein levels correlate with diabetic phenotype in NZO mice--an optimized rapid workflow for SRM-based quantification. J Proteome Res 12:1331-1343
Vranakis I, Goniotakis I, Psaroulaki A, Sandalakis V, Tselentis Y, Gevaert K, Tsiotis G. (2013) Proteome studies of bacterial antibiotic resistance mechanisms. J Proteomics
Wasinger VC, Zeng M, Yau Y. (2013) Current status and advances in quantitative proteomic mass spectrometry. Int J Proteomics 2013:180605
Weisser H, Nahnsen S, Grossmann J, Nilse L, Quandt A, Brauer H, Sturm M, Kenar E, Kohlbacher O, Aebersold R, Malmstrom L. (2013) An Automated Pipeline for High-Throughput Label-Free Quantitative Proteomics. J Proteome Res (in press)
Wrzesinski K, I RL, Kulej K, Sprenger RR, Bjorndal B, Christensen BJ, Berge RK, O NJ, Rogowska-Wrzesinska A. (2013) Proteomics identifies molecular networks affected by tetradecylthioacetic acid and fish oil supplemented diets. J Proteomics 84:61-77
Xue L, Wang P, Wang L, Renzi E, Radivojac P, Tang H, Arnold R, Zhu JK, Tao WA (2013) Quantitative measurement of phosphoproteome response to osmotic stress in arabidopsis based on Library-Assisted eXtracted Ion Chromatogram (LAXIC). Mol Cell Proteomics 12:2354-2369
Zattas D, Adle DJ, Rubenstein EM, Hochstrasser M. (2013) N-terminal acetylation of the yeast Derlin Der1 is essential for Hrd1 ubiquitin-ligase activity toward luminal ER substrates. Mol Biol Cell 24:890-900
2012
Azimzadeh O, Scherthan H, Yentrapalli R, Barjaktarovic Z, Ueffing M, Conrad M, Neff F, Calzada-Wack J, Aubele M, Buske C, Atkinson MJ, Hauck SM, Tapio S. (2012) Label-free protein profiling of formalin-fixed paraffin-embedded (FFPE) heart tissue reveals immediate mitochondrial impairment after ionising radiation. J Proteomics 75:2384-2395
Bantscheff M, Lemeer S, Savitski MM, Kuster B. (2012) Quantitative mass spectrometry in proteomics: critical review update from 2007 to the present. Anal Bioanal Chem 404:939-965
Bastiaan-Net S, Chanput W, Hertz A, Zwittink RD, Mes JJ, Wichers HJ. (2012) Biochemical and functional characterization of recombinant fungal immunomodulatory proteins (rFIPs). Int Immunopharmacol 15:167-175
Bleijerveld OB, Wijten P, Cappadona S, McClellan EA, Polat AN, Raijmakers R, Sels JW, Colle L, Grasso S, van den Toorn HW, van Breukelen B, Stubbs A, Pasterkamp G, Heck AJ, Hoefer IE, Scholten A. (2012) Deep proteome profiling of circulating granulocytes reveals bactericidal/permeability-increasing protein as a biomarker for severe atherosclerotic coronary stenosis. J Proteome Res 11:5235-5244
Broodman I, de Costa D, Stingl C, Dekker LJ, VanDuijn MM, Lindemans J, van Klaveren RJ, Luider TM. (2012) Mass spectrometry analyses of kappa and lambda fractions result in increased number of complementarity-determining region identifications. Proteomics 12:183-191
Dechavanne C, Guillonneau F, Chiappetta G, Sago L, Levy P, Salnot V, Guitard E, Ehrenmann F, Broussard C, Chafey P, Le Port A, Vinh J, Mayeux P, Dugoujon JM, Lefranc MP, Migot-Nabias F. (2012) Mass Spectrometry Detection of G3m and IGHG3 Alleles and Follow-Up of Differential Mother and Neonate IgG3. PLoS One 7:e46097
Eisenacher M, Kohl M, Turewicz M, Koch MH, Uszkoreit J, Stephan C. (2012) Search and decoy: the automatic identification of mass spectra. Methods Mol Biol 893:445-488
Fila J, Matros A, Radau S, Zahedi RP, Capkova V, Mock HP, Honys D. (2012) Revealing phosphoproteins playing role in tobacco pollen activated in vitro. Proteomics 12:3229-3250
