Support for ChemSpider

About this plug-in
ChemSpider is a web-based chemical structure database with access to over 32 million structures from hundreds of data sources. This method makes use of those ChemSpider web services, automatically exporting data from Progenesis QI to ChemSpider for searching according to the parameters you select, importing the results, and assigning them against the correct compounds within the software. This service is intended for users with a valid support contract, or users who are evaluating, and may be restricted to those users in the future.
Selecting this search method
At the Identify Compounds step of the workflow, use the drop-down menu to select ChemSpider:
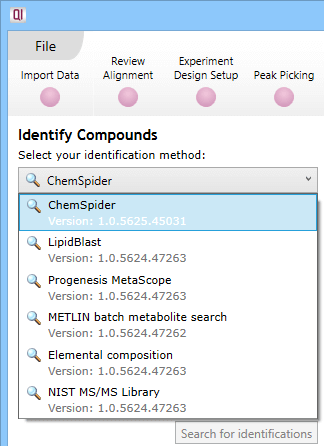
The drop-down selection dialog at Identify Compounds.
Filter the compounds
You may wish to select a filtered set of compounds for this process, to reduce the processing and curation time. This is because ChemSpider can search a very large range of data and return a large number of results depending upon the search settings. This can be done using tag filtering. For example, you may wish to only search compounds that are significantly altered between control and treatment groups. The search will be carried out for all compounds present in the current filter (or all compounds in the experiment, if no filter is applied).
Set the search parameters
The search parameters are grouped into parameter sets. A default parameter set is provided, with suggested settings. You can use this default set, customise it, or create your own.
Search parameter set selection.
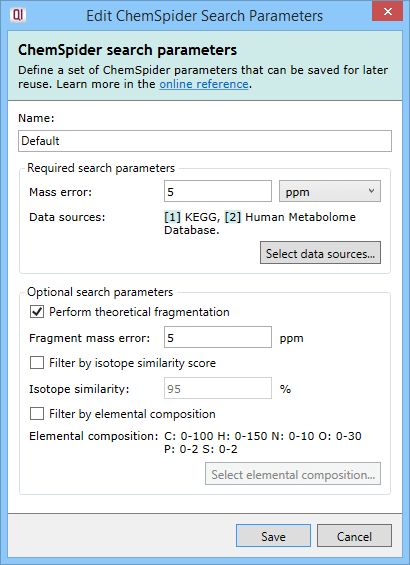
Search parameter set dialog.
The ChemSpider method requires just two selections to be made: the mass error tolerance and the data sources to be searched.
The mass error tolerance, in ppm or Da, is the maximum difference between the mass for the compound in the databases searched and the observed mass. Where a compound has been successfully deconvoluted, the observed neutral mass is searched directly against the database neutral mass.
Where only a single m/z is available, and its adduct form is unclear, a single neutral mass cannot be assigned for searching against the database(s). Instead, this compound will be searched multiple times. Each time, it is treated as if it is derived from a different adduct; the neutral mass corresponding to that original m/z and assumed adduct is calculated and searched, and this is repeated for all the adducts you defined in the experiment setup that could impart the correct observed charge state for the compound. This allows attempted identification even for compounds without a known neutral mass.
The second parameter is the selection of one or more data sources from the available list. On clicking the Select data sources button (as seen in the image above), you will see the following dialog:
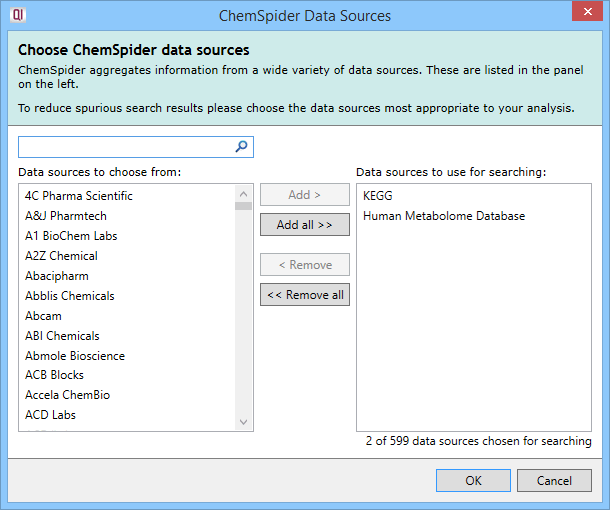
The ChemSpider Data Sources selection box that appears on clicking on the Select data sources button.
