How can I edit the results of peak picking?
If whilst reviewing your data in Review Deconvolution you’ve found an example where the peak picking has missed an isotope for an adduct ion, you’ll want to edit this following the instructions below:
- At Review Deconvolution add the adduct where the isotope has been missed to the correct compound by right-clicking and selecting Add to Compound.
- Tag the compound and apply a filter so that only this compound is visible.
- Editing can only be performed on single-adduct compounds, so you’ll need to remove the adduct you wish to edit from the compound by right-clicking on it and selecting Remove from Compound — you can leave the other adducts that don’t require editing as part of the compound. The table will now display 2 tagged compounds one of which is the single adduct requiring editing that you just removed.
- Click on Review Compounds at the top of the screen to change stages in the workflow.
- Select the adduct that requires editing from the table and then click Review selected compound.
- Click on Edit adduct to access the editing tools and then click on the plus sign that appears above the missing isotope to add it in:
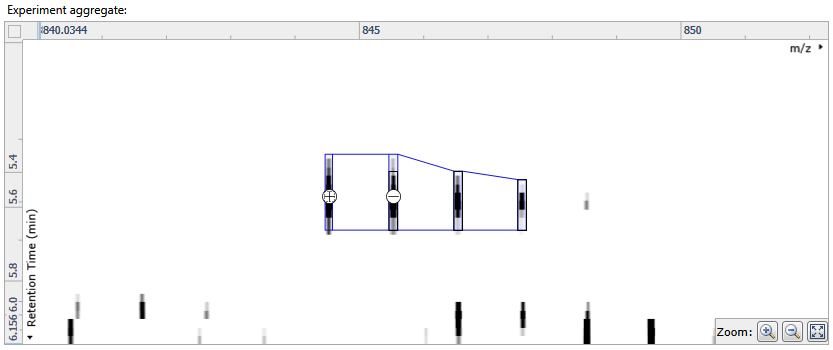
- Click on Save edits to save your changes.
- Go back to Review Deconvolution and add the edited adduct to the compound as described in step 1 – the table will update to list only the single compound.
- Finally, clear the filter to display all compounds.