Filiou MD, Martins-de-Souza D, Guest PC, Bahn S, Turck CW. (2012) To label or not to label: applications of quantitative proteomics in neuroscience research. Proteomics 12:736-747
Fischer R, Trudgian DC, Wright C, Thomas G, Bradbury LA, Brown MA, Bowness P, Kessler BM. (2012) Discovery of candidate serum proteomic and metabolomic biomarkers in ankylosing spondylitis. Mol Cell Proteomics 11:M111 013904
Frei AP, Jeon OY, Kilcher S, Moest H, Henning LM, Jost C, Pluckthun A, Mercer J, Aebersold R, Carreira EM, Wollscheid B. (2012) Direct identification of ligand-receptor interactions on living cells and tissues. Nat Biotechnol 30:997-1001
Glatter T, Ludwig C, Ahrne E, Aebersold R, Heck AJ, Schmidt A. (2012) Large-scale quantitative assessment of different in-solution protein digestion protocols reveals superior cleavage efficiency of tandem Lys-C/trypsin proteolysis over trypsin digestion. J Proteome Res 11:5145-5156
Greer E, Martin AC, Pendle A, Colas I, Jones AM, Moore G, Shaw P. (2012) The Ph1 locus suppresses Cdk2-type activity during premeiosis and meiosis in wheat. Plant Cell 24:152-162
Gundisch S, Hauck S, Sarioglu H, Schott C, Viertler C, Kap M, Schuster T, Reischauer B, Rosenberg R, Verhoef C, Mischinger HJ, Riegman P, Zatloukal K, Becker KF. (2012) Variability of protein and phosphoprotein levels in clinical tissue specimens during the preanalytical phase. J Proteome Res 11:5748-5762
Hauck SM, Hofmaier F, Dietter J, Swadzba ME, Blindert M, Amann B, Behler J, Kremmer E, Ueffing M, Deeg CA. (2012) Label-free LC-MSMS analysis of vitreous from autoimmune uveitis reveals a significant decrease in secreted Wnt signalling inhibitors DKK3 and SFRP2. J Proteomics 75:4545-4554
Hummel M, Cordewener JH, de Groot JC, Smeekens S, America AH, Hanson J. (2012) Dynamic protein composition of Arabidopsis thaliana cytosolic ribosomes in response to sucrose feeding as revealed by label free MSE proteomics. Proteomics 12:1024-1038
Karhemo PR, Ravela S, Laakso M, Ritamo I, Tatti O, Makinen S, Goodison S, Stenman UH, Holtta E, Hautaniemi S, Valmu L, Lehti K, Laakkonen P. (2012) An optimized isolation of biotinylated cell surface proteins reveals novel players in cancer metastasis. J Proteomics 77:87-100
Kolmeder CA, de Been M, Nikkila J, Ritamo I, Matto J, Valmu L, Salojarvi J, Palva A, Salonen A, de Vos WM. (2012) Comparative metaproteomics and diversity analysis of human intestinal microbiota testifies for its temporal stability and expression of core functions. PLoS One 7:e29913
Krishnan N, Lam TT, Fritz A, Rempinski D, O'Loughlin K, Minderman H, Berezney R, Marzluff WF, Thapar R. (2012) The prolyl isomerase Pin1 targets stem-loop binding protein (SLBP) to dissociate the SLBP-histone mRNA complex linking histone mRNA decay with SLBP ubiquitination. Mol Cell Biol 32:4306-4322
Lemeer S, Hahne H, Pachl F, Kuster B. (2012) Software tools for MS-based quantitative proteomics: a brief overview. Methods Mol Biol 893:489-499
Li B, Takahashi D, Kawamura Y, Uemura M. (2012) Comparison of plasma membrane proteomic changes of Arabidopsis suspension-cultured cells (T87 Line) after cold and ABA treatment in association with freezing tolerance development. Plant Cell Physiol 53:543-554
Maat P, VanDuijn M, Brouwer E, Dekker L, Zeneyedpour L, Luider T, Smitt PS. (2012) Mass spectrometric detection of antigen-specific immunoglobulin peptides in paraneoplastic patient sera. J Autoimmun 38:354-360
Marguerat S, Schmidt A, Codlin S, Chen W, Aebersold R, Bahler J. (2012) Quantitative Analysis of Fission Yeast Transcriptomes and Proteomes in Proliferating and Quiescent Cells. Cell 151:671-683