You can select or deselect data sources individually, use multiple selections (e.g. Shft- or Ctrl-clicking), select or deselect all data sources at once, and can track down data sources of interest using the Search box. Once selected, the data sources are listed in the parameter set dialog and on the main page in the search parameter set drop-down, as was shown earlier (e.g. KEGG, Human Metabolome Database). Up to five will be listed by name, and any after that presented as "...and X more".
The next three selections—whether or not to perform theoretical fragmentation on the results or to perform two different kinds of filtering—are optional.
Theoretical fragmentation is performed using the same algorithm as in the MetaScope plug-in—your experimental fragmentation data is compared to theoretical fragmentation patterns generated by the simulated breaking of bonds in the structures of possible identifications and a fragmentation score is assigned to them. If you choose to perform theoretical fragmentation, you'll need to specify the fragment mass error tolerance in ppm, which is the maximum difference between the mass of a fragment as determined by the fragmentation algorithm and the observed mass of the fragment.
Isotope similarity filtering discards the results whose isotope similarity score is below the configured cut-off value. The score is calculated between the theoretical and observed isotope patterns in the same manner as for the MetaScope scoring engine.
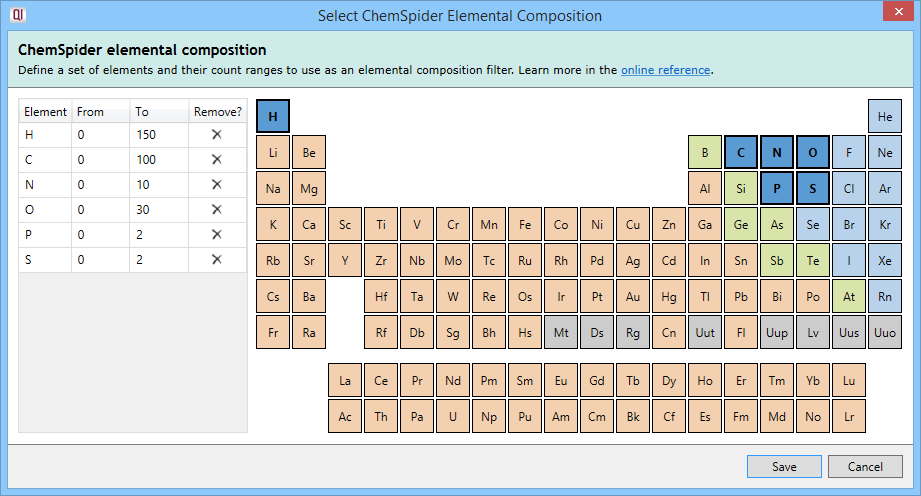
Elemental composition filtering compares the formula returned by ChemSpider to a specification consisting of elements and their allowed count ranges. Results that do not conform to the specification are dropped. On clicking the Select elemental composition button, you will see the following dialog:
Clicking on an element in the periodic table will add it to the specification, allowing it to be considered as part of the possible molecular formulae. The element will then be listed in the left table.
The From and To fields in that table are customisable and allow you to specify the minimum and maximum number of atoms of each element (range) in the molecular formulae.
To remove an element from the specification, you can press the Delete key with that element selected in the left table, click on the cross button in the Remove? column of that table, or click on the element in the periodic table a second time.
