What are additional compound properties files and how do I use them?
Progenesis MetaScope provides the ability to augment SDF or CSV compound databases with so-called Additional compound properties files.
These allow you to separate out properties provided in externally sourced databases (e.g. neutral mass and structure), from those produced in-house (e.g. retention time and CCS), thereby allowing you to update the externally sourced part without losing these additional properties.
Contents
- What is an additional compound properties file?
- Why use an additional compound properties file?
- How do I use an additional compound properties file?
- How do I create my own additional compound properties file?
- How do I append to an existing additional compound properties file?
What is an additional compound properties file?
An additional compound properties file is simply a CSV file. Each row represents a particular adducted form of a particular compound, and it contains the following columns:
| Column name | Description |
|---|---|
| Name | The common name of the compound |
| Compound ID | The unique compound ID (this may be a MassBank ID, PubChem ID, HMDB ID etc.) |
| Retention time (min) | The retention time of this compound (measured in minutes) |
| CCS (angstrom^2) | The Collisional Cross Sectional area of this particular adduct of the given compound (measured in Å2) |
| Adduct | The adduct form that this row refers to. The format of this field is like in these examples: "M+H", "M-NH4" |
For example, here is the contents of the Analgesic Mix - RT & CCS.csv file included with the installation of Progenesis QI:
| Name | Compound ID | Retention time (min) | CCS (angstrom^2) | Adduct |
|---|---|---|---|---|
| Acetanilide | 14708992 | 4.5 | 115.1 | M+H |
| Paracetamol | 46506142 | 3.1 | 118.8 | M+H |
| Caffeine | 46511425 | 3.78 | 129.4 | M+H |
| Phenacetin | 49854487 | 5.22 | 132.6 | M+H |
An additional compound properties file allows you to store retention time and CCS values separately from your main SDF or CSV database file.
Why use an additional compound properties file?
There are a few of advantages to using additional compound properties files rather than entering the information directly in an SDF file:
- Separation of intrinsic and non-intrinsic properties
- Using an additional compound properties file, you separate those properties of a compound which are intrinsic and constant for everyone, from those that are specific to your experimental method/machine etc. Thus, you can still share your main SDF database, without mixing in properties that are likely to be different for other researchers.
- Ease of updating externally sourced compound databases
- If you obtain an SDF file from an external source (e.g. ChemSpider), and later wish to update this SDF file to a newer version provided by ChemSpider, the use of an additional compound properties file means you won't have to re-enter all your RT and CCS values in the updated SDF.
- Ability to record adduct specific data
- Different adducted forms of a compound may have different CCS values, and storing this in an SDF file (which only concerns the neutral compound) is problematic. By using an additional compound properties file, it becomes easy to record the different adduct values.
How do I use an additional compound properties file?
To use an additional compound properties file in a MetaScope search, you simply need to choose the file to use in the Read additional compound properties from this file section of the MetaScope search parameters dialog, as shown here:
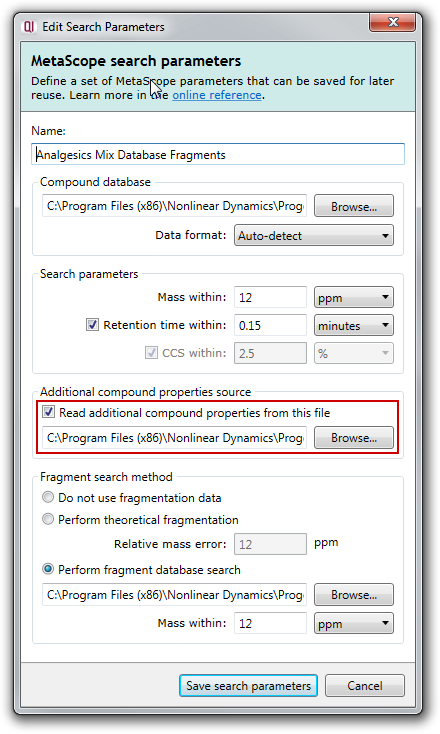
Note that the CCS within option is disabled in the screenshot above. Progenesis QI will detect whether your experiment contains CCS data, and only enable this option if it does.
For every compound present in the SDF or CSV database, Progenesis QI will look in the additional compound properties file for a row with a matching compound ID (or multiple rows for multiple adducts). If a matching row is found, the retention time and CCS values therein will be compared to your experimental measured values.
The retention time and CCS values read from the additional compound properties file will be displayed in the Possible identifications area, along with the difference between the measured values.