McDonagh B, Dominguez-Martin MA, Gomez-Baena G, Lopez-Lozano A, Diez J, Barcena JA, Garcia Fernandez JM. (2012) Nitrogen starvation induces extensive changes in the redox proteome of Prochlorococcus sp. strain SS120. Environ Microbiol Rep 4:257-267
Meding S, Balluff B, Elsner M, Schone C, Rauser S, Nitsche U, Maak M, Schafer A, Hauck SM, Ueffing M, Langer R, Hofler H, Friess H, Rosenberg R, Walch A. (2012) Tissue-based proteomics reveals FXYD3, S100A11 and GSTM3 as novel markers for regional lymph node metastasis in colon cancer. J Pathol (In press)
Meleady P, Gallagher M, Clarke C, Henry M, Sanchez N, Barron N, Clynes M. (2012) Impact of miR-7 over-expression on the proteome of Chinese hamster ovary cells. J Biotechnol 160:251-262
Merl J, Ueffing M, Hauck SM, von Toerne C. (2012) Direct comparison of MS-based label-free and SILAC quantitative proteome profiling strategies in primary retinal Muller cells. Proteomics 12:1902-1911
Michaux C, Martini C, Shioya K, Ahmed Lecheheb S, Budin-Verneuil A, Cosette P, Sanguinetti M, Hartke A, Verneuil N, Giard JC. (2012) CspR, a cold shock RNA-binding protein involved in the long-term survival and the virulence of Enterococcus faecalis. J Bacteriol 194:6900-6908
Mustafa DA, Dekker LJ, Stingl C, Kremer A, Stoop M, Sillevis Smitt PA, Kros JM, Luider TM. (2012) A proteome comparison between physiological angiogenesis and angiogenesis in glioblastoma. Mol Cell Proteomics 11:M111 008466
Nesvizhskii AI. (2012) Computational and informatics strategies for identification of specific protein interaction partners in affinity purification mass spectrometry experiments. Proteomics 12:1639-1655
O'Connell K, Prencipe M, O'Neill A, Corcoran C, Rani S, Henry M, Dowling P, Meleady P, O'Driscoll L, Watson W, O'Connor R. (2012) The use of LC-MS to identify differentially expressed proteins in docetaxel-resistant prostate cancer cell lines. Proteomics 12:2115-2126
Peltoniemi H, Natunen S, Ritamo I, Valmu L, Rabina J. (2012) Novel data analysis tool for semiquantitative LC-MS-MS2 profiling of N-glycans. Glycoconj J 30:159-170
Qi D, Brownridge P, Xia D, Mackay K, Gonzalez-Galarza FF, Kenyani J, Harman V, Beynon RJ, Jones AR. (2012) A software toolkit and interface for performing stable isotope labeling and top3 quantification using Progenesis LC-MS. Omics 16:489-495
Rosenling T, Stoop MP, Attali A, van Aken H, Suidgeest E, Christin C, Stingl C, Suits F, Horvatovich P, Hintzen RQ, Tuinstra T, Bischoff R, Luider TM. (2012) Profiling and identification of cerebrospinal fluid proteins in a rat EAE model of multiple sclerosis. J Proteome Res 11:2048-2060
Sitek B, Waldera-Lupa DM, Poschmann G, Meyer HE, Stuhler K. (2012) Application of label-free proteomics for differential analysis of lung carcinoma cell line A549. Methods Mol Biol 893:241-248
Stoop MP, Rosenling T, Attali A, Meesters RJ, Stingl C, Dekker LJ, van Aken H, Suidgeest E, Hintzen RQ, Tuinstra T, van Gool A, Luider TM, Bischoff R. (2012) Minocycline effects on the cerebrospinal fluid proteome of experimental autoimmune encephalomyelitis rats. J Proteome Res 11:4315-4325
Street JM, Barran PE, Mackay CL, Weidt S, Balmforth C, Walsh TS, Chalmers RT, Webb DJ, Dear JW. (2012) Identification and proteomic profiling of exosomes in human cerebrospinal fluid. J Transl Med 10:5
Takahashi D, Kawamura Y, Yamashita T, Uemura M. (2012) Detergent-resistant plasma membrane proteome in oat and rye: similarities and dissimilarities between two monocotyledonous plants. J Proteome Res 11:1654-1665
