How do I export compound data for external analysis?
While Progenesis QI allows you to carry out univariate and multivariate statistical tests within the software, and provides multivariate methods of visualisation of your data, it is possible you may wish to carry out different analyses to suit your specific situation. To this end, the software allows the export of your compound data. You can export whole-compound data, or even individual isotope information from all the adducted forms of the compound. To keep your visualisations and records consistent with analyses performed within the software, you can then re-import lists of compounds of interest from any external analyses back into Progenesis QI.
You may also wish to export lists of identifications in your experiment for text-based data mining, presentational or recording purposes, and this is also possible.
Exporting compound data from the software
The export options are accessed at the Identify Compounds or Review Compounds stages, using the File menu:
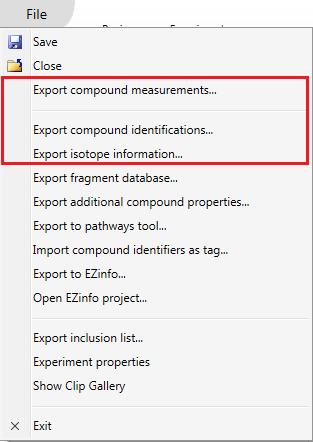
The data export options available. Note that EZinfo only appears if this additional statistical software is also installed. Those relevant to this FAQ are highlighted by the box.
The options include:
- Export compound measurements… – produce a csv file containing compound abundances, properties, and information on differences in the compounds across experimental conditions
- Export compound identifications… – produce a csv file containing compound identifications and properties
- Export isotope information… – produce an xml file containing m/z and abundance information for every isotope of every adducted form of every compound
Each option has its own dedicated FAQ, linked from the list.
This FAQ deals with the export of compound and identification data for general purposes, and so only covers the boxed export options from the file menu on the figure above. For more on fragment databases or additional compound properties, which support the generation of identifications in the first place, see the existing FAQs.
Likewise, the specific external analysis options you can see in the figure – pathways tools and EZinfo – have their own dedicated FAQ pages.
Are all my data exported?
For compounds, all those matching the current tag filter have their data exported. If you have not applied a tag filter, all compounds have their data exported. If you have applied one, but wish to export everything, you should clear the filter first.
For the data fields associated with each compound, some apply to all runs, such as retention time. However, some fields, such as normalised abundance, are run-specific. In these cases, such run-specific data are exported only for runs in your current experiment design. If you wish to see the data for all runs, but have an experiment design active that only includes some of them, then you may wish to switch to (or make and switch to) one that includes them all prior to export. Likewise, some fields, such as ANOVA (p) values, depend on the currently active experiment design, and will update as it is altered – this is worth bearing in mind if revisiting any exports later.
Importing compound lists back into the software
Once a list of compound identifiers of interest has been generated externally, you can re-import your results as a tag. This allows you to record externally derived results alongside your software-derived results, and visualise them together. To do this, create a simple text file with the relevant compound identifiers listed in a column, and select File | Import compound identifiers as tag... as shown below at the Identify compounds or Review compounds stages. A pop-up dialog will guide you through the process.
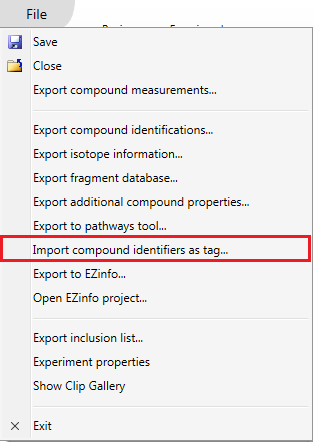
The Import compound identifiers as tag option on the file menu.
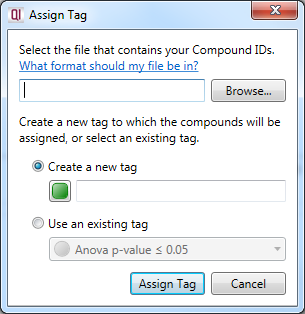
The dialog box that will appear on selecting the Import compound identifiers as tag option.
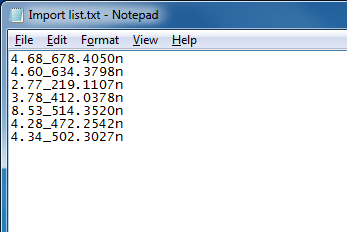
A simple text-based file suitable for compound identifier import, with format corresponding to those used for the compound in the software.






