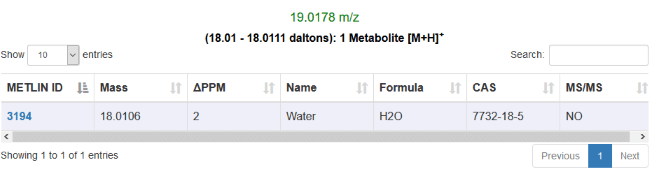
Support for METLIN batch metabolite search
Support for this compound search method is provided as standard.
About this plug-in
METLIN is a metabolite database for metabolomics. This identification method integrates with the batch search interface available on the METLIN website. It does this by copying the masses of the selected features to your clipboard, allowing you to paste them into the METLIN batch search interface.
METLIN not only provides information on the names, formulae and theoretical masses of metabolites; it also provides a link to a web page containing identifiers for the compound on various other databases such as KEGG and HMDB.
IMPORTANT: CSV export
The ability to export search results to a CSV file has been removed from the METLIN batch search interface. Unfortunately, this means it is no longer possible to import your METLIN search results into Progenesis QI.
If you are interested in this feature being reinstated, please contact Scripps.
Using this search method
- Make sure you have registered for a METLIN account.
- At the Identify Compounds screen, select the METLIN batch metabolite search method.
- Click the Copy masses to clipboard button. Progenesis QI will open the METLIN homepage in your web browser.
-
Log in to METLIN using your METLIN account:
-
In the METLIN menu bar, click Batch Search.

-
In the Masses box, right click and select Paste to paste the masses from Progenesis QI.
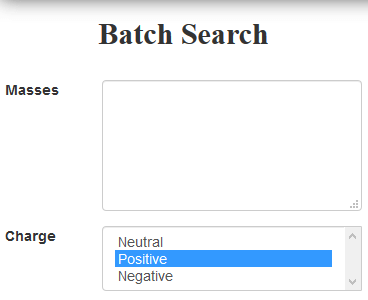
-
Click the Search button.
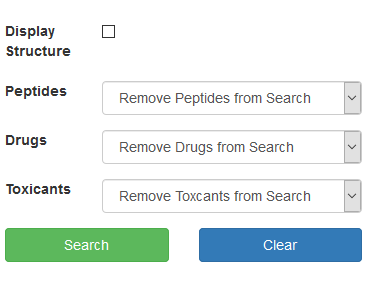
-
Your search results will then appear in the right hand panel.