Tuli L, Tsai TH, Varghese RS, Xiao JF, Cheema A, Ressom HW. (2012) Using a spike-in experiment to evaluate analysis of LC-MS data. Proteome Sci 10:13
Vasilj A, Gentzel M, Ueberham E, Gebhardt R, Shevchenko A. (2012) Tissue proteomics by one-dimensional gel electrophoresis combined with label-free protein quantification. J Proteome Res 11:3680-3689
Wang H, Alvarez S, Hicks LM. (2012) Comprehensive comparison of iTRAQ and label-free LC-based quantitative proteomics approaches using two Chlamydomonas reinhardtii strains of interest for biofuels engineering. J Proteome Res 11:487-501
Zimmer A, Bouley J, Le Mignon M, Pliquet E, Horiot S, Turfkruyer M, Baron-Bodo V, Horak F, Nony E, Louise A, Moussu H, Mascarell L, Moingeon P. (2012) A regulatory dendritic cell signature correlates with the clinical efficacy of allergen-specific sublingual immunotherapy. J Allergy Clin Immunol 129:1020-1030
2011
Altun M, Kramer HB, Willems LI, McDermott JL, Leach CA, Goldenberg SJ, Kumar KG, Konietzny R, Fischer R, Kogan E, Mackeen MM, McGouran J, Khoronenkova SV, Parsons JL, Dianov GL, Nicholson B, Kessler BM. (2011) Activity-based chemical proteomics accelerates inhibitor development for deubiquitylating enzymes. Chem Biol 18:1401-1412
Beck F, Lewandrowski U, Wiltfang M, Feldmann I, Geiger J, Sickmann A, Zahedi RP. (2011) The good, the bad, the ugly: validating the mass spectrometric analysis of modified peptides. Proteomics 11:1099-1109
Blanchet L, Smolinska A, Attali A, Stoop MP, Ampt KA, van Aken H, Suidgeest E, Tuinstra T, Wijmenga SS, Luider T, Buydens LM. (2011) Fusion of metabolomics and proteomics data for biomarkers discovery: case study on the experimental autoimmune encephalomyelitis. BMC Bioinformatics 12:254
Christin C, Bischoff R, Horvatovich P. (2011) Data processing pipelines for comprehensive profiling of proteomics samples by label-free LC-MS for biomarker discovery. Talanta 83:1209-1224
Dekker LJ, Zeneyedpour L, Brouwer E, van Duijn MM, Sillevis Smitt PA, Luider TM. (2011) An antibody-based biomarker discovery method by mass spectrometry sequencing of complementarity determining regions. Anal Bioanal Chem 399:1081-1091
English JA, Pennington K, Dunn MJ, Cotter DR. (2011) The neuroproteomics of schizophrenia. Biol Psychiatry 69:163-172
Gil GC, Brennan J, Throckmorton DJ, Branda SS, Chirica GS. (2011) Automated analysis of mouse serum peptidome using restricted access media and nanoliquid chromatography-tandem mass spectrometry. J Chromatogr B Analyt Technol Biomed Life Sci 879:1112-1120
Glatter T, Schittenhelm RB, Rinner O, Roguska K, Wepf A, Junger MA, Kohler K, Jevtov I, Choi H, Schmidt A, Nesvizhskii AI, Stocker H, Hafen E, Aebersold R, Gstaiger M. (2011) Modularity and hormone sensitivity of the Drosophila melanogaster insulin receptor/target of rapamycin interaction proteome. Mol Syst Biol 7:547
Hansson J, Panchaud A, Favre L, Bosco N, Mansourian R, Benyacoub J, Blum S, Jensen ON, Kussmann M. (2011) Time-resolved quantitative proteome analysis of in vivo intestinal development. Mol Cell Proteomics 10:M110 005231
Ijsselstijn L, Dekker LJ, Koudstaal PJ, Hofman A, Sillevis Smitt PA, Breteler MM, Luider TM. (2011) Serum clusterin levels are not increased in presymptomatic Alzheimer's disease. J Proteome Res 10:2006-2010
Ijsselstijn L, Dekker LJ, Stingl C, van der Weiden MM, Hofman A, Kros J, Koudstaal PJ, Sillevis Smitt P, Ikram MA, Breteler MM, Luider T. (2011) Serum Levels of Pregnancy Zone Protein are Elevated in Presymptomatic Alzheimer's Disease. J Proteome Res 10:4902-4910