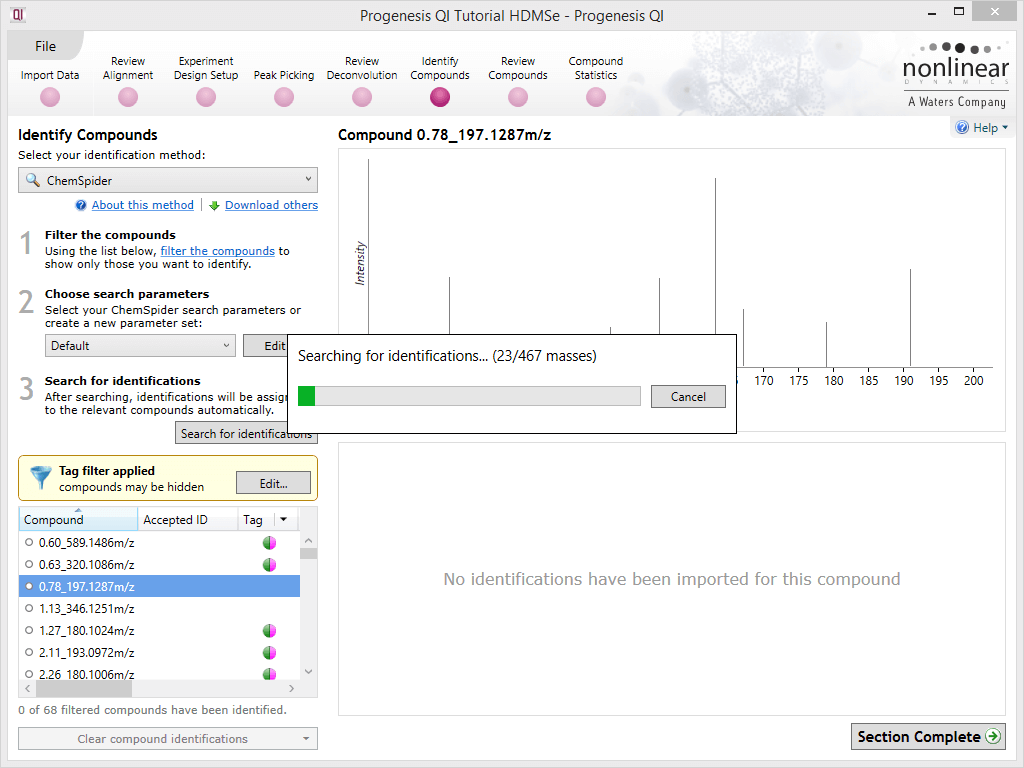
Search for identifications
Once your parameters are set up, clicking on this button will begin the search process.
Minimising search time
Given the number of databases (potentially) searched, ChemSpider searches can take a long time to complete. To minimise this, you can pre-filter your compounds, as described above. You may also wish to narrow your database selection only to highly relevant databases rather than including a larger number, and narrow your mass error tolerance as much as is reasonably possible for your instrument. These measures will reduce the search space, and return results more rapidly.
