Exporting peptide ion measurements for external analysis
There are two stages within the Progenesis QI for proteomics workflow where you can export peptide ion data: pre-identification and post-identification. The content of these two exports are referred to as peptide ion data and peptide ion measurements, respectively. As you'll read below, the former of these exports can include identification data...
Export peptide ion data
"Peptide ions" correspond to isotopic clusters derived from a single charge state of a given peptide and depending on where you are in your workflow, may be identified or unidentified. The Export peptide ion data export allows you to export any peptide ions, regardless of whether or not they have an identification.
This export is available at the Review Peak Picking, Peptide Ion Statistics and Identify Peptides windows by selecting File | Export peptide ion data... as shown below:
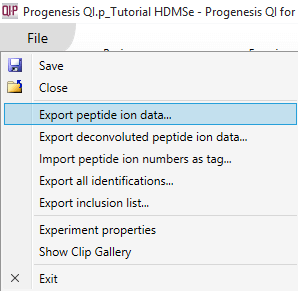
The export can be customised by using the list that then appears. Data are available on the nature of the peptide ion itself (including m/z, retention time, mass and charge, for example, based on the aggregate); on the inter-group differences associated with the peptide ion and its variability (including the ANOVA value for the current experiment design group comparison for that peptide ion, maximum fold change for that design across any two groups, highest and lowest group mean, and the maximum group CV); and quantitative peptide ion data. The raw and normalised abundance are the key columns in this last category for external analyses, and spectral count data are available for the highest-scoring identification associated with the peptide ion. Unchecking the final tick-box guarantees that you will export everything, by ignoring the current tag filter.
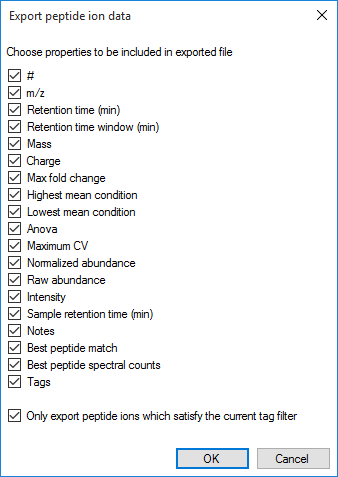
The column selection dialog box for Export peptide ion data.
A csv (comma-separated values) file is then generated; you will be given a pop-up dialog prompting you to name and save the file, and then the option to open the file immediately.
In the file generated by the export, the rows are your peptide ions (isotopic groups of peaks corresponding to a single peptide ion). These are as numbered by the software, for example at the Review Peak Picking and Peptide Ion Statistics windows. The columns contain the selected data outputs for each peptide ion.
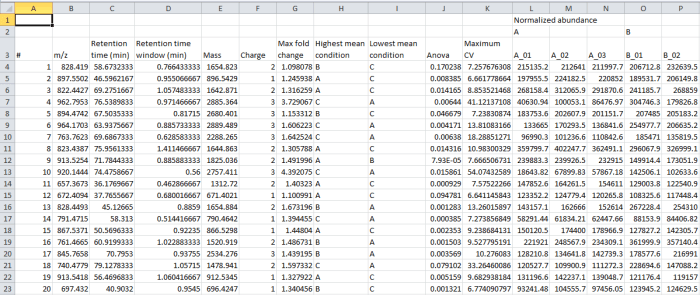
Screenshot of the csv export resulting from the Export peptide ion data option.
Export peptide ion measurements
In this peptide-ion based export each row is a single charge state of a now-identified peptide ion. With the focus on identifications, the ions are grouped by the peptide sequence they are derived from.
This export can be accessed in a similar manner to the Export peptide ion data export, at the Review Proteins and Protein Statistics windows, by selecting File | Export peptide ion measurements... from the menu.
Alternatively, use the Export peptide ion measurements button in step 4 on the side-bar at the Review Proteins window:

The data columns available include quantitative information on the peptide ion, identification data and also the spectral counting data for each peptide, as below:
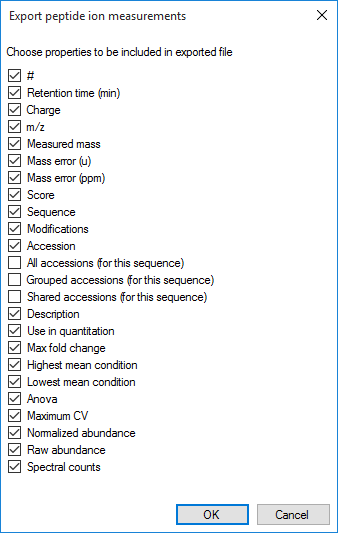
The column selection dialog box for Export peptide ion measurements.
The Use in quantitation column indicates whether a peptide ion was selected for use in protein abundance calculation by the quantitation scheme being employed (for example, using unique peptides only or Hi-N will result in only a subset of peptide ions contributing).
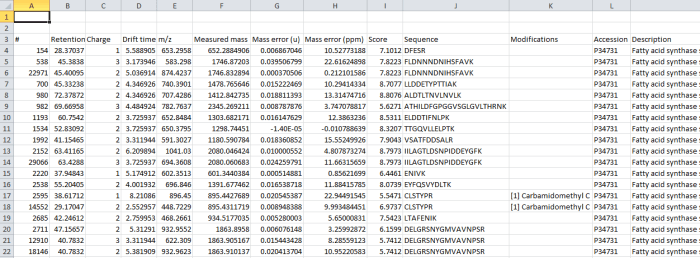
Screenshot of the csv export resulting from the Export peptide ion measurements option.
Re-importing your results from external analyses
For both of the peptide ion exports, once you have carried out your external analyses, you can re-import your results as a tag. This allows you to record externally derived results alongside your software-derived results, and visualise them together. To do this, create a simple text file with the relevant peptide ion numbers listed in a column, and select File | Import peptide ion numbers as tag... as shown below. The pop-up box will guide you through the process.
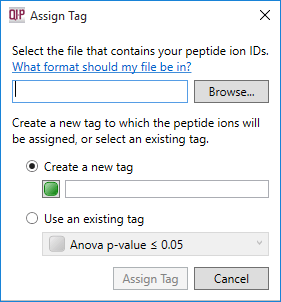
The dialog box that will appear on selecting the Import peptide ion numbers as tag option.
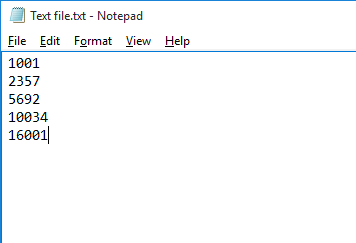
A simple text-based file suitable for peptide ion import with numerical identifiers corresponding to those used for the peptide ion in the software.





