Exporting protein measurements for external analysis
Once you have identified proteins present and the software has 'rolled up' peptide-ion-based data to the protein level, it is possible to export protein-level results for external analyses in a csv format.
These data can be exported at the Review Proteins and Protein Statistics stages by selecting File | Export protein measurements... from the menu.
Alternatively, they can be exported at the Review Proteins window by using the Export protein measurements button in step 4 on the side-bar:

In this case, the rows are your proteins listed by accession. The columns contain the selected outputs for that protein. These can include quantitative protein data as provided at the Review Proteins and Protein Statistics windows, along with spectral counting MS2 data where appropriate, as shown below. The normalised abundance is available in this export for external analysis.
The column selection dialog box for Export protein measurements.
Note: the Spectral counts tick-box is not available if you are quantitating proteins using the Hi-N method, as these data would be misleading when only a limited number of the peptides are being considered. If you do wish to export these data, switch to either relative quantitation using non-conflicting (unique) peptides or relative quantitation using all peptides at the Protein options dialog.
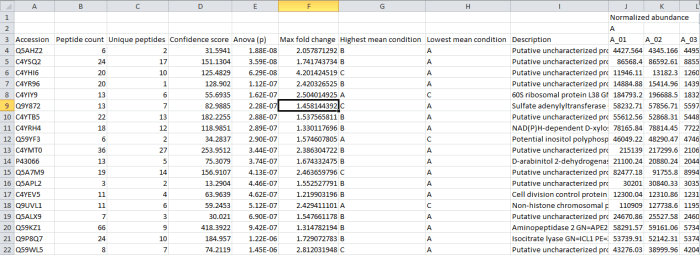
Screenshot of the csv export resulting from the Export protein measurements option.
Re-importing your results
Analogously to the Import peptide ion numbers as tag... option, it is possible to bring back into the software a list of proteins of interest derived from external analyses. This is done by creating a list of accessions, and then selecting File | Import protein accessions as tag... from the menu. Note that tags at the protein and peptide ion level are independent.
The dialog box that will appear on selecting the Import protein accessions as tag option.