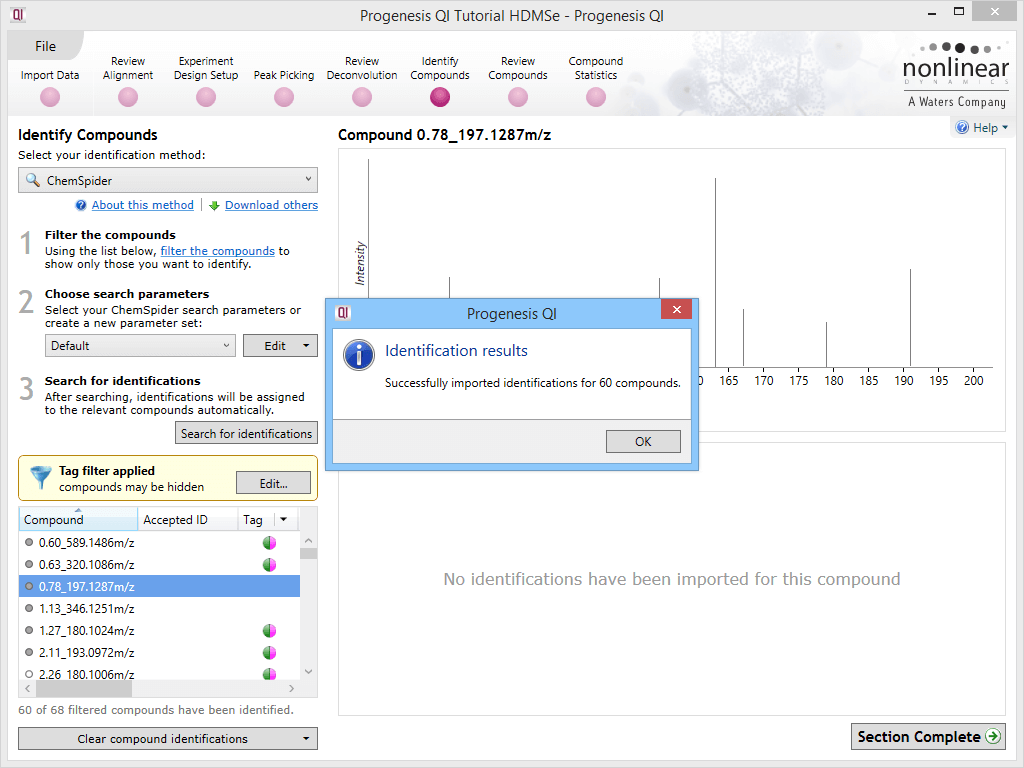
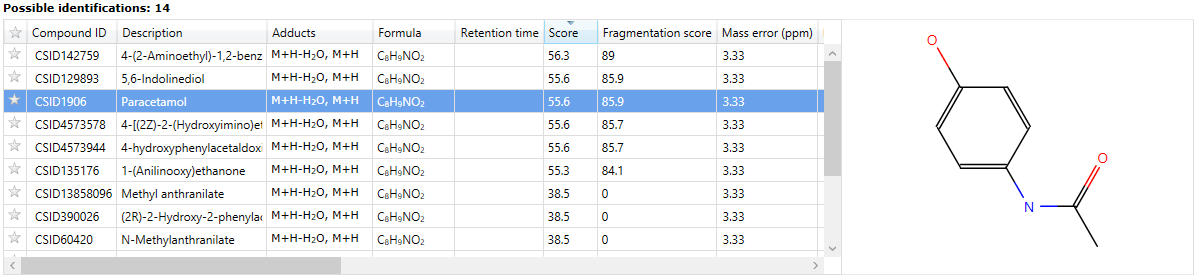
Results
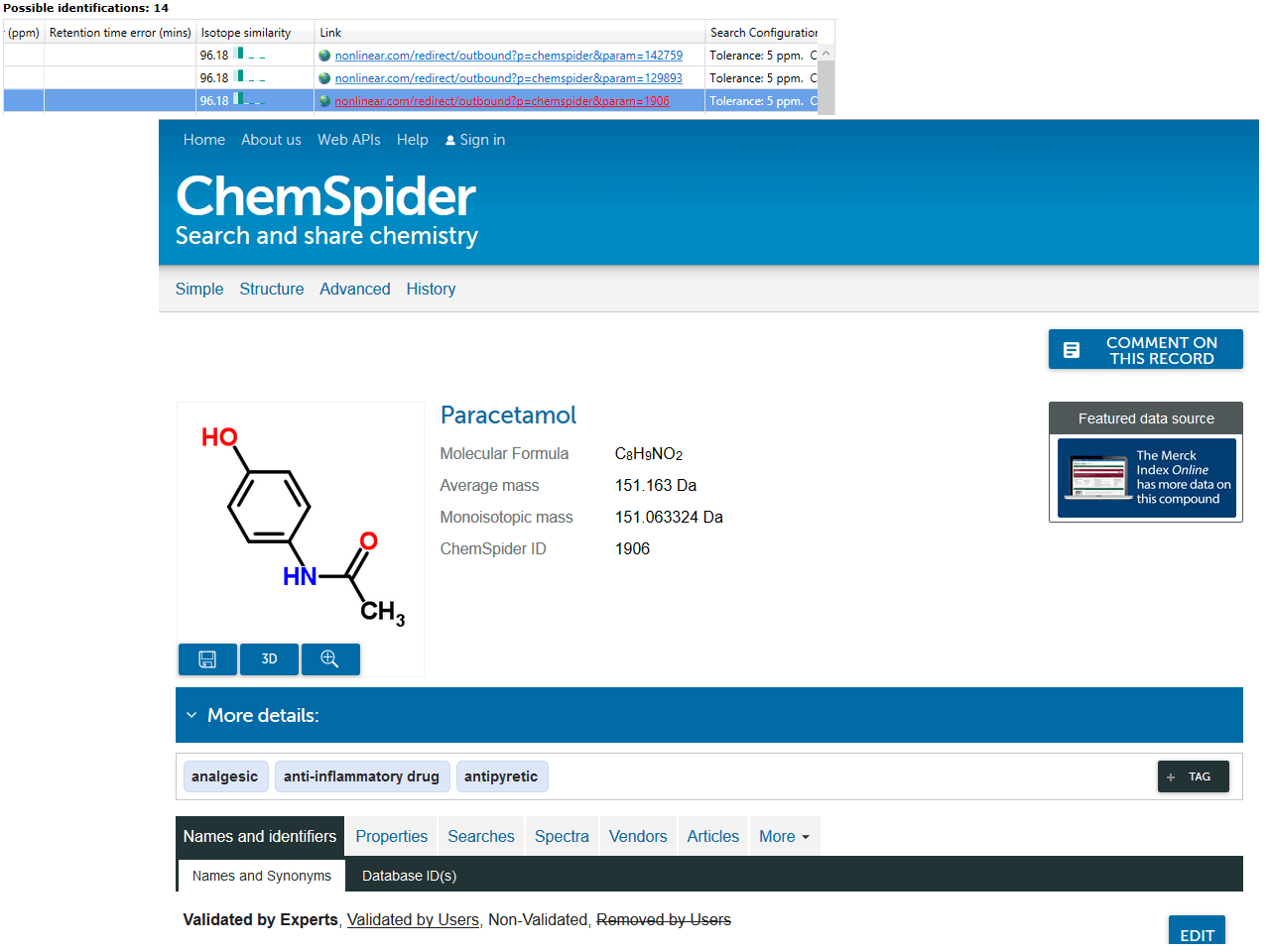
Identifications are obtained, then imported and matched against the relevant compounds automatically. The hits from a given compound will be denoted by ChemSpider IDs (CSIDs); these values will return more information if entered as search terms into ChemSpider, including the data sources for the compound. To save manually entering the data, the Link field of the table also provides a direct path to the relevant page at ChemSpider.
Multiple searches
Note that if you perform one ChemSpider search but select multiple databases, this is imported as one search by the software. The external database that the identity was discovered in is not marked, as the CSID is a unique identifier across all of ChemSpider. To determine where a ChemSpider hit resulted from in this case, the CSID or link should be used to query ChemSpider directly, where the data sources containing that CSID are displayed.
If you perform two separate ChemSpider searches, with different parameters, and obtain new identification data, then results from the two searches will be assigned a unique letter by the search they came from, to differentiate which results came from which search.
When letters are assigned, a compound that would have been identified by more than one of the searches would be given the letter of the first search that discovered it (i.e. a compound returned by both searches A and B carried out in that order would be assigned the identifier for search A).