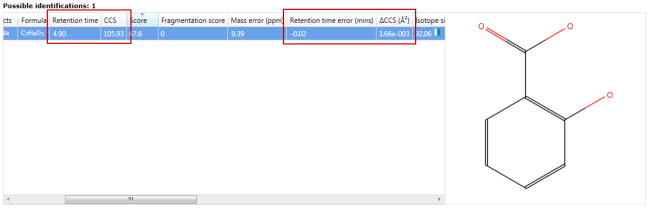
You can also choose to enable or disable searching by retention time and CCS in the Search parameters section of the MetaScope search parameters dialog:
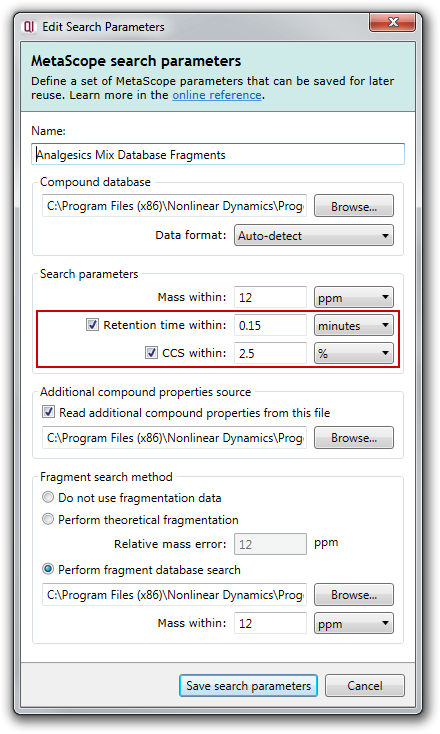
How do I create my own additional compound properties file?
To create an additional compound properties file using measured retention times and CCS values, first follow the normal identification workflow.
Once you have accepted your compound identifications, choose the Export additional compound properties... item in the File menu:
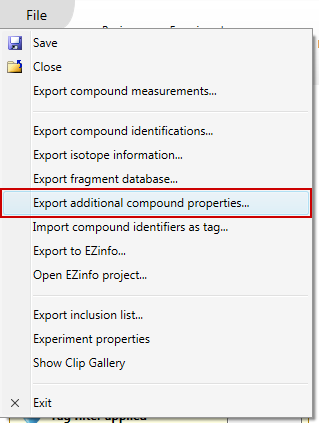
This will bring up a dialog showing you all of your accepted compound identifications. You can choose to export all or a subset of these by using the checkboxes in the first column:
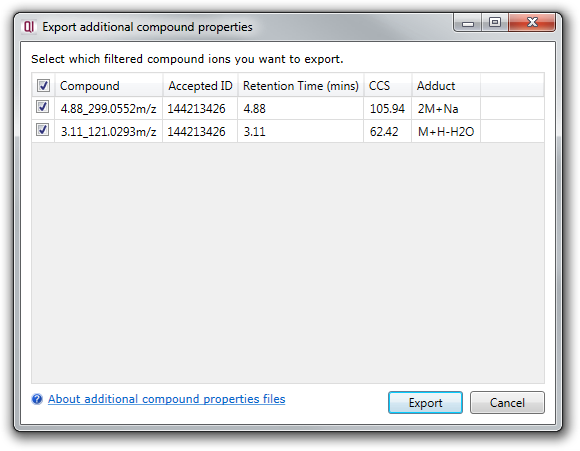
Clicking the Export button will prompt you to choose a filename, then carry out the export for you.
You can now use the generated file to assist searches in future experiments. Simply browse to the generated CSV file in the Additional compound properties source section of the MetaScope search parameters dialog (see above).
How do I append to an existing additional compound properties file?
Suppose you have already generated your additional compound properties file using the above method, but you wish to extend it by adding new data from a recent experiment.
This is easy to do in Progenesis QI. When exporting the additional compound properties file, choose the existing CSV file when prompted for a filename.
Progenesis QI will then ask you if you wish to overwrite or append to the existing file:
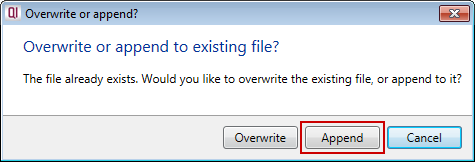
To append your new data, choose Append, and the new data will be added to the end of your additional compound properties file.
Warning: Choosing Overwrite in the above dialog will completely overwrite the existing file. Make sure this is what you intend to do before choosing this option, and we recommend making a backup of the file just in case you need it again.






