How can I export multi-fraction experiment results for use in another graphing or stats package?
In your recombined fractionation experiment proceed with the analysis as far as the Review Proteins screen.
On this screen you can export three different types of results:
- Protein measurements
- Peptide measurements
- Peptide ion measurements
Each of these exports are accessed via the File menu; the image below shows the option to Export protein measurements. In all cases, the data are only exported for the active tag filter (by default) and experimental grouping, but the values are not recalculated in any way – the restriction is purely a content filter. To export all data: i) use a grouping that contains all runs to avoid restricting the columns; and ii) clear any active tag filters.
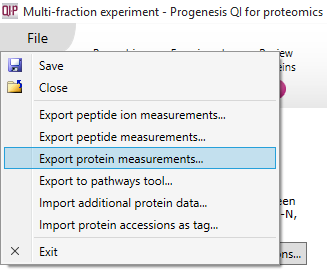
Exporting protein measurements from Review Proteins (multi-fraction experiment).
For each of these three exports you will be presented with a content-selection dialog, allowing you to tailor what values are included in your export. For example, Export protein measurements:
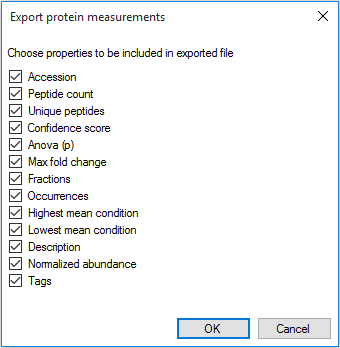
The column selection dialog box for Export protein measurements (multi-fraction experiment).
All of these exports produce data in the csv format.





