Support for Progenesis MetaScope
Support for this compound search method is provided as standard.
About this plug-in
This identification method is intended to support identifications from a number of different compound databases. It allows you to choose a .SDF, .CSV, .XLS or .XLSX file containing identifications and information on a selection of compounds, and search for matches by neutral mass.
It also allows you to search retention time and CCS values using additional properties files, and fragmentation data using either theoretical fragmentation patterns or fragment databases.
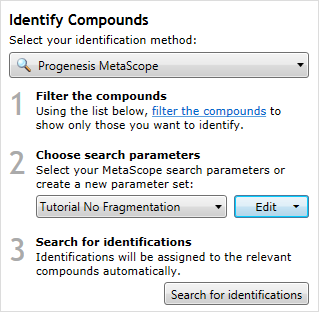
The MetaScope search method, with example parameters chosen.
The use of this method has the following steps:
- In the Identify Compounds screen, filter the list of compounds to show only those that you want to identify.
- Choose a set of search parameters from the Choose search parameters drop-down (you might need to first create or edit a search parameter set. See the section on search parameters below for how to do this).
- Click the Search for identifications button.
When the import and search is complete, a prompt will tell you how many compounds have been identified. You can then continue with your analysis, identifying further compounds if necessary by searching with other parameter sets.
Search parameters
MetaScope comes with a number of built-in search parameter sets, to show you the kind of parameters you can set. However, you'll probably want to create your own or modify the built in parameter sets.
To edit or create a new search parameter set, choose the appropriate option in the drop-down to the right of the parameter set name:
This will pop up a dialog where you can fill in the various search parameters:
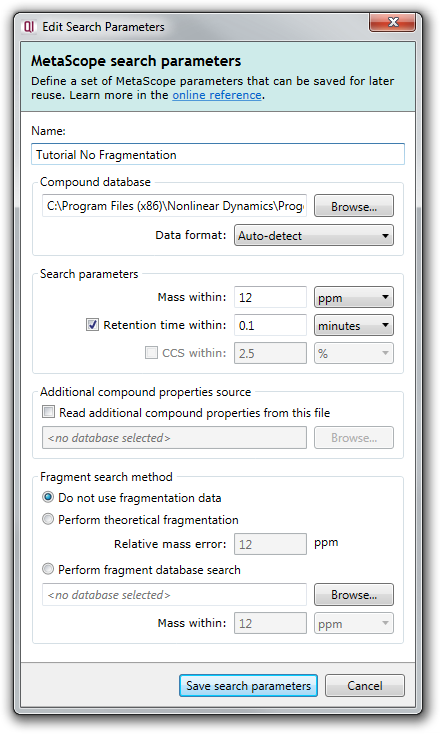
Briefly, the options available to you are:
- Name
- Give the search parameter set a useful name to refer to later.
- Compound database
- Choose either an SDF or CSV/XLS/XLSX file containing compound information.
- Mass within
- Choose the threshold (in ppm or Da) within which the neutral mass must lie to constitute a match.
- Retention time within
- Choose the threshold (in minutes or %) within which the retention time must lie to constitute a match. Can be turned on or off using the checkbox.
- CCS within
-
Choose the threshold (in % or Å2) within which the CCS must lie to constitute a match. Can be turned on or off using the checkbox. Will be disabled if the experiment contains no CCS data.
Note: for compounds with multiple adducts, only one of the adducts' CCS values needs to be within the threshold. Also, because CCS calibration is only applied within a finite m/z range, adducts with m/z less than 150 Da will use a CCS threshold of 10% regardless of the user entered threshold.
- Additional compound properties source
- Allows you to load additional compound properties (e.g. RT and CCS) from an external file. See What are additional compound properties files? for more information.
- Fragment search method
- Allows you to choose between no fragmentation search, search against theoretical fragmentation, or search against a fragment database. See What are fragment databases? for more information.
After entering your search parameters, click Save search parameters to save your search parameter set for use in this and future experiments.
SDF Files
The SDF format contains structural information and other associated data about the metabolites in the database.
Here are some good sources of metabolite SDF databases which can be searched with MetaScope:
Dealing with problems
If your SDF database file cannot be read in full, the following dialog is displayed:
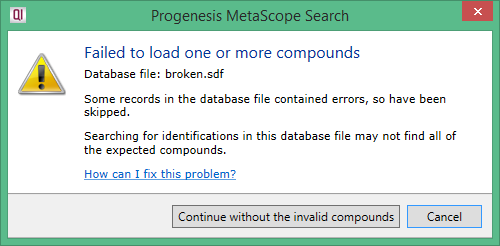
You can choose to continue the search without the components that couldn't be read, although the search may not find all of the expected compounds this way. The Progenesis SDF Studio can help you fix the problems with the SDF file.
Excel File Format
If you are using a .CSV, .XLS or .XLSX file for MetaScope identifications, it should contain the following fields:
| Field Name | Required? |
|---|---|
| Neutral Mass | Yes |
| Compound Id | Yes |
| Description | No |
| Formula | No |
| Url | No |
| Retention Time (min) | If Retention time within checked. |
Field names are case insensitive and can occur in any order.
You can also download an example database.
Note: you will need to have Microsoft Excel 2007 (or later) installed to perform the search.
MSP Files
MSP files contain information about fragmentation masses and intensities for a set of precursors. In MetaScope, you can use an MSP file either as a standalone compound database, or you can use it to provide experimental fragmentation information for an SDF or an Excel database.
To use an MSP as a standalone database, it needs several specific fields.
| Field Name | Purpose |
|---|---|
| Name | Required - uniquely identifies the compound. |
| Formula | Used to calculate a neutral mass. With a neutral mass, search results can be found regardless of the adduct ion form. |
| PrecursorMZ | Required if Formula is not provided, used to search for possible matching precursor ions. |
| Precursor_type | The name of the adduct ion which was detected in an experimental database. Allows a neutral mass to be calculated even if Formula is not provided. The values in this field will need to match the names of the adducts you've specified in your experiment. |
| Charge | If provided, only matches this entry against experimental compounds with a matching charge. |
| Num Peaks | Required - the number of fragments in this entry. |
Field names are case insensitive. Num Peaks must be the last entry before the list of fragments, but otherwise they can be in any order.
You can create your own fragment databases in Progenesis QI. You can also download an example database.




