Support for Metabolic Profiling CCS Library

About this plug-in
The Metabolic Profiling CCS Library search plugin bundles Waters Metabolic Profiling CCS libraries and performs a combination of neutral mass, MS/MS and CCS based searches. It uses the same search and scoring algorithms as Progenesis MetaScope.
Set the search parameters
Searches using the Metabolic Profiling CCS Library search plugin require the selection of three search parameters:
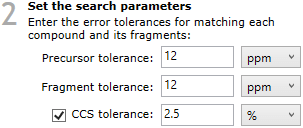

The Metabolic Profiling CCS Library parameter selection options.
- Precursor tolerance
-
This value, in ppm or Da, is the maximum difference between the mass for the compound in the Metabolic Profiling library and the observed mass.
Where a compound has been successfully deconvoluted, the neutral mass is searched directly against the neutral mass in the Metabolic Profiling CCS library.
Where only a single m/z is available, and its adduct status is unclear, a single neutral mass cannot be assigned for searching against the Metabolic Profiling CCS library. Instead, this compound will be searched multiple times. Each time, it is treated as if it is derived from a different adduct; the neutral mass corresponding to that original m/z and assumed adduct is calculated and searched, and this is repeated for all the adducts you defined in the experiment setup that could impart the correct observed charge state for the compound. This allows attempted identification even for compounds without a known neutral mass.
- Fragment tolerance
-
This threshold (in ppm or Da) is the maximum difference between the mass of a fragment in the Metabolic Profiling MS/MS library and the observed mass of the fragment.
The Metabolic Profiling libraries contain MS/MS data for many commonly adducted forms of compounds. Progenesis QI will search only for fragments that are derived from the adducted precursors it has determined are present for that compound, to reduce false positives (hits against fragments that would not be present as the precursor for them is not present).
- CCS within
-
This threshold (in % or Å2) is the maximum difference between the CCS for the compound in the Metabolic Profiling library and the observed CCS. For compounds with multiple adducts, only one of the adducts' CCS values needs to be within the threshold.
Because CCS calibration is only applied within a finite m/z range, adducts with m/z less than 150 Da will use a CCS threshold of 10% regardless of the user entered threshold.
Search for identifications
Clicking this button will show the Select Compound Classes window. If you do not wish to search the entire library, check the box next to Restrict IDs to the following compound classes. You'll then be able to select specific compound classes to be searched, by checking the boxes next to their names.
You can use the Find box to search for the compound class you're looking for and the results get highlighted in the list. Use the Select all and Clear all buttons to select all or none of the available classes, respectively.
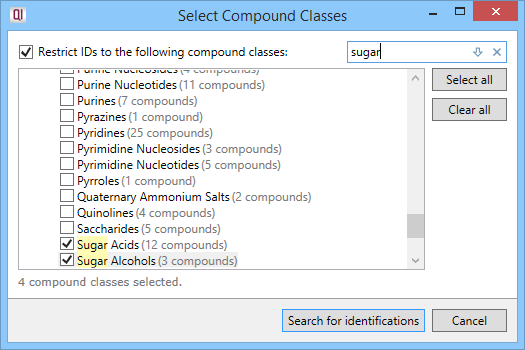
The Select Compound Classes window
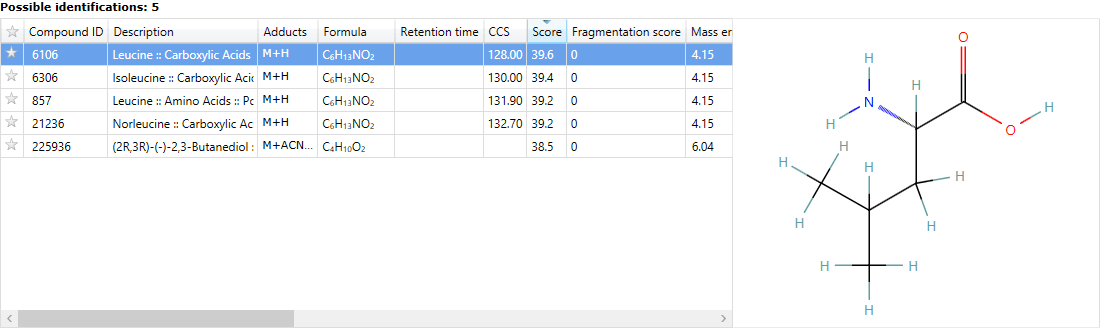
When you're done, clicking Search for identifications button will begin the search, and automatically import the results against your compounds at the Possible identifications window at the Identify Compounds screen.
You can visualise the matching of measured to database-predicted fragments for each potential ID using the 'mirror plot' on the same screen, after clicking on a search result. Matched fragments are shown in red, unmatched in grey; and hovering the cursor over a fragment match will display relevant information in a tooltip.
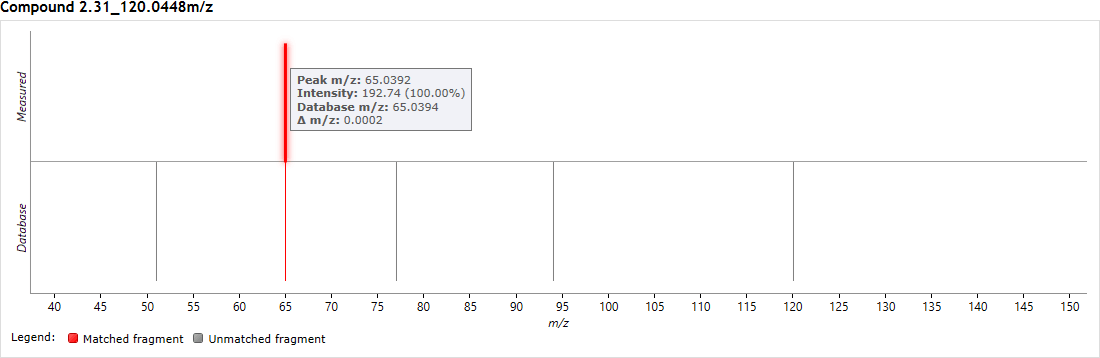
Fragment matching for Metabolic Profiling results can be visualised in a mirror plot above the Possible identifications table.