Le Bihan T, Martin SF, Chirnside ES, van Ooijen G, Barrios-Llerena ME, O'Neill JS, Shliaha PV, Kerr LE, Millar AJ. (2011) Shotgun proteomic analysis of the unicellular alga Ostreococcus tauri. J Proteomics 74:2060-2070
Levin Y. (2011) The role of statistical power analysis in quantitative proteomics. Proteomics 11:2565-2567
Matros A, Kaspar S, Witzel K, Mock HP. (2011) Recent progress in liquid chromatography-based separation and label-free quantitative plant proteomics. Phytochemistry 72:963-974
Mutsaers CA, Wishart TM, Lamont DJ, Riessland M, Schreml J, Comley LH, Murray LM, Parson SH, Lochmuller H, Wirth B, Talbot K, Gillingwater TH. (2011) Reversible molecular pathology of skeletal muscle in spinal muscular atrophy. Hum Mol Genet 20:4334-4344
Neilson KA, Ali NA, Muralidharan S, Mirzaei M, Mariani M, Assadourian G, Lee A, van Sluyter SC, Haynes PA. (2011) Less label, more free: approaches in label-free quantitative mass spectrometry. Proteomics 11:535-553
Olsson N, Wingren C, Mattsson M, James P, O'Connell D, Nilsson F, Cahill DJ, Borrebaeck CA. (2011) Proteomic analysis and discovery using affinity proteomics and mass spectrometry. Mol Cell Proteomics 10:M110 003962
Prapainop K, Wentworth P, Jr. (2011) A shotgun proteomic study of the protein corona associated with cholesterol and atheronal-B surface-modified quantum dots. Eur J Pharm Biopharm 77:353-359
Reiland S, Finazzi G, Endler A, Willig A, Baerenfaller K, Grossmann J, Gerrits B, Rutishauser D, Gruissem W, Rochaix JD, Baginsky S. (2011) Comparative phosphoproteome profiling reveals a function of the STN8 kinase in fine-tuning of cyclic electron flow (CEF). Proc Natl Acad Sci U S A 108:12955-12960
Sandin M, Krogh M, Hansson K, Levander F. (2011) Generic workflow for quality assessment of quantitative label-free LC-MS analysis. Proteomics 11:1114-1124
Schmidt A, Beck M, Malmstrom J, Lam H, Claassen M, Campbell D, Aebersold R. (2011) Absolute quantification of microbial proteomes at different states by directed mass spectrometry. Mol Syst Biol 7:510
Titulaer MK, de Costa D, Stingl C, Dekker LJ, Sillevis Smitt PA, Luider TM. (2011) Label-free peptide profiling of Orbitrap full mass spectra. BMC Res Notes 4:21
Trudgian DC, Ridlova G, Fischer R, Mackeen MM, Ternette N, Acuto O, Kessler BM, Thomas B. (2011) Comparative evaluation of label-free SINQ normalized spectral index quantitation in the central proteomics facilities pipeline. Proteomics 11:2790-2797
Wada K, Ogiwara A, Nagasaka K, Tanaka N, Komatsu Y. (2011) i-RUBY: a novel software for quantitative analysis of highly accurate shotgun-proteomics liquid chromatography/tandem mass spectrometry data obtained without stable-isotope labeling of proteins. Rapid Commun Mass Spectrom 25:960-968
Wiedl T, Arni S, Roschitzki B, Grossmann J, Collaud S, Soltermann A, Hillinger S, Aebersold R, Weder W. (2011) Activity-based proteomics: identification of ABHD11 and ESD activities as potential biomarkers for human lung adenocarcinoma. J Proteomics 74:1884-1894
Wu Z, Doondeea JB, Gholami AM, Janning MC, Lemeer S, Kramer K, Eccles SA, Gollin SM, Grenman R, Walch A, Feller SM, Kuster B. (2011) Quantitative chemical proteomics reveals new potential drug targets in head and neck cancer. Mol Cell Proteomics 10:M111 011635
Yun BW, Feechan A, Yin M, Saidi NB, Le Bihan T, Yu M, Moore JW, Kang JG, Kwon E, Spoel SH, Pallas JA, Loake GJ. (2011) S-nitrosylation of NADPH oxidase regulates cell death in plant immunity. Nature 478:264-268
Zabron AA, Horneffer-van der Sluis VM, Wadsworth CA, Laird F, Gierula M, Thillainayagam AV, Vlavianos P, Westaby D, Taylor-Robinson SD, Edwards RJ, Khan SA. (2011) Elevated levels of neutrophil gelatinase-associated lipocalin in bile from patients with malignant pancreatobiliary disease. Am J Gastroenterol 106:1711-1717
2010
Abdul-Salam VB, Wharton J, Cupitt J, Berryman M, Edwards RJ, Wilkins MR. (2010) Proteomic analysis of lung tissues from patients with pulmonary arterial hypertension. Circulation 122:2058-2067
Affolter M, Grass L, Vanrobaeys F, Casado B, Kussmann M. (2010) Qualitative and quantitative profiling of the bovine milk fat globule membrane proteome. J Proteomics 73:1079-1088
de Costa D, Broodman I, Vanduijn MM, Stingl C, Dekker LJ, Burgers PC, Hoogsteden HC, Sillevis Smitt PA, van Klaveren RJ, Luider TM. (2010) Sequencing and quantifying IgG fragments and antigen-binding regions by mass spectrometry. J Proteome Res 9:2937-2945
Dicker L, Lin X, Ivanov AR. (2010) Increased power for the analysis of label-free LC-MS/MS proteomics data by combining spectral counts and peptide peak attributes. Mol Cell Proteomics 9:2704-2718
Dowsey AW, English JA, Lisacek F, Morris JS, Yang GZ, Dunn MJ. (2010) Image analysis tools and emerging algorithms for expression proteomics. Proteomics 10:4226-4257
Hauck SM, Dietter J, Kramer RL, Hofmaier F, Zipplies JK, Amann B, Feuchtinger A, Deeg CA, Ueffing M. (2010) Deciphering membrane-associated molecular processes in target tissue of autoimmune uveitis by label-free quantitative mass spectrometry. Mol Cell Proteomics 9:2292-2305
Lalmahomed Z, Roest H, Woltman K, Verhoef C, Luider T, IJzermans J (2010) Identification of urinary biomarkers for colorectal liver metastases. Annals of Oncology 21 (supplement 6): vi31–vi121
Levin Y, Bahn S. (2010) Quantification of proteins by label-free LC-MS/MS. Methods Mol Biol 658:217-231
Magdeldin S, Li H, Yoshida Y, Satokata I, Maeda Y, Yokoyama M, Enany S, Zhang Y, Xu B, Fujinaka H, Yaoita E, Yamamoto T. (2010) Differential proteomic shotgun analysis elucidates involvement of water channel aquaporin 8 in presence of alpha-amylase in the colon. J Proteome Res 9:6635-6646
Nanjo Y, Skultety L, Ashraf Y, Komatsu S. (2010) Comparative proteomic analysis of early-stage soybean seedlings responses to flooding by using gel and gel-free techniques. J Proteome Res 9:3989-4002
Stoop MP, Coulier L, Rosenling T, Shi S, Smolinska AM, Buydens L, Ampt K, Stingl C, Dane A, Muilwijk B, Luitwieler RL, Sillevis Smitt PA, Hintzen RQ, Bischoff R, Wijmenga SS, Hankemeier T, van Gool AJ, Luider TM. (2010) Quantitative proteomics and metabolomics analysis of normal human cerebrospinal fluid samples. Mol Cell Proteomics 9:2063-2075
Strassberger V, Fugmann T, Neri D, Roesli C. (2010) Chemical proteomic and bioinformatic strategies for the identification and quantification of vascular antigens in cancer. J Proteomics 73:1954-1973
VanDuijn MM, Dekker LJ, Zeneyedpour L, Smitt PA, Luider TM. (2010) Immune responses are characterized by specific shared immunoglobulin peptides that can be detected by proteomic techniques. J Biol Chem 285:29247-29253
Zhang R, Barton A, Brittenden J, Huang J, Crowther D. (2010) Evaluation for Computational Platforms of LC-MS Based Label-Free Quantitative Proteomics: A Global View. J Proteomics Bioinform 3:260-265
2009
Dakna M, He Z, Yu WC, Mischak H, Kolch W. (2009) Technical, bioinformatical and statistical aspects of liquid chromatography-mass spectrometry (LC-MS) and capillary electrophoresis-mass spectrometry (CE-MS) based clinical proteomics: a critical assessment. J Chromatogr B Analyt Technol Biomed Life Sci 877:1250-